We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
Sandbox Reserved 951
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 1: | Line 1: | ||
<StructureSection load='1LCI' size='450' side='right' caption='Crystal structure of firefly luciferase' scene='Insert optional scene name here'> | <StructureSection load='1LCI' size='450' side='right' caption='Crystal structure of firefly luciferase' scene='Insert optional scene name here'> | ||
| + | == <div class="center" style="width: auto; margin-left: auto; margin-right: auto;">Firefly Luciferase</div> == | ||
==Biological context== | ==Biological context== | ||
| Line 32: | Line 33: | ||
====Interactions with ligands==== | ====Interactions with ligands==== | ||
| - | The active site is not strictly highlighted according to the actual state of studies but some residues and motifs strongly modified have been determined and this conformation enables to find the active site. Many of these conserved residues are located on the core of the β-barrel, on the small C-terminal domain and in the surface of the N-terminal domain, which forms a <scene name='60/604470/Depression_c-ter_and_n-ter/1'>depression</scene>. However, this depression is too large to enable interactions between residues and substrates, so it is thought that a conformational change occurs and sandwiches the substrates, forming the active site. This conformational change provides a suitable environment for light production because of exclusion of water molecules from the active site, favouring intramolecular reactions. Residues also follow a <scene name='60/604470/Cleft/1'>cleft</scene> caused by of the <scene name='60/604470/Beta_sheet_b/3'>Beta-sheet B</scene> against the <scene name='60/604470/Beta_barrel/3'>Beta-barrel</scene>.<ref name = | + | The active site is not strictly highlighted according to the actual state of studies but some residues and motifs strongly modified have been determined and this conformation enables to find the active site. Many of these conserved residues are located on the core of the β-barrel, on the small C-terminal domain and in the surface of the N-terminal domain, which forms a <scene name='60/604470/Depression_c-ter_and_n-ter/1'>depression</scene>. However, this depression is too large to enable interactions between residues and substrates, so it is thought that a conformational change occurs and sandwiches the substrates, forming the active site. This conformational change provides a suitable environment for light production because of exclusion of water molecules from the active site, favouring intramolecular reactions. Residues also follow a <scene name='60/604470/Cleft/1'>cleft</scene> caused by of the <scene name='60/604470/Beta_sheet_b/3'>Beta-sheet B</scene> against the <scene name='60/604470/Beta_barrel/3'>Beta-barrel</scene>.<ref name ="first">PMID:8805533</ref> |
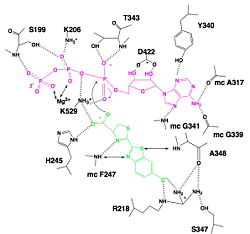
[[Image:Luciferin_bounding_to_luciferase.jpg|250px|right|thumb|Hydrogen bonding between Luciferase and substrates luciferin (green), ATP (violet) and Mg2+,<ref name="second">[http://www.photobiology.info/ Photobiology]</ref>]] | [[Image:Luciferin_bounding_to_luciferase.jpg|250px|right|thumb|Hydrogen bonding between Luciferase and substrates luciferin (green), ATP (violet) and Mg2+,<ref name="second">[http://www.photobiology.info/ Photobiology]</ref>]] | ||
Current revision
| |||||||||||
References
- ↑ Welsh DK, Kay SA. Bioluminescence imaging in living organisms. Curr Opin Biotechnol. 2005 Feb;16(1):73-8. PMID:15722018 doi:http://dx.doi.org/10.1016/j.copbio.2004.12.006
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 2.6 Conti E, Franks NP, Brick P. Crystal structure of firefly luciferase throws light on a superfamily of adenylate-forming enzymes. Structure. 1996 Mar 15;4(3):287-98. PMID:8805533
- ↑ 3.0 3.1 Marques SM, Esteves da Silva JC. Firefly bioluminescence: a mechanistic approach of luciferase catalyzed reactions. IUBMB Life. 2009 Jan;61(1):6-17. PMID:18949818 doi:10.1002/iub.134
- ↑ 4.0 4.1 4.2 Photobiology
- ↑ Hosseinkhani S. Molecular enigma of multicolor bioluminescence of firefly luciferase. Cell Mol Life Sci. 2011 Apr;68(7):1167-82. doi: 10.1007/s00018-010-0607-0. Epub, 2010 Dec 28. PMID:21188462 doi:http://dx.doi.org/10.1007/s00018-010-0607-0
- ↑ Inouye S. Firefly luciferase: an adenylate-forming enzyme for multicatalytic functions. Cell Mol Life Sci. 2010 Feb;67(3):387-404. Epub 2009 Oct 27. PMID:19859663 doi:10.1007/s00018-009-0170-8