|
|
| (3 intermediate revisions not shown.) |
| Line 1: |
Line 1: |
| | + | |
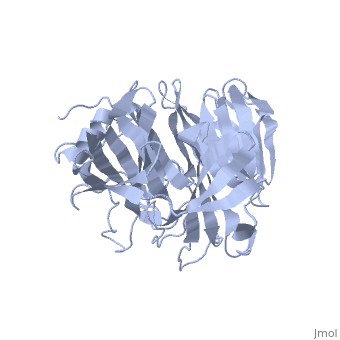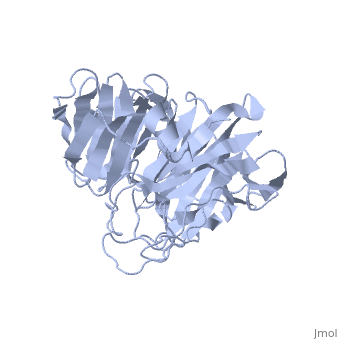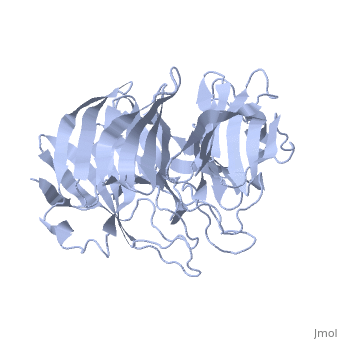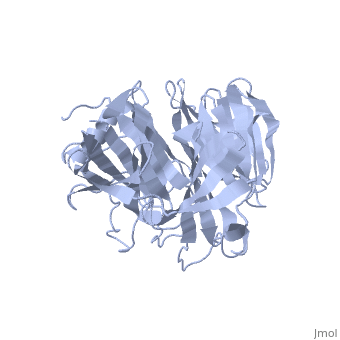
| | ==Crystal Structure of the N-terminal Domain of Nup159== | | ==Crystal Structure of the N-terminal Domain of Nup159== |
| - | <StructureSection load='1xip' size='340' side='right' caption='[[1xip]], [[Resolution|resolution]] 2.50Å' scene=''> | + | <StructureSection load='1xip' size='340' side='right'caption='[[1xip]], [[Resolution|resolution]] 2.50Å' scene=''> |
| | == Structural highlights == | | == Structural highlights == |
| - | <table><tr><td colspan='2'>[[1xip]] is a 1 chain structure with sequence from [http://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1XIP OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1XIP FirstGlance]. <br> | + | <table><tr><td colspan='2'>[[1xip]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Saccharomyces_cerevisiae Saccharomyces cerevisiae]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1XIP OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1XIP FirstGlance]. <br> |
| - | </td></tr><tr id='NonStdRes'><td class="sblockLbl"><b>[[Non-Standard_Residue|NonStd Res:]]</b></td><td class="sblockDat"><scene name='pdbligand=MSE:SELENOMETHIONINE'>MSE</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.5Å</td></tr> |
| - | <tr id='gene'><td class="sblockLbl"><b>[[Gene|Gene:]]</b></td><td class="sblockDat">NUP159 ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=4932 Saccharomyces cerevisiae])</td></tr> | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=MSE:SELENOMETHIONINE'>MSE</scene></td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=1xip FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1xip OCA], [http://www.rcsb.org/pdb/explore.do?structureId=1xip RCSB], [http://www.ebi.ac.uk/pdbsum/1xip PDBsum]</span></td></tr> | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1xip FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1xip OCA], [https://pdbe.org/1xip PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1xip RCSB], [https://www.ebi.ac.uk/pdbsum/1xip PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1xip ProSAT]</span></td></tr> |
| | </table> | | </table> |
| | == Function == | | == Function == |
| - | [[http://www.uniprot.org/uniprot/NU159_YEAST NU159_YEAST]] Functions as a component of the nuclear pore complex (NPC). NPC components, collectively referred to as nucleoporins (NUPs), can play the role of both NPC structural components and of docking or interaction partners for transiently associated nuclear transport factors. Active directional transport is assured by both, a Phe-Gly (FG) repeat affinity gradient for these transport factors across the NPC and a transport cofactor concentration gradient across the nuclear envelope (GSP1 and GSP2 GTPases associated predominantly with GTP in the nucleus, with GDP in the cytoplasm). NUP159 plays an important role in several nuclear export pathways including poly(A)+ RNA, pre-ribosome, and protein export.<ref>PMID:9736720</ref> <ref>PMID:9843582</ref> <ref>PMID:9891088</ref> <ref>PMID:10523319</ref> <ref>PMID:10801828</ref> <ref>PMID:10952996</ref> <ref>PMID:11387327</ref> <ref>PMID:11739405</ref> <ref>PMID:11689687</ref> <ref>PMID:12543930</ref> <ref>PMID:12604785</ref> <ref>PMID:15039779</ref> <ref>PMID:15574330</ref> | + | [https://www.uniprot.org/uniprot/NU159_YEAST NU159_YEAST] Functions as a component of the nuclear pore complex (NPC). NPC components, collectively referred to as nucleoporins (NUPs), can play the role of both NPC structural components and of docking or interaction partners for transiently associated nuclear transport factors. Active directional transport is assured by both, a Phe-Gly (FG) repeat affinity gradient for these transport factors across the NPC and a transport cofactor concentration gradient across the nuclear envelope (GSP1 and GSP2 GTPases associated predominantly with GTP in the nucleus, with GDP in the cytoplasm). NUP159 plays an important role in several nuclear export pathways including poly(A)+ RNA, pre-ribosome, and protein export.<ref>PMID:9736720</ref> <ref>PMID:9843582</ref> <ref>PMID:9891088</ref> <ref>PMID:10523319</ref> <ref>PMID:10801828</ref> <ref>PMID:10952996</ref> <ref>PMID:11387327</ref> <ref>PMID:11739405</ref> <ref>PMID:11689687</ref> <ref>PMID:12543930</ref> <ref>PMID:12604785</ref> <ref>PMID:15039779</ref> <ref>PMID:15574330</ref> |
