This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Diguanylate cyclase
From Proteopedia
(Difference between revisions)
(New page: <StructureSection load='1stp' size='340' side='right' caption='Caption for this structure' scene=''> == Function == '''Diguanylate cyclase''' (DGC) catalyzes the conversion of GTP to c...) |
|||
| (21 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
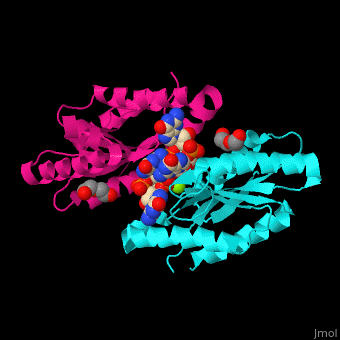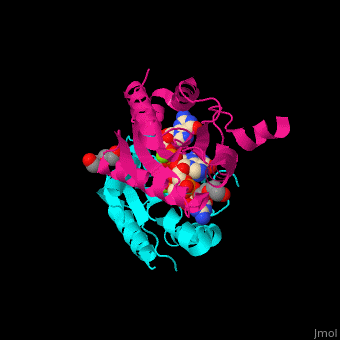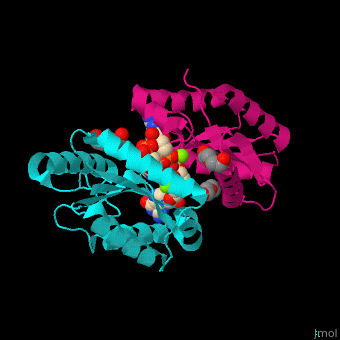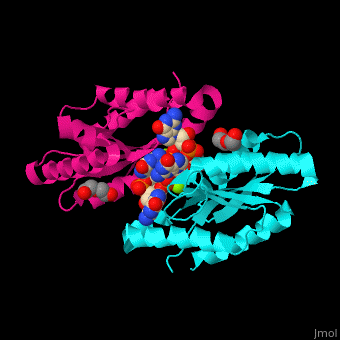
| - | + | <StructureSection load='' size='350' side='right' caption='Diguanylate cyclase GGDEF domain complex with cyclic di-GMP, Mg+2 ion (green) and PEG (PDB code [[3qyy]]).' scene='67/677040/Cv/1'> | |
| - | <StructureSection load=' | + | |
== Function == | == Function == | ||
| - | '''Diguanylate cyclase''' (DGC) catalyzes the conversion of GTP to | + | '''Diguanylate cyclase''' (DGC) catalyzes the conversion of GTP to [[C-di-GMP]] and diphosphate. <ref>PMID:17697992</ref> For more details see<br /> |
| - | + | [[PleD catalysis]]<br /> | |
| - | + | [[PleD activation]]<br /> | |
| + | [[PleD allosteric product inhibition]]<br /> | ||
| + | [[C-di-GMP signaling]]. | ||
== Relevance == | == Relevance == | ||
| + | |||
| + | DGC from ''Borrelia burgdorferi'' is involved in the transmission of spirochetes from ticks to mammals causing Lyme Disease. | ||
== Structural highlights == | == Structural highlights == | ||
| + | |||
| + | DGC contains the sequence <scene name='67/677040/Cv/4'>motif GGDEF</scene> in its active site. <scene name='67/677040/Cv/5'>Cyclic di-GMP and Mg+2 ions interact with two monomers</scene> of DGC.<ref>PMID:22120736</ref> | ||
</StructureSection> | </StructureSection> | ||
| Line 21: | Line 26: | ||
*DGC | *DGC | ||
| - | **[[4urg]], [[ | + | **[[4urg]], [[6khu]] – TmDGC GGDEF domain 82-248 – ''Thermotoga maritima''<br /> |
**[[4urq]] – TmDGC GGDEF domain (mutant)<br /> | **[[4urq]] – TmDGC GGDEF domain (mutant)<br /> | ||
| - | **[3ign]] – | + | **[[3ign]] – MaDGC GGDEF domain – ''Marinobacter aquaeolei''<br /> |
| - | **[[ | + | **[[3h9w]] – MaDGC N terminal<br /> |
| + | **[[4zmu]] – PaDGC – ''Pseudomonas aeruginosa''<br /> | ||
| + | **[[3hvw]], [[4iob]] – DGC GGDEF domain 254-414<br /> | ||
| + | **[[7a7e]] – PaDGC GGDEF domain (mutant)<br /> | ||
