This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Priyanka Basak/Sandbox
From Proteopedia
(Difference between revisions)
| (30 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | == | + | ==Introduction== |
| - | + | <StructureSection load='3tj8' size='340' side='right' caption='''Hp''UreE,PDB ID: 1TJ8 ' scene=''> | |
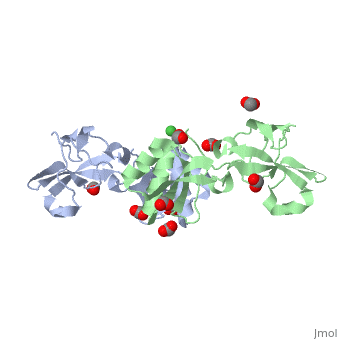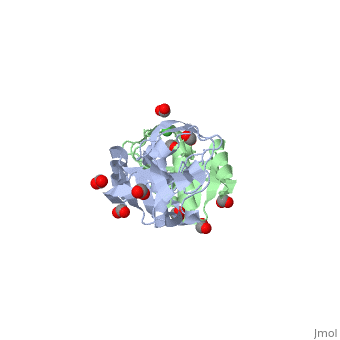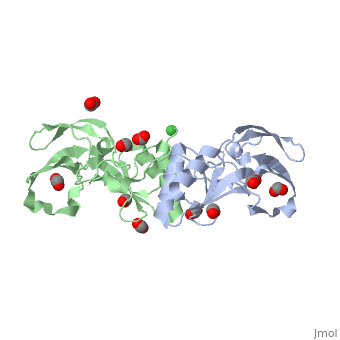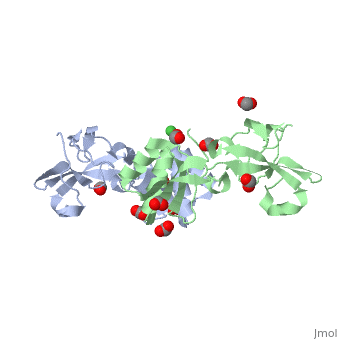
| - | <StructureSection load='3tj8' size='340' side='right' caption=' | + | UreE is an essential metallochaeprone for ''Helicobacter pylori'' (''H.pylori''). ''H.pylori'' is a class I human pathogen that is involved in causing gastric ulcers and stomach cancers. Its two nickel dependent enzymes - Urease and NiFe-H2ase, provides it a unique ability to survive and colonize in acidic human stomach. Urease catalyzes the conversion of urea to ammonia and carbamate which neutralizes its local acidic environment. |
| - | + | ||
| - | + | ||
| - | + | ||
== Function == | == Function == | ||
| - | + | Several accessory proteins play a crucial role in the maturation of Urease. UreE is a urease-specific nickel metallochaperone that is involved in the delivery nickel to urease. | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
== Structural highlights == | == Structural highlights == | ||
| + | Apo-UreE from ''H.pylori'' is a dimer.''Hp''UreE can bind one Ni(II) per dimer <scene name='60/609775/Nickel_bound_uree/5'>Nickel bound UreE</scene>. Nickel is bound to UreE in an octahedral fashion with one His from one subunit and two His from other subunit <scene name='60/609775/Nickel_bound_uree_2/2'>(UreE-Nickel Binding site)</scene>.In presence of Ni, its oligomeric state changes to a tetramer (dimer of dimer). | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | </StructureSection> | ||
== References == | == References == | ||
| - | < | + | <http://www.ncbi.nlm.nih.gov/pubmed/20681615/> |
| - | + | [http://] | |
Current revision
Introduction
| |||||||||||