This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
JMS/Sandbox25
From Proteopedia
(Difference between revisions)
(New page: ==Your Heading Here (maybe something like 'Structure')== <StructureSection load='Conevo_1.pdb' size='340' side='right' caption='Caption for this structure' scene=''> This is a default text...) |
|||
| (17 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
==Your Heading Here (maybe something like 'Structure')== | ==Your Heading Here (maybe something like 'Structure')== | ||
| - | <StructureSection load='Conevo_1.pdb' size='340' side='right' caption='Caption for this structure' scene=''> | + | <StructureSection load='Conevo_1.pdb' size='340' side='right' caption='Caption for this structure' scene='70/700369/Golden_comp_1/1'> |
| - | + | ||
You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | You may include any references to papers as in: the use of JSmol in Proteopedia <ref>DOI 10.1002/ijch.201300024</ref> or to the article describing Jmol <ref>PMID:21638687</ref> to the rescue. | ||
| - | == | + | == Structural highlights == |
| - | == | + | key: w = whale; p = pig; s = seal; c = cat; m = muskrate; g = guniae pig. |
| - | = | + | <scene name='70/700369/Golden_comp_1/2'>all areas with possible change in net charge</scene> 8,12,31,53,57,82,84,87,113,116,121,122,132,140. |
| - | == | + | <scene name='70/700369/Golden_comp_1/3'>only times created a hydrophobic patch</scene> was w53p, c121s and g122m. in every other case removing a hydrophobic patch in each case! |
| + | <scene name='70/700369/Golden_comp_1/4'>funny at 84</scene>, muskrat lost a hydrophic patch but becomes less charged. | ||
| + | <scene name='70/700369/Golden_comp_1/6'>could have increased the charge AND the net charge at once ... but didn't!</scene> in blue. in red, is a conserved negatively charge aa - could been both or just increase in charge. | ||
| - | + | <scene name='70/700369/Golden_comp_1/7'>showing lost hydrophobic patch, increase in charge of muskrat 84 in purple, and the three increase in hydophobic patches in green</scene> | |
| + | p:w 12,53,87,113,116 | ||
| + | c:s 8,31,57,82,116,121 | ||
| + | g:m 12,82,84,113,122,132 | ||
| + | |||
| + | wm 12,113 | ||
| + | sm 82 | ||
| + | |||
| + | <scene name='70/700369/Golden_comp_1/9'>showing where each evolution step occurred by aquatic mammal.</scene> | ||
| + | |||
| + | <scene name='70/700369/Globin/1'>dude!!</scene>a backbone of 3 myoglobin pairs, oxy and deoxy wt hemoglobin, and deoxy sickle cell, and the horse myoglobin dimer. | ||
| + | |||
| + | showing 1mbn with the sites that have evolved. and hbB from sickle, with the e6v in halo. <scene name='70/700369/Globin/2'>funny no i6+ in aquatic mammals.</scene> | ||
| + | |||
| + | w whale -> r c cat -> q s seal -> N g gunea pig -> o p pig m muskrat | ||
| + | z oaa oxy y oab | ||
| + | x doa deoxy w dob v doc u dod | ||
| + | a-h ds{a-h} sickle cell | ||
| + | t horse a s horse b | ||
| + | |||
| + | <scene name='70/700369/Globin/5'>whale horse comparison</scene> | ||
| + | |||
| + | <scene name='70/700369/Globin/6'>whale to horse comp. emphasizing halves of protein</scene>. next up is interface. | ||
| + | |||
| + | note the v to i mutation in all three pairs! why?? also, horse is i.. | ||
</StructureSection> | </StructureSection> | ||
| + | |||
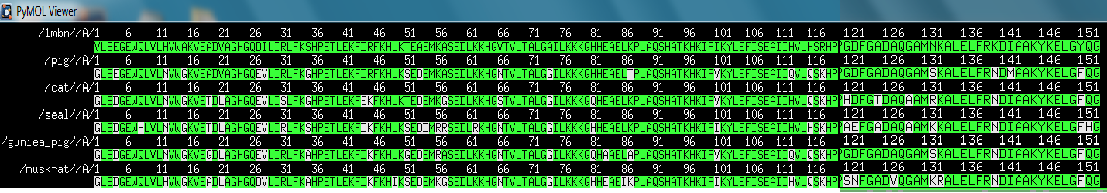
| + | pymol sequence alignment | ||
| + | [[Image:Comp_seq_1.png]] | ||
== References == | == References == | ||
<references/> | <references/> | ||
Current revision
Your Heading Here (maybe something like 'Structure')
| |||||||||||
References
- ↑ Hanson, R. M., Prilusky, J., Renjian, Z., Nakane, T. and Sussman, J. L. (2013), JSmol and the Next-Generation Web-Based Representation of 3D Molecular Structure as Applied to Proteopedia. Isr. J. Chem., 53:207-216. doi:http://dx.doi.org/10.1002/ijch.201300024
- ↑ Herraez A. Biomolecules in the computer: Jmol to the rescue. Biochem Mol Biol Educ. 2006 Jul;34(4):255-61. doi: 10.1002/bmb.2006.494034042644. PMID:21638687 doi:10.1002/bmb.2006.494034042644