| + | **[[4zmm]] – PaDGC GGDEF domain + C-diGMP<br /> | ||
| + | **[[5m3c]] – PaDGC GGDEF domain + GTP<br /> | ||
| + | **[[7e6g]] – PaDGC GGDEF domain + SiaC<br /> | ||
| + | **[[3n53]] – DGC – ''Pelobacter carbinolicus''<br /> | ||
| + | **[[5h5o]] – XcDGC residues 1-150 – ''Xanthomonas campestris''<br /> | ||
| + | **[[3qyy]] – XcDGC GGDEF domain + cyclic di-GMP <br /> | ||
| + | **[[4urs]] – TmDGC GGDEF domain + cyclic di-GMP<br /> | ||
| + | **[[3t9o]] – EcDGC YDEH CZB domain – ''Escherichia coli''<br /> | ||
| + | **[[3tvk]], [[4zve]], [[4zvg]], [[4zvh]] – EcDGC YDEH GGDEF domain <br /> | ||
| + | **[[4h54]] – EcDGC YDEH (mutant) <br /> | ||
| + | **[[4zva]], [[4zvb]] – EcDGC globin domain <br /> | ||
| + | **[[4zvc]], [[4zvd]] – EcDGC mid domain <br /> | ||
| + | **[[4zvf]] – EcDGC GGDEF domain 297-460 + GTP derivative<br /> | ||
| + | **[[6eib]] – DGC GGDEF domain – ''Vibrio cholerae''<br /> | ||
| + | **[[6et7]] – IdDGC (mutant) - ''Idiomarina''<br /> | ||
| + | **[[5llw]], [[5lly]] – IdDGC + phytochromobilin <br /> | ||
| + | **[[5llx]] – IdDGC + phytochromobilin + GTP <br /> | ||
| + | **[[6saw]], [[6sax]] – IdDGC chromophore-binding domain 3-312<br /> | ||
| + | **[[6hbz]] – DGCB – ''Bdellovibrio bacteriovorus''<br /> | ||
| + | **[[6tts]] – CvDGCB GGDEF domain 176-353 + C-diGMP – ''Caulobacter vibrioides'' <br /> | ||
| + | **[[6ttr]] – CvDGCB GGDEF+coiled-coil domains 153-353 (mutant) + C-diGMP <br /> | ||
| + | **[[7k5n]] – DGC LBD domain 39-286 + Pro – ''Aeromonas caviae''<br /> | ||
*Response regulator PleD | *Response regulator PleD | ||
| - | **[[1w25]], [[2wb4]] – CvDGC + | + | **[[1w25]], [[2wb4]] – CvDGC + cyclic di-GMP <br /> |
| - | **[[2v0n]] – CvDGC + | + | **[[2v0n]] – CvDGC + cyclic di-GMP + GTP<br /> |
}} | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D Structures of diguanylate cyclase
Updated on 21-December-2022
References
- ↑ Stock AM. Diguanylate cyclase activation: it takes two. Structure. 2007 Aug;15(8):887-8. PMID:17697992 doi:http://dx.doi.org/10.1016/j.str.2007.07.003
- ↑ Yang CY, Chin KH, Chuah ML, Liang ZX, Wang AH, Chou SH. The structure and inhibition of a GGDEF diguanylate cyclase complexed with (c-di-GMP)(2) at the active site. Acta Crystallogr D Biol Crystallogr. 2011 Dec;67(Pt 12):997-1008. Epub 2011 Nov, 18. PMID:22120736 doi:10.1107/S090744491104039X