This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Ectonucleoside triphosphate diphosphohydrolase
From Proteopedia
(Difference between revisions)
| (2 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
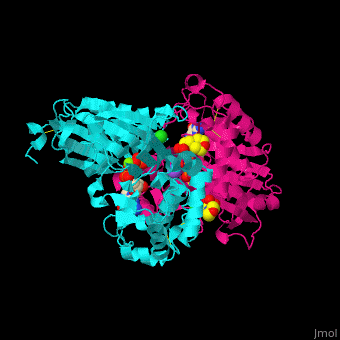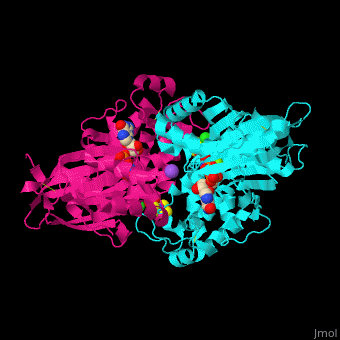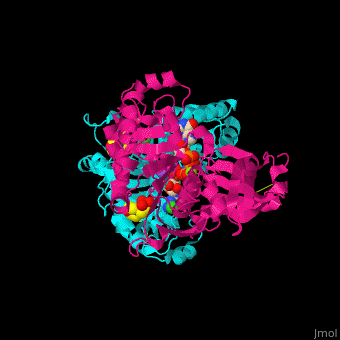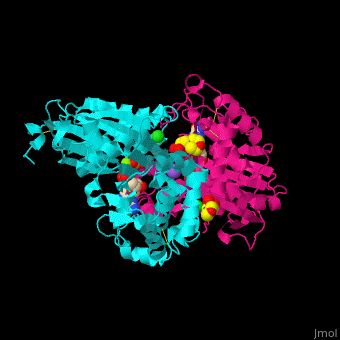
| - | <StructureSection load='4brg' size=' | + | <StructureSection load='4brg' size='400' side='right' caption='Structure of NTDPase1 complex with GMPPNP, Na+ (purple), Cl- (large green), Mg+2 (small green) ions (PDB code [[4brg]]).' scene='59/597004/Cv/1'> |
| - | == Function == | ||
'''Ectonucleoside triphosphate diphosphohydrolase (NTPDase)''' hydrolyzes nucleoside 5’-triphosphate to nucleoside 5’-phosphate and 2 phosphates. NTPDase can hydrolyze ATP and ADP. NTPDase has an important role in regulating neutrophil chemotaxis. NTPDase 1,2,3,8 are cell surface enzymes with catalytic sites facing extracellularly while other NTPDases act intracellularly. NTPDase require Ca<sup>+2</sup> or Mg<sup>+2</sup> for their activity. The NTPDases can be differentiated according to their substrate specificity, divalent cation usage and product formation.<ref>PMID:23830739</ref> | '''Ectonucleoside triphosphate diphosphohydrolase (NTPDase)''' hydrolyzes nucleoside 5’-triphosphate to nucleoside 5’-phosphate and 2 phosphates. NTPDase can hydrolyze ATP and ADP. NTPDase has an important role in regulating neutrophil chemotaxis. NTPDase 1,2,3,8 are cell surface enzymes with catalytic sites facing extracellularly while other NTPDases act intracellularly. NTPDase require Ca<sup>+2</sup> or Mg<sup>+2</sup> for their activity. The NTPDases can be differentiated according to their substrate specificity, divalent cation usage and product formation.<ref>PMID:23830739</ref> | ||
| - | == Disease == | ||
| - | |||
| - | == Relevance == | ||
| - | |||
| - | == Structural highlights == | ||
</StructureSection> | </StructureSection> | ||
| Line 71: | Line 65: | ||
**[[4a5a]], [[4kh4]] – TgNTPDase (mutant) + AMPPNP<br /> | **[[4a5a]], [[4kh4]] – TgNTPDase (mutant) + AMPPNP<br /> | ||
**[[4kh5]] – TgNTPDase (mutant) + adenosine-imido-diphosphate<br /> | **[[4kh5]] – TgNTPDase (mutant) + adenosine-imido-diphosphate<br /> | ||
| + | |||
| + | *NTPDase4 | ||
| + | |||
| + | **[[6wg5]] – hNTPDase <br /> | ||
}} | }} | ||
Current revision
| |||||||||||
3D structures of ectonucleoside triphosphate diphosphohydrolase
Updated on 12-July-2021
References
- ↑ Zebisch M, Krauss M, Schafer P, Lauble P, Strater N. Crystallographic Snapshots along the Reaction Pathway of Nucleoside Triphosphate Diphosphohydrolases. Structure. 2013 Jul 2. pii: S0969-2126(13)00200-1. doi:, 10.1016/j.str.2013.05.016. PMID:23830739 doi:10.1016/j.str.2013.05.016