| | == Evolutionary Conservation == | | == Evolutionary Conservation == |
| | [[Image:Consurf_key_small.gif|200px|right]] | | [[Image:Consurf_key_small.gif|200px|right]] |
| | Check<jmol> | | Check<jmol> |
| | <jmolCheckbox> | | <jmolCheckbox> |
| - | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/xi/1xip_consurf.spt"</scriptWhenChecked> | + | <scriptWhenChecked>; select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/xi/1xip_consurf.spt"</scriptWhenChecked> |
| | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> |
| | <text>to colour the structure by Evolutionary Conservation</text> | | <text>to colour the structure by Evolutionary Conservation</text> |
| | </jmolCheckbox> | | </jmolCheckbox> |
| - | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1xip ConSurf]. |
| | <div style="clear:both"></div> | | <div style="clear:both"></div> |
| | <div style="background-color:#fffaf0;"> | | <div style="background-color:#fffaf0;"> |
| Line 27: |
Line 28: |
| | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> |
| | </div> | | </div> |
| | + | <div class="pdbe-citations 1xip" style="background-color:#fffaf0;"></div> |
| | | | |
| | ==See Also== | | ==See Also== |
| | *[[Nucleoporin|Nucleoporin]] | | *[[Nucleoporin|Nucleoporin]] |
| | + | *[[Nucleoporin 3D structures|Nucleoporin 3D structures]] |
| | == References == | | == References == |
| | <references/> | | <references/> |
| | __TOC__ | | __TOC__ |
| | </StructureSection> | | </StructureSection> |
| | + | [[Category: Large Structures]] |
| | [[Category: Saccharomyces cerevisiae]] | | [[Category: Saccharomyces cerevisiae]] |
| - | [[Category: Berger, J M]] | + | [[Category: Berger JM]] |
| - | [[Category: Erzberger, J P]] | + | [[Category: Erzberger JP]] |
| - | [[Category: Weirich, C S]] | + | [[Category: Weirich CS]] |
| - | [[Category: Weis, K]] | + | [[Category: Weis K]] |
| - | [[Category: Beta-propeller]]
| + | |
| - | [[Category: Transport protein]]
| + | |
| Structural highlights
Function
NU159_YEAST Functions as a component of the nuclear pore complex (NPC). NPC components, collectively referred to as nucleoporins (NUPs), can play the role of both NPC structural components and of docking or interaction partners for transiently associated nuclear transport factors. Active directional transport is assured by both, a Phe-Gly (FG) repeat affinity gradient for these transport factors across the NPC and a transport cofactor concentration gradient across the nuclear envelope (GSP1 and GSP2 GTPases associated predominantly with GTP in the nucleus, with GDP in the cytoplasm). NUP159 plays an important role in several nuclear export pathways including poly(A)+ RNA, pre-ribosome, and protein export.[1] [2] [3] [4] [5] [6] [7] [8] [9] [10] [11] [12] [13]
Evolutionary Conservation
Check, as determined by ConSurfDB. You may read the explanation of the method and the full data available from ConSurf.
Publication Abstract from PubMed
Nuclear export of mRNA in eukaryotic cells is mediated by soluble transport factors and components of the nuclear pore complex (NPC). The cytoplasmically oriented nuclear pore protein Nup159 plays a critical role in mRNA export through its conserved N-terminal domain (NTD). Here, we report the crystal structure of the Nup159 NTD, refined to 2.5 A. The structure reveals an unusually asymmetric seven-bladed beta-propeller that is structurally conserved throughout eukarya. Using structure-based conservation analysis, we have targeted specific surface residues for mutagenesis. Residue substitutions in a conserved loop of the NTD abolish in vitro binding to Dbp5, a DEAD box helicase required for mRNA export. In vivo, these mutations cause Dbp5 mislocalization and block mRNA export. These findings suggest that the Nup159 NTD functions in mRNA export as a binding platform, tethering shuttling Dbp5 molecules at the nuclear periphery and locally concentrating this mRNA remodeling factor at the cytoplasmic face of the NPC.
