We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
SAM-dependent methyltransferase
From Proteopedia
(Difference between revisions)
| (18 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
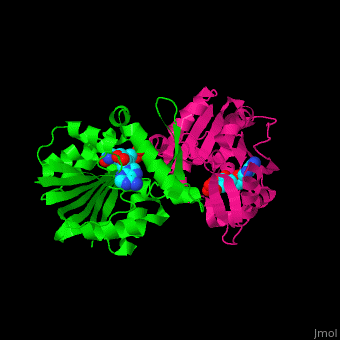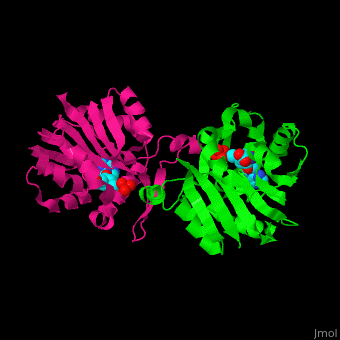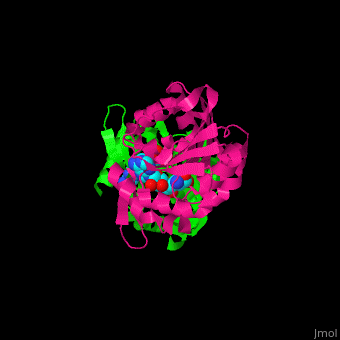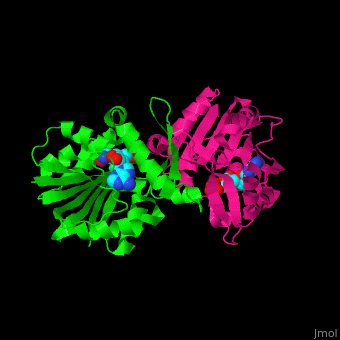
| - | + | <StructureSection load='' size='350' side='right' scene='51/510230/Cv/3' caption='SAM-dependent methyltransferase dimer complex with S-adenosyl-L-homocysteine and sulfate [[3ou6]]'> | |
== Function == | == Function == | ||
| - | '''SAM-dependent methyltransferase''' (SDM) utilizes the methyl donor S-adenosyl-L-methionine (SAM) as a cofactor to methylate proteins, small molecules, lipids and nucleic acids. SAM forms S-adenosyl-L-homocysteine (SAH) upon demethylation. About 120 members of the SDM family have been identified. They differ in their substrate specificity and the atom targeted for methylation (N, O, C, S)<ref>PMID:23180741</ref>. | + | '''SAM-dependent methyltransferase''' or '''S-adenosylmethionine-dependent methyltransferase''' (SDM) utilizes the methyl donor S-adenosyl-L-methionine (SAM) as a cofactor to methylate proteins, small molecules, lipids and nucleic acids. SAM forms S-adenosyl-L-homocysteine (SAH) upon demethylation. About 120 members of the SDM family have been identified. They differ in their substrate specificity and the atom targeted for methylation ('''N, O, C, S''')<ref>PMID:23180741</ref>. |
| + | For '''Chemotaxis receptor methyltransferase CheR''' see details in [[Molecular Playground/CheR]].<ref>PMID:9628482</ref>.<br /> | ||
| + | *SEE ALSO [[Chemotaxis protein]]. | ||
== Structural highlights == | == Structural highlights == | ||
| - | The core of the SDM fold contains alternating β strands and α helices. | + | The <scene name='51/510230/Cv/5'>core of the SDM fold contains alternating β strands and α helices</scene>. SDM <scene name='51/510230/Cv/6'>active site is located between the 2 monomers</scene><ref>PMID:20876132</ref>. Water molecules are shown as red spheres. |
==3D structures of SAM-dependent methyltrasferase== | ==3D structures of SAM-dependent methyltrasferase== | ||
| + | [[SAM-dependent methyltrasferase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | *SAM-dependent O-methyltransferase | ||
| - | |||
| - | **[[1o54]] – TmSDM – ''Thermotoga maritima''<br /> | ||
| - | **[[2hnk]] – SDM – ''Leptospira interrogans''<br /> | ||
| - | **[[3r3h]] – SDM – ''Legionella pneumophila'' | ||
| - | |||
| - | *SAM-dependent N-methyltransferase | ||
| - | |||
| - | **[[1vlm]] – TmSDM (mutant) | ||
| - | |||
| - | *SAM-dependent methyltransferase | ||
| - | |||
| - | **[[1wzn]] – SDM – ''Pyrococcus horikoshii''<br /> | ||
| - | **[[2igt]] – SDM –'' Agrobacterium tumefaciens''<br /> | ||
| - | **[[3cgg]] – CgSDM – ''Corynebacterium glutamicum''<br /> | ||
| - | **[[3d2l]] - SDM – ''Exiguobacterium sibiricum''<br /> | ||
| - | **[[3dh0]] - SDM – ''Aquifex aeolicus''<br /> | ||
| - | **[[3dp7]] - SDM – ''Bacterioides vulgatus''<br /> | ||
| - | **[[3ggd]] - SDM – ''Anabaena variabilis''<br /> | ||
| - | **[[3sm3]] - SDM (mutant) – ''Methanosarcina mazei''<br /> | ||
| - | **[[3bkx]] – SDM – ''Lactobacillus casei''<br /> | ||
| - | **[[3kr9]] – SpSDM – ''Streptococcus pneumonia''<br /> | ||
| - | |||
| - | *SAM-dependent methyltransferase complex with SAM/SAH | ||
| - | |||
| - | **[[3ku1]] – SpSDM + SAM<br /> | ||
| - | **[[3lga]] – PaSDM + SAH – ''Pyrococcus abyssi''<br /> | ||
| - | **[[3mb5]] – PaSDM + SAM<br /> | ||
| - | **[[3ou2]] – SlSDM + SAH – ''Streptomyces luridus''<br /> | ||
| - | **[[3ou6]] – SlSDM + SAM<br /> | ||
| - | **[[3ou7]] – SlSDM + SAM + phosphonic acid derivative<br /> | ||
| - | **[[3h2b]] – CgSDM + pyrophosphate + SAH<br /> | ||
| - | **[[5bp7]] – SlSDM + SAH – ''Geobacter sulfurreducens''<br /> | ||
| - | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Struck AW, Thompson ML, Wong LS, Micklefield J. S-adenosyl-methionine-dependent methyltransferases: highly versatile enzymes in biocatalysis, biosynthesis and other biotechnological applications. Chembiochem. 2012 Dec 21;13(18):2642-55. doi: 10.1002/cbic.201200556. Epub 2012, Nov 23. PMID:23180741 doi:http://dx.doi.org/10.1002/cbic.201200556
- ↑ Djordjevic S, Stock AM. Chemotaxis receptor recognition by protein methyltransferase CheR. Nat Struct Biol. 1998 Jun;5(6):446-50. PMID:9628482
- ↑ Lee JH, Bae B, Kuemin M, Circello BT, Metcalf WW, Nair SK, van der Donk WA. Characterization and structure of DhpI, a phosphonate O-methyltransferase involved in dehydrophos biosynthesis. Proc Natl Acad Sci U S A. 2010 Oct 12;107(41):17557-62. Epub 2010 Sep 27. PMID:20876132 doi:10.1073/pnas.1006848107