NADH peroxidase
From Proteopedia
(Difference between revisions)
| (7 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | + | <StructureSection load='' size='450' side='right' scene='48/486476/Cv/1' caption='NADH peroxidase containing FAD complex with NAD, [[2npx]]'> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
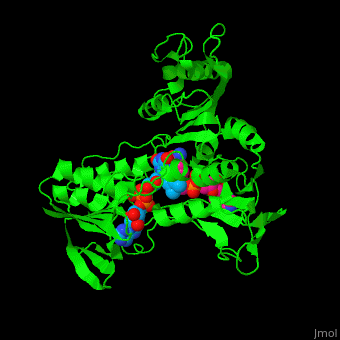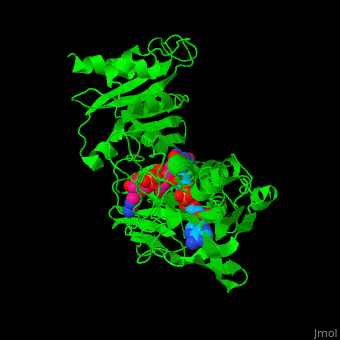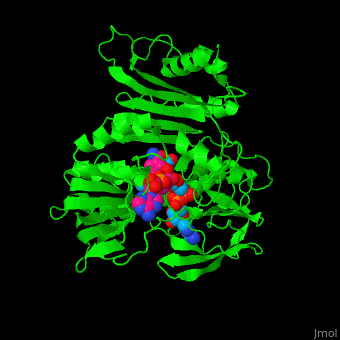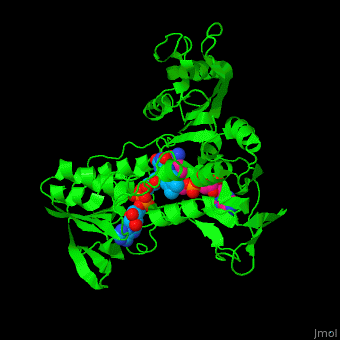
| + | '''NADH peroxidase''' (NPO) catalyzes the conversion of NADH to NAD+ using hydrogen peroxide. See [[NAD]]. NPO eliminates the potentially toxic hydrogen peroxide and defends the cell against H<sub>2</sub>O<sub>2</sub>-mediated oxidative stress<ref>PMID:8425532</ref>. NPO provides an additional pathway for regeneration of NAD+ which is essential to the fermentative metabolism. FAD is the cofactor in this reaction. | ||
| + | *<scene name='48/486476/Cv/13'>NAD binding site</scene>. Water molrcules are shown as red spheres. | ||
| + | *<scene name='48/486476/Cv/14'>NAD/FAD interactions</scene>. | ||
| + | *<scene name='48/486476/Cv/15'>FAD binding site</scene>. | ||
| + | *<scene name='48/486476/Cv/16'>Whole binding site</scene>. | ||
| + | *<scene name='48/486476/Cv/17'>NAD binding pocket</scene>. | ||
| + | *<scene name='48/486476/Cv/18'>FAD binding pocket</scene>. | ||
| + | </StructureSection> | ||
==3D structures of NADH peroxidase== | ==3D structures of NADH peroxidase== | ||
| Line 13: | Line 14: | ||
[[2npx]] – EfNPO + NAD<br /> | [[2npx]] – EfNPO + NAD<br /> | ||
[[1nhp]], [[1nhq]], [[1nhr]], [[1nhs]], [[1f8w]] – EfNPO (mutant) | [[1nhp]], [[1nhq]], [[1nhr]], [[1nhs]], [[1f8w]] – EfNPO (mutant) | ||
| + | |||
| + | == References == | ||
| + | <references/> | ||
| + | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
3D structures of NADH peroxidase
1npx, 1joa – EfNPO – Enterococcus faecalis
2npx – EfNPO + NAD
1nhp, 1nhq, 1nhr, 1nhs, 1f8w – EfNPO (mutant)
References
- ↑ Stehle T, Claiborne A, Schulz GE. NADH binding site and catalysis of NADH peroxidase. Eur J Biochem. 1993 Jan 15;211(1-2):221-6. PMID:8425532