This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Connexin
From Proteopedia
(Difference between revisions)
| (8 intermediate revisions not shown.) | |||
| Line 35: | Line 35: | ||
| - | =Differences between wild type and mutant connexin 26 | + | ==Differences between wild type and mutant connexin 26== |
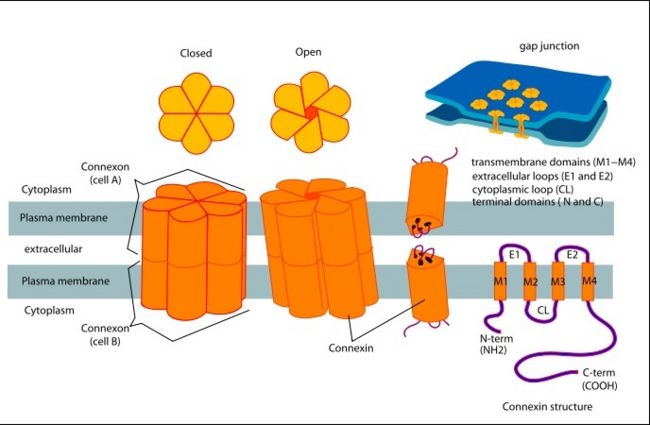
In general, single site mutations are spread fairly evenly across the whole protein with TM2 having the highest mutation density (number of amino acids with NHLS mutations divided by the total number of amino acids in the domain) at 67% to M1 and E1, having the lowest density of mutations with their respective domains at 33%. According to this criterion, TM4 has a mutation density of 40%. Of the four transmembrane helices, M1, M2 and M3 have attracted the most attention, because of the controversies involved in models with different helix assignments, based on lower resolution cryo-electron crystallographic structures and scanning cysteine accessibility mutagenesis. Far less is known about TM4 and how side chains interact with the other helices and with the lipid bilayer.<ref name='mutant int'/> | In general, single site mutations are spread fairly evenly across the whole protein with TM2 having the highest mutation density (number of amino acids with NHLS mutations divided by the total number of amino acids in the domain) at 67% to M1 and E1, having the lowest density of mutations with their respective domains at 33%. According to this criterion, TM4 has a mutation density of 40%. Of the four transmembrane helices, M1, M2 and M3 have attracted the most attention, because of the controversies involved in models with different helix assignments, based on lower resolution cryo-electron crystallographic structures and scanning cysteine accessibility mutagenesis. Far less is known about TM4 and how side chains interact with the other helices and with the lipid bilayer.<ref name='mutant int'/> | ||
| Line 43: | Line 43: | ||
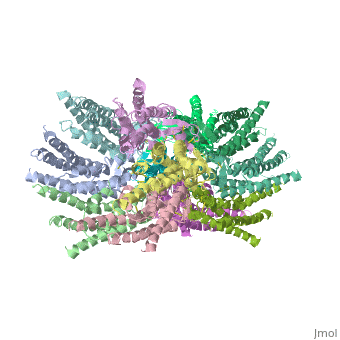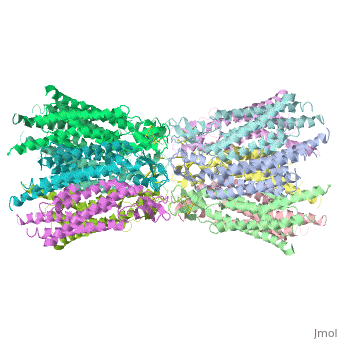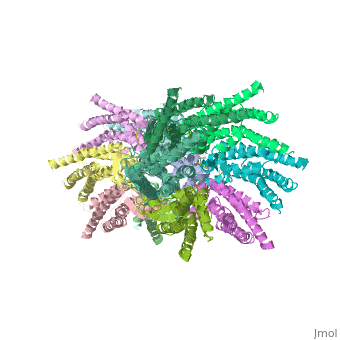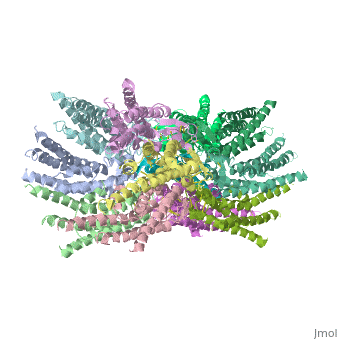
Functional analysis of the Cx26M34A channels revealed that these channels are predominantly closed, with the residual electrical conductance showing normal voltage gating. N-terminal deletion mutants with and without the M34A mutation showed no electrical activity in paired Xenopus oocytes and significantly decreased dye permeability in HeLa cells. Comparing this closed structure with the published X-ray structure of wild-type Cx26, which is proposed to be in an open state, revealed a radial outward shift in the transmembrane helices in the closed state, presumably to accommodate the N-terminal plug occluding the pore. Because both Cx26del2-7 and Cx26M34Adel2-7 channels are closed, the N terminus appears to have a prominent role in stabilizing the open configuration. <ref name='pdb'/> | Functional analysis of the Cx26M34A channels revealed that these channels are