This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox bcce04
From Proteopedia
(Difference between revisions)
| (17 intermediate revisions not shown.) | |||
| Line 2: | Line 2: | ||
<StructureSection load='1iat' size='450' side='right' scene='' caption=''> | <StructureSection load='1iat' size='450' side='right' scene='' caption=''> | ||
| - | '''Imaginaryase''' (alternatively known as '''imaginary isomerase''' or '''imaginary- | + | '''Imaginaryase''' (alternatively known as '''imaginary isomerase''' or '''imaginary-6-phosphate isomerase''') are a group of enzymes of the isomerase family ([[EC]] 5.3.a.b), so named for their main function in metabolism. In this pathway imaginary isomerase (IGI) is used to inter-convert imaginary-6-phosphate and reality-6-phosphate. |
| - | + | Imaginaryase is also known for a list of activities: | |
| - | * | + | * Something cool <ref>2</ref>. |
| - | * | + | * Something cooler<ref>3</ref>. |
| - | * | + | * Something my prof thinks is cool <ref>4</ref>. |
| - | + | * Something really boring <ref>5</ref>. | |
| - | + | ||
| - | * | + | |
| - | + | ||
| - | + | ||
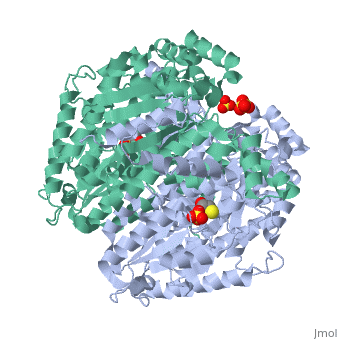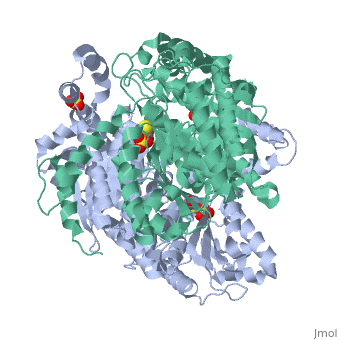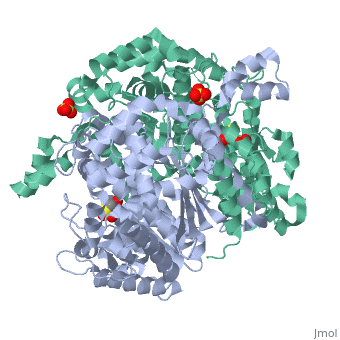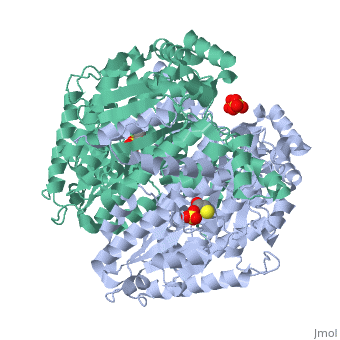
| - | Phosphoglucose isomerase exists in the cell usually as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Dimer/1'>homodimer</scene>, nevertheless outside of the cell, it has been isolated as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Monomer/1'>monomeric</scene> structure. PGI has essentially an identical fold in all of the characterized species (see '''Figure 1'''). The <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Sec_struct/1'>secondary structure</scene> of phosphoglucose isomerase is charaterized by an αβα conformation, on each of its two domains. The smaller domain is characterized by 5 parallel β-sheets, while the larger domain if formed out of 6 parallel/antiparallel β-sheets. Furthermore, another characteristic trait is a residue extension at the C-terminus, which wraps around the other monomer in the dimeric conformation. A "hook" that can potentially be involved in the previously mentioned extracellular activities. | ||
| - | + | =='''Structure'''== | |
| - | + | Imaginary isomerase exists in cells usually as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Dimer/1'>homodimer</scene>, nevertheless outside of cells, it has been isolated as a <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Monomer/1'>monomeric</scene> structure. The <scene name='Stancu_Phosphoglucoisomerase_Sandbox_1/Sec_struct/1'>secondary structure</scene> of imaginary isomerase is characterized by a conformation, on each of its two domains. Fill in the information here to give more detail. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
</StructureSection> | </StructureSection> | ||
__NOTOC__ | __NOTOC__ | ||
=='''Mechanism'''== | =='''Mechanism'''== | ||
| - | The proposed reaction mechanism of PGI for the reversible conversion of glucose-6-phosphate to fructose 6-phosphate involves an acid/base catalysis by the enzyme. The basic mechanism involves the isomerization of an aldose to a ketose. This is performed by a ring opening, followed by an isomeration of the opened ring, then a ring closing. A detailed step by step mechanism of this process can be seen as follows <ref> Voet D, Voet J, and Pratt C. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. Print.</ref>: | ||
