Cytochrome c
From Proteopedia
(Difference between revisions)
| (9 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' scene='Cytochrome_c/Cyt_c/1' caption='Cytochrome c with heme complex with sulfate (PDB code [[3cp5]])'> |
| - | The '''cytochrome ''c''''' (cyt ''c'') proteins are a superfamily belonging to the class of [http://en.wikipedia.org/wiki/All-α_proteins all-α proteins], which are denoted as such by having an α-helical core. Each protein in this superfamily also contains one or more covalently-bound [http://en.wikipedia.org/wiki/Heme heme prosthetic groups].<ref>PMID:11697912</ref><ref name=main /> The cyt ''c'' superfamily contains many different families, some of which are better characterized than others. These families include monodomain and multi-domain C-type cytochromes, such as [http://proteopedia.org/wiki/index.php/1etp cyt c4], a diheme C-type cytochrome, and [http://proteopedia.org/wiki/index.php/2ozy NrfB], a pentaheme C-type cytochrome. In particular, the monoheme cyt ''c'' from ''Rhodothermus marinus'' has been previously studied and provides an excellent example of how some protein characteristics and structures can be extremely diverse, yet conserved, through evolution.<br /> For details on decaheme cyt see [[MtrF]].<br /> For details on Cyt c7 see [[Cytochrome c 7]]. | + | The '''cytochrome ''c''''' (cyt ''c'') proteins are a superfamily belonging to the class of [http://en.wikipedia.org/wiki/All-α_proteins all-α proteins], which are denoted as such by having an α-helical core. Each protein in this superfamily also contains one or more covalently-bound [http://en.wikipedia.org/wiki/Heme heme prosthetic groups].<ref>PMID:11697912</ref><ref name=main /> The cyt ''c'' superfamily contains many different families, some of which are better characterized than others. These families include monodomain and multi-domain C-type cytochromes, such as [http://proteopedia.org/wiki/index.php/1etp cyt c4], a diheme C-type cytochrome, and [http://proteopedia.org/wiki/index.php/2ozy NrfB], a pentaheme C-type cytochrome. In particular, the monoheme cyt ''c'' from ''Rhodothermus marinus'' has been previously studied and provides an excellent example of how some protein characteristics and structures can be extremely diverse, yet conserved, through evolution.<br /> For details on decaheme cyt see [[MtrF]].<br /> For details on Cyt c7 see [[Cytochrome c 7]].<br /> |
| + | See also [[Cytochrome C -Adis]], [[Hemeproteins]], [[Cytochrome C (Hebrew)]], [[Cytochrome C (arabic)]]. | ||
== Introduction == | == Introduction == | ||
| Line 14: | Line 15: | ