This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Nitrate reductase
From Proteopedia
(Difference between revisions)
| (14 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
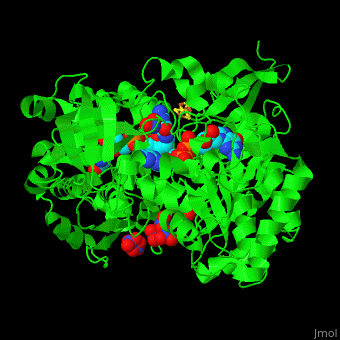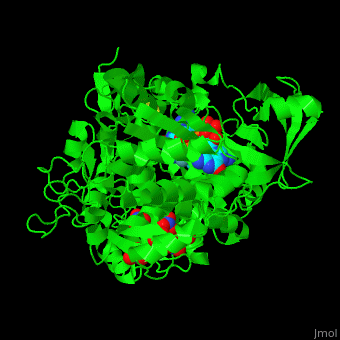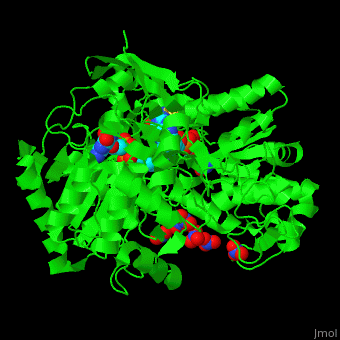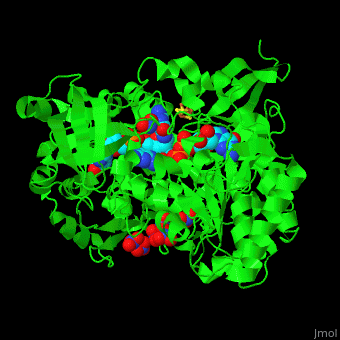
| - | + | <StructureSection load='' size='350' side='right' caption='Bacterial periplasmic dissimilatory nitrate reductase containing Fe4S4 cluster with Mo, NO3- and molybdepterin guanine nucleotide (PDB entry [[2jiq]])' scene='49/490062/Cv/3'> | |
| - | + | == Function == | |
| - | + | '''Nitrate reductase''' (NR) catalyzes the reduction of NO<sub>3</sub> to NO<sub>2</sub> using NADPH<ref>PMID:13152110</ref>. NR active site contains Mo atom. Four types of NR are known:<br /> | |
| - | + | '''eukaryotic assimilatory NR'''<br /> | |
| - | + | '''bacterial cytoplasmic assimilatory''' (NAC)<br /> | |
| - | + | '''bacterial membrane-bound respiratory''' (NAR)<br /> | |
| - | '''Nitrate reductase''' (NR) catalyzes the reduction of | + | '''bacterial periplasmic dissimilatory''' (NAP).<br /> |
| + | The cofactors of NR are Molybdopterin (MPT) in eukaryotic NR and bis-Molybdopterin guanine dinucleotide (MGD) in bacterial NR. | ||
| + | |||
| + | == Structural highlights == | ||
| + | <scene name='49/490062/Cv/7'>Bacterial periplasmic dissimilatory nitrate reductase containing Fe4S4 cluster with Mo, NO3 and 2 molybdepterin guanine nucleotides</scene>. NAP active site includes a <scene name='49/490062/Cv/8'>six-coordinated Mo atom which coordinates to 2 MGD molecules and Cys140</scene>. The <scene name='49/490062/Cv/9'>NO3 molecule is guided to the active site by a funnel leading to the Mo atom</scene><ref>PMID:18327621</ref>. | ||
| + | |||
| + | </StructureSection> | ||
==3D structures of nitrate reductase== | ==3D structures of nitrate reductase== | ||
| Line 21: | Line 27: | ||
**[[1ogy]] - NAP + heme + MGD + Mo– ''Rhodobacter sphaeroides''<br /> | **[[1ogy]] - NAP + heme + MGD + Mo– ''Rhodobacter sphaeroides''<br /> | ||
**[[2nya]] - EcNAP + MGD + Mo – ''Escherichia coli''<br /> | **[[2nya]] - EcNAP + MGD + Mo – ''Escherichia coli''<br /> | ||
| - | **[[3ml1]], [[3o5a]] - NAP catalytic subunit + NAPB + heme + MGD + Mo – ''Ralstonia eutropha'' | + | **[[3ml1]], [[3o5a]] - NAP catalytic subunit + NAPB + heme + MGD + Mo – ''Ralstonia eutropha''<br /> |
| + | **[[5yly]] – NR cytochrome b5 reductase domain + FAD – ''Ulva prolifera''<br /> | ||
*Bacterial respiratory NR | *Bacterial respiratory NR | ||
| Line 36: | Line 43: | ||
**[[2bih]], [[2bii]] – NR MoCo-binding domain + MPT – ''Pichia angusta'' | **[[2bih]], [[2bii]] – NR MoCo-binding domain + MPT – ''Pichia angusta'' | ||
| + | |||
| + | *Eukaryotic NR | ||
| + | |||
| + | **[[1cne]] – cNR + FAD – corn<br /> | ||
| + | **[[1cnf]], [[2cnd]] – cNR (mutant) + FAD + ADP<br /> | ||
| + | |||
}} | }} | ||
| + | == References == | ||
| + | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D structures of nitrate reductase
Updated on 23-July-2019
References
- ↑ NICHOLAS DJ, NASON A. Molybdenum and nitrate reductase. II. Molybdenum as a constituent of nitrate reductase. J Biol Chem. 1954 Mar;207(1):353-60. PMID:13152110
- ↑ Najmudin S, Gonzalez PJ, Trincao J, Coelho C, Mukhopadhyay A, Cerqueira NM, Romao CC, Moura I, Moura JJ, Brondino CD, Romao MJ. Periplasmic nitrate reductase revisited: a sulfur atom completes the sixth coordination of the catalytic molybdenum. J Biol Inorg Chem. 2008 Mar 8;. PMID:18327621 doi:10.1007/s00775-008-0359-6