The N-terminal domain of Nup159 forms a beta-propeller that functions in mRNA export by tethering the helicase Dbp5 to the nuclear pore.,Weirich CS, Erzberger JP, Berger JM, Weis K Mol Cell. 2004 Dec 3;16(5):749-60. PMID:15574330[14]
From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.
See Also
References
- ↑ Hurwitz ME, Strambio-de-Castillia C, Blobel G. Two yeast nuclear pore complex proteins involved in mRNA export form a cytoplasmically oriented subcomplex. Proc Natl Acad Sci U S A. 1998 Sep 15;95(19):11241-5. PMID:9736720
- ↑ Belgareh N, Snay-Hodge C, Pasteau F, Dagher S, Cole CN, Doye V. Functional characterization of a Nup159p-containing nuclear pore subcomplex. Mol Biol Cell. 1998 Dec;9(12):3475-92. PMID:9843582
- ↑ Seedorf M, Damelin M, Kahana J, Taura T, Silver PA. Interactions between a nuclear transporter and a subset of nuclear pore complex proteins depend on Ran GTPase. Mol Cell Biol. 1999 Feb;19(2):1547-57. PMID:9891088
- ↑ Hodge CA, Colot HV, Stafford P, Cole CN. Rat8p/Dbp5p is a shuttling transport factor that interacts with Rat7p/Nup159p and Gle1p and suppresses the mRNA export defect of xpo1-1 cells. EMBO J. 1999 Oct 15;18(20):5778-88. PMID:10523319 doi:10.1093/emboj/18.20.5778
- ↑ Bailer SM, Balduf C, Katahira J, Podtelejnikov A, Rollenhagen C, Mann M, Pante N, Hurt E. Nup116p associates with the Nup82p-Nsp1p-Nup159p nucleoporin complex. J Biol Chem. 2000 Aug 4;275(31):23540-8. PMID:10801828 doi:http://dx.doi.org/10.1074/jbc.M001963200
- ↑ Strasser K, Bassler J, Hurt E. Binding of the Mex67p/Mtr2p heterodimer to FXFG, GLFG, and FG repeat nucleoporins is essential for nuclear mRNA export. J Cell Biol. 2000 Aug 21;150(4):695-706. PMID:10952996
- ↑ Allen NP, Huang L, Burlingame A, Rexach M. Proteomic analysis of nucleoporin interacting proteins. J Biol Chem. 2001 Aug 3;276(31):29268-74. Epub 2001 May 31. PMID:11387327 doi:http://dx.doi.org/10.1074/jbc.M102629200
- ↑ Gleizes PE, Noaillac-Depeyre J, Leger-Silvestre I, Teulieres F, Dauxois JY, Pommet D, Azum-Gelade MC, Gas N. Ultrastructural localization of rRNA shows defective nuclear export of preribosomes in mutants of the Nup82p complex. J Cell Biol. 2001 Dec 10;155(6):923-36. Epub 2001 Dec 10. PMID:11739405 doi:http://dx.doi.org/10.1083/jcb.200108142
- ↑ Bailer SM, Balduf C, Hurt E. The Nsp1p carboxy-terminal domain is organized into functionally distinct coiled-coil regions required for assembly of nucleoporin subcomplexes and nucleocytoplasmic transport. Mol Cell Biol. 2001 Dec;21(23):7944-55. PMID:11689687 doi:http://dx.doi.org/10.1128/MCB.21.23.7944-7955.2001
- ↑ Allen NP, Patel SS, Huang L, Chalkley RJ, Burlingame A, Lutzmann M, Hurt EC, Rexach M. Deciphering networks of protein interactions at the nuclear pore complex. Mol Cell Proteomics. 2002 Dec;1(12):930-46. PMID:12543930
- ↑ Denning DP, Patel SS, Uversky V, Fink AL, Rexach M. Disorder in the nuclear pore complex: the FG repeat regions of nucleoporins are natively unfolded. Proc Natl Acad Sci U S A. 2003 Mar 4;100(5):2450-5. Epub 2003 Feb 25. PMID:12604785 doi:10.1073/pnas.0437902100
- ↑ Strawn LA, Shen T, Shulga N, Goldfarb DS, Wente SR. Minimal nuclear pore complexes define FG repeat domains essential for transport. Nat Cell Biol. 2004 Mar;6(3):197-206. Epub 2004 Feb 22. PMID:15039779 doi:10.1038/ncb1097
- ↑ Weirich CS, Erzberger JP, Berger JM, Weis K. The N-terminal domain of Nup159 forms a beta-propeller that functions in mRNA export by tethering the helicase Dbp5 to the nuclear pore. Mol Cell. 2004 Dec 3;16(5):749-60. PMID:15574330 doi:10.1016/j.molcel.2004.10.032
- ↑ Weirich CS, Erzberger JP, Berger JM, Weis K. The N-terminal domain of Nup159 forms a beta-propeller that functions in mRNA export by tethering the helicase Dbp5 to the nuclear pore. Mol Cell. 2004 Dec 3;16(5):749-60. PMID:15574330 doi:10.1016/j.molcel.2004.10.032
|