predominantly closed, with the residual electrical conductance showing normal voltage gating. N-terminal deletion mutants with and without the M34A mutation showed no electrical activity in paired Xenopus oocytes and significantly decreased dye permeability in HeLa cells. Comparing this closed structure with the published X-ray structure of wild-type Cx26, which is proposed to be in an open state, revealed a radial outward shift in the transmembrane helices in the closed state, presumably to accommodate the N-terminal plug occluding the pore. Because both Cx26del2-7 and Cx26M34Adel2-7 channels are closed, the N terminus appears to have a prominent role in stabilizing the open configuration. <ref name='pdb'/> | ||
| + | =3D structures of connexin= | ||
| + | [[Connexin 3D structure]] | ||
| + | </StructureSection> | ||
| - | + | = References = | |
<references/> | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Zonta F, Buratto D, Cassini C, Bortolozzi M, Mammano F. Molecular dynamics simulations highlight structural and functional alterations in deafness-related M34T mutation of connexin 26. Front Physiol. 2014 Mar 4;5:85. doi: 10.3389/fphys.2014.00085. eCollection 2014. PMID:24624091 doi:http://dx.doi.org/10.3389/fphys.2014.00085
- ↑ 2.0 2.1 2.2 2.3 Suga M, Maeda S, Nakagawa S, Yamashita E, Tsukihara T. A description of the structural determination procedures of a gap junction channel at 3.5 A resolution. Acta Crystallogr D Biol Crystallogr. 2009 Aug;65(Pt 8):758-66. Epub 2009, Jul 10. PMID:19622859 doi:http://dx.doi.org/10.1107/S0907444909014711
- ↑ http://en.wikipedia.org/wiki/Connexin
- ↑ Sokolov M, Brownstein Z, Frydman M, Avraham KB. Apparent phenotypic anticipation in autosomal dominant connexin 26 deafness. J Basic Clin Physiol Pharmacol. 2014 Sep;25(3):289-92. doi:, 10.1515/jbcpp-2014-0053. PMID:25153233 doi:http://dx.doi.org/10.1515/jbcpp-2014-0053
- ↑ 5.0 5.1 Ambrosi C, Walker AE, Depriest AD, Cone AC, Lu C, Badger J, Skerrett IM, Sosinsky GE. Analysis of trafficking, stability and function of human connexin 26 gap junction channels with deafness-causing mutations in the fourth transmembrane helix. PLoS One. 2013 Aug 15;8(8):e70916. doi: 10.1371/journal.pone.0070916. eCollection, 2013. PMID:23967136 doi:http://dx.doi.org/10.1371/journal.pone.0070916
- ↑ Kikuchi T, Adams JC, Miyabe Y, So E, Kobayashi T. Potassium ion recycling pathway via gap junction systems in the mammalian cochlea and its interruption in hereditary nonsyndromic deafness. Med Electron Microsc. 2000;33(2):51-6. PMID:11810458 doi:http://dx.doi.org/10.1007/s007950000009
- ↑ 7.0 7.1 7.2 7.3 7.4 Oshima A, Tani K, Toloue MM, Hiroaki Y, Smock A, Inukai S, Cone A, Nicholson BJ, Sosinsky GE, Fujiyoshi Y. Asymmetric Configurations and N-terminal Rearrangements in Connexin26 Gap Junction Channels. J Mol Biol. 2011 Jan 21;405(3):724-35. Epub 2010 Nov 20. PMID:21094651 doi:10.1016/j.jmb.2010.10.032
Proteopedia Page Contributors and Editors (what is this?)
Safaa Salah Hussiesy, Michal Harel, Doaa Naffaa, Jaime Prilusky