This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
TRNA methyltransferase
From Proteopedia
(Difference between revisions)
| (11 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
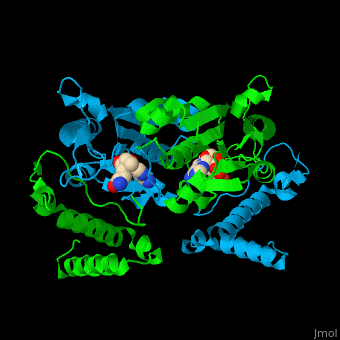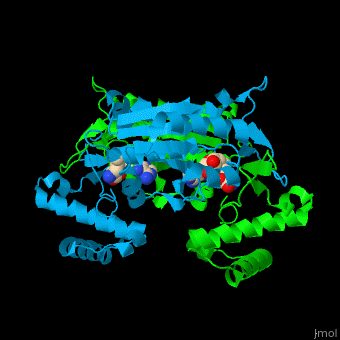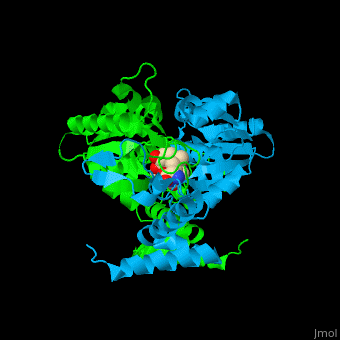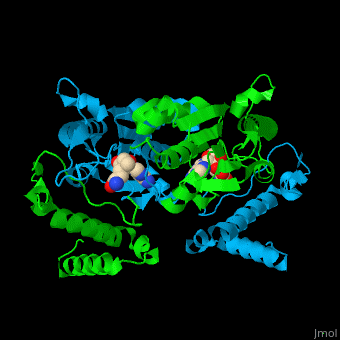
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' caption='Structure of tRNA methyltransferase for N1 of guanine 37 complex with S-adenosylhomocysteine (PDB entry [[1uak]])' scene='55/553109/Cv/1'> |
== Function == | == Function == | ||
'''tRNA methyltransferase''' (Trm) methylates specific positions on tRNA. tRNA methylation is a post-transcription modification. The methyl donor is S-adenosylmethionine (SAM) and the product is the methylated tRNA and S-adenosylhomocysteine (SAH)<ref>PMID:25626150</ref>. S-adenosylornithine or sinefugin (SFG) is an inhibitor of the reaction. | '''tRNA methyltransferase''' (Trm) methylates specific positions on tRNA. tRNA methylation is a post-transcription modification. The methyl donor is S-adenosylmethionine (SAM) and the product is the methylated tRNA and S-adenosylhomocysteine (SAH)<ref>PMID:25626150</ref>. S-adenosylornithine or sinefugin (SFG) is an inhibitor of the reaction. | ||
| Line 6: | Line 6: | ||
== Structural highlights == | == Structural highlights == | ||
| - | Trm active site is located at the cleft at the subunit interface<ref>PMID:12773376</ref>. | + | Trm <scene name='55/553109/Cv/4'>active site</scene> is located at the cleft at the subunit interface<ref>PMID:12773376</ref>. Water molecules are shown as red spheres. |
| - | + | ||
| - | + | ||
==3D structures of tRNA methyltransferase== | ==3D structures of tRNA methyltransferase== | ||
| + | [[tRNA methyltransferase 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | **[[3bt7]] – yTrm + RNA<br /> | ||
| - | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Swinehart WE, Jackman JE. Diversity in mechanism and function of tRNA methyltransferases. RNA Biol. 2015;12(4):398-411. doi: 10.1080/15476286.2015.1008358. PMID:25626150 doi:http://dx.doi.org/10.1080/15476286.2015.1008358
- ↑ Ahn HJ, Kim HW, Yoon HJ, Lee BI, Suh SW, Yang JK. Crystal structure of tRNA(m1G37)methyltransferase: insights into tRNA recognition. EMBO J. 2003 Jun 2;22(11):2593-603. PMID:12773376 doi:10.1093/emboj/cdg269