This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1d66
From Proteopedia
(Difference between revisions)
| (6 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| + | |||
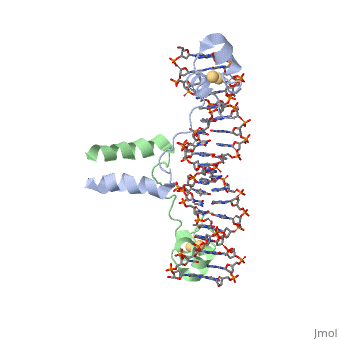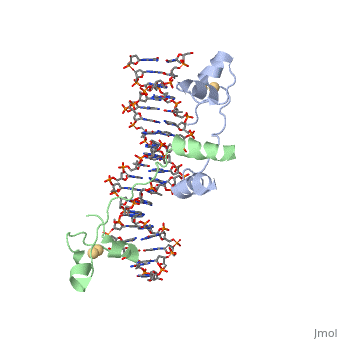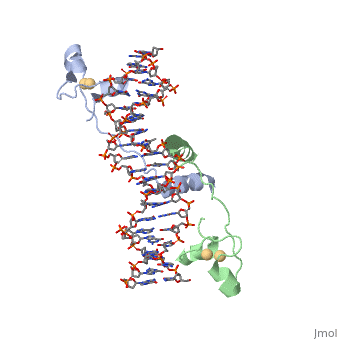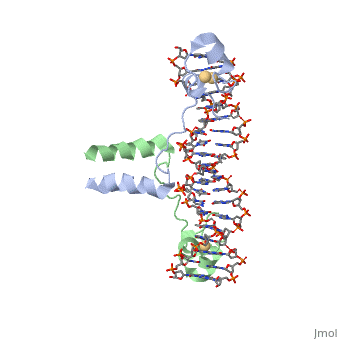
==DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX== | ==DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX== | ||
| - | <StructureSection load='1d66' size='340' side='right' caption='[[1d66]], [[Resolution|resolution]] 2.70Å' scene=''> | + | <StructureSection load='1d66' size='340' side='right'caption='[[1d66]], [[Resolution|resolution]] 2.70Å' scene=''> |
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1d66]] is a 4 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1d66]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1D66 OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1D66 FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=CD:CADMIUM+ION'>CD</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.7Å</td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CD:CADMIUM+ION'>CD</scene></td></tr> |
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1d66 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1d66 OCA], [https://pdbe.org/1d66 PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1d66 RCSB], [https://www.ebi.ac.uk/pdbsum/1d66 PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1d66 ProSAT]</span></td></tr> | ||
</table> | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/GAL4_YEAST GAL4_YEAST] This protein is a positive regulator for the gene expression of the galactose-induced genes such as GAL1, GAL2, GAL7, GAL10, and MEL1 which code for the enzymes used to convert galactose to glucose. It recognizes a 17 base pair sequence in (5'-CGGRNNRCYNYNCNCCG-3') the upstream activating sequence (UAS-G) of these genes. | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
Check<jmol> | Check<jmol> | ||
<jmolCheckbox> | <jmolCheckbox> | ||
| - | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/d6/1d66_consurf.spt"</scriptWhenChecked> | + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/d6/1d66_consurf.spt"</scriptWhenChecked> |
<scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
<text>to colour the structure by Evolutionary Conservation</text> | <text>to colour the structure by Evolutionary Conservation</text> | ||
</jmolCheckbox> | </jmolCheckbox> | ||
| - | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/ | + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1d66 ConSurf]. |
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
| - | A specific DNA complex of the 65-residue, N-terminal fragment of the yeast transcriptional activator, GAL4, has been analysed at 2.7 A resolution by X-ray crystallography. The protein binds as a dimer to a symmetrical 17-base-pair sequence. A small, Zn(2+)-containing domain recognizes a conserved CCG triplet at each end of the site through direct contacts with the major groove. A short coiled-coil dimerization element imposes 2-fold symmetry. A segment of extended polypeptide chain links the metal-binding module to the dimerization element and specifies the length of the site. The relatively open structure of the complex would allow another protein to bind coordinately with GAL4. | ||
| - | |||
| - | DNA recognition by GAL4: structure of a protein-DNA complex.,Marmorstein R, Carey M, Ptashne M, Harrison SC Nature. 1992 Apr 2;356(6368):408-14. PMID:1557122<ref>PMID:1557122</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
==See Also== | ==See Also== | ||
| - | *[[FirstGlance in Jmol|FirstGlance in Jmol]] | ||
*[[Gal3-Gal80-Gal4|Gal3-Gal80-Gal4]] | *[[Gal3-Gal80-Gal4|Gal3-Gal80-Gal4]] | ||
*[[Hydrogen in macromolecular models|Hydrogen in macromolecular models]] | *[[Hydrogen in macromolecular models|Hydrogen in macromolecular models]] | ||
| - | *[[User:Eric Martz/Introduction to Structural Bioinformatics I|User:Eric Martz/Introduction to Structural Bioinformatics I]] | ||
| - | *[[User:Eric Martz/Introduction to Structural Bioinformatics I%2C 2014|User:Eric Martz/Introduction to Structural Bioinformatics I%2C 2014]] | ||
| - | *[[User:Eric Martz/JSmol Notes|User:Eric Martz/JSmol Notes]] | ||
| - | *[[User:Wayne Decatur/Biochem642 Molecular Visualization 2010 Fall Sessions|User:Wayne Decatur/Biochem642 Molecular Visualization 2010 Fall Sessions]] | ||
| - | *[[User:Wayne Decatur/Biochem642 Molecular Visualization Sessions|User:Wayne Decatur/Biochem642 Molecular Visualization Sessions]] | ||
| - | *[[User:Wayne Decatur/UNH BCHEM833 Structural Analysis Workshop Session Fall 2012|User:Wayne Decatur/UNH BCHEM833 Structural Analysis Workshop Session Fall 2012]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| + | [[Category: Large Structures]] | ||
[[Category: Saccharomyces cerevisiae]] | [[Category: Saccharomyces cerevisiae]] | ||
| - | [[Category: Carey | + | [[Category: Carey M]] |
| - | [[Category: Harrison | + | [[Category: Harrison SC]] |
| - | [[Category: Marmorstein | + | [[Category: Marmorstein R]] |
| - | [[Category: Ptashne | + | [[Category: Ptashne M]] |
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
DNA RECOGNITION BY GAL4: STRUCTURE OF A PROTEIN/DNA COMPLEX
| |||||||||||