| - | |||
| - | '''Step 1.''' The substrate binds to the enzyme. | ||
| - | |||
| - | '''Step 2.''' The residue Lys518 (or His388b) acts as an enzymatic acid catalyzing the opening of the ring. | ||
| - | |||
| - | '''Step 3.''' Conserved Glu357 abstracts the acidic proton from C2 forming a cis-enendiol intermediate. | ||
| - | |||
| - | '''Step 4.''' Glu357 donates back the proton at the C1 position. | ||
| - | |||
| - | '''Step 5.''' Lys518 (or His388b) abstracts back the proton from the sugar ring oxygen, resulting in a ring closure, to give the product. | ||
| - | |||
| - | [[Image:Mech.png]] | ||
=='''Regulation and Inhibition'''== | =='''Regulation and Inhibition'''== | ||
| - | '''Regulation''' of phosphoglucoisomerase is only done by the relative concentrations of glucose-6-phosphate and fructose 6-phosphate, towards equilibrium. Nevertheless, it was found that the kinetic parameters of PGI does depend on the pH and temperature of the environment. | ||
| - | The following kinetic parameters are proposed for rabbit PGI at pH 8.5 and 30°C <ref>PMID: 5647261</ref> | ||
| + | ==3D structures of Imaginary isomerase== | ||
| - | + | ==Additional Resources== | |
| - | |||
| - | It is interesting to point the regulation of PGI in other aspects that are not involved in metabolism. For example, PGI acts as a "cytokine" outside the cell in that it can be used as a cell signalling protein. PGI has been found to to be associated with AMF cells, which is found to regulate tumor cell motility. Regulation of these extracellular "cytokine" PGI/AMF can be seen. The amount of PGI/AMF that is secreted inside and outside the cell based on infection <ref> Funasaka T, Hu H, Yanagawa T, Hogan V, Raz A. Down-Regulation of Phosphoglucose Isomerase/Autocrine Motility Factors Results in Mesenchymal-to-Epithelial Transition of Human Lung Fibrosarcoma Cells. (2007) Cancer Res, 76(9) </ref>. | ||
| - | |||
| - | |||
| - | '''Inhibition''' of the phosphoglucoisomerase regulated reaction of glucose-6-phosphate to fructose-6-phosphate can also occur. Competitive competition can take place from inhibitors such as 5PAH. 5PAH resembles PGI, differing only in a nitrogen atom at the first carbon position. 5PAH is reported to have a Ki of .0000002 M <ref> Arsenieva D, Hardre R, Salmon L, Jeffery CJ. The crystal structure of rabbit phosphoglucose isomerase complex with 5-phospho-D-arabinonohydroxamic acid. (2002),PNAS, 99(9) </ref>. | ||
| - | =='''Links'''== | ||
| - | |||
| - | |||
| - | * Crystal Structure of human phosphoglucose isomerase (PDB=[[1iat]]) | ||
| - | |||
| - | * Crystal Structure of rabbit phosphoglucose isomerase complexed fructose 6-phosphate (PDB=[[1hox]]<ref>PMID:11425306</ref>) | ||
| - | |||
| - | * Other available structures [[1dqr]], [[1g98]], [[1gzd]], [[1gzv]], [[1hm5]], [[1iri]], [[1jiq]], [[1jlh]], [[1koj]], [[1n8t]], [[1nuh]], [[1xtb]]. | ||
| - | |||
| - | ==3D structures of phosphoglucose isomerase== | ||
| - | |||
| - | ===PGI=== | ||
| - | |||
| - | [[2pgi]], [[1b0z]] – GsPGI – ''Geobacillus stearothermophilus''<br /> | ||
| - | [[1dqr]], [[1hm5]], [[1n8t]] – rPGI – rabbit<br /> | ||
| - | [[1iat]], [[1jlh]] – hPGI – human<br /> | ||
| - | [[1qxj]], [[1x8e]], [[3sxw]] – PfPGI – ''Pyrococcus furiosus''<br /> | ||
| - | [[1j3p]], [[1j3q]] – TlPGI – ''Thermococcus litoralis''<br /> | ||
| - | [[1q50]] – LmPGI – ''Leishmania Mexicana''<br /> | ||
| - | [[1u0e]], [[2cvp]] – mPGI – mouse<br /> | ||
| - | [[2q8n]] – PGI – ''Thermotoga maritima''<br /> | ||
| - | [[3hjb]] – PGI – ''Vibrio cholera''<br /> | ||
| - | [[3ifs]] – PGI – ''Bacillus anthracis''<br /> | ||
| - | [[2wu8]] – PGI – ''Mycobacterium tuberculosis''<br /> | ||
| - | [[3ljk]] – FtPGI (mutant) – ''Francisella tularensis''<br /> | ||
| - | [[3nbu]] – PGI – ''Escherichia coli'' | ||
| - | |||
| - | ===PGI complex with fructose-6-phosphate=== | ||
| - | |||
| - | [[1hox]] – rPGI + fructose-6-phosphate<br /> | ||
| - | [[1t10]] - LmPGI + fructose-6-phosphate<br /> | ||
| - | [[2gc2]] - PfPGI + fructose-6-phosphate<br /> | ||
| - | [[2cxs]], [[2cxt]] - mPGI + fructose-6-phosphate<br /> | ||
| - | [[3m5p]] - FtPGI (mutant) + fructose-6-phosphate<br /> | ||
| - | |||
| - | ===PGI complex with sorbitol-6-phosphate=== | ||
| - | |||
| - | [[1xtb]] - rPGI + sorbitol-6-phosphate<br /> | ||
| - | [[2gc1]] - PfPGI + sorbitol-6-phosphate<br /> | ||
| - | [[2cxq]] - mPGI + sorbitol-6-phosphate<br /> | ||
| - | |||
| - | ===PGI complex with phosphoarabinose=== | ||
| - | |||
| - | [[1gzd]], [[1gzv]] – PGI + phosphoarabinose – pig<br /> | ||
| - | [[1c7r]] - GsPGI + phosphoarabinose<br /> | ||
| - | [[2cxp]] – mPGI + phosphoarabinose<br /> | ||
| - | [[1nuh]] - hPGI + phosphoarabinose<br /> | ||
| - | [[1qsr]], [[1x7n]], [[1x82]] - PfPGI + phosphoarabinose<br /> | ||
| - | [[2gc0]] - PfPGI + phosphoarabinose derivative<br /> | ||
| - | [[1koj]] – rPGI + phosphoarabinose derivative | ||
| - | |||
| - | ===PGI complex with gluconate-6-phosphate=== | ||
| - | |||
| - | [[1qy4]] - PfPGI + gluconate-6-phosphate<br /> | ||
| - | [[1j3r]] - TlPGI + gluconate-6-phosphate<br /> | ||
| - | [[2cxr]] - mPGI + gluconate-6-phosphate<br /> | ||
| - | [[3q7i]] - FtPGI (mutant) + gluconate-6-phosphate<br /> | ||
| - | |||
| - | ===PGI complex with glucose-6-phosphate=== | ||
| - | |||
| - | [[1u0f]] - mPGI + glucose-6-phosphate<br /> | ||
| - | [[3ff1]] - PGI + glucose-6-phosphate – ''Staphylococcus aureus''<br /> | ||
| - | [[2o2c]] - TbPGI + glucose-6-phosphate – ''Trypanosoma brucei'' | ||
| - | |||
| - | ===Other PGI binary complexes=== | ||
| - | |||
| - | [[2o2d]] – TbPGI + citrate<br /> | ||
| - | [[1u0g]], [[2cxo]] - mPGI + erythrose-4-phosphate<br /> | ||
| - | [[3q88]] – FtPGI (mutant) + ribose bisphosphate<br /> | ||
| - | [[1c7q]] – GsPGI + phosphate inhibitor<br /> | ||
| - | [[2cxn]], [[2cxu]] – mPGI + phosphate<br /> | ||
| - | [[1g98]] – rPGI + transition state analog<br /> | ||
| - | [[2gc3]] - PfPGI + mannose-6-phosphate<br /> | ||
| - | |||
| - | ==Additional Resources== | ||
| - | For additional information, see: [[Carbohydrate Metabolism]] | ||
| - | <br /> | ||
==References== | ==References== | ||
<references/> | <references/> | ||
Current revision
Imaginary Protein
| |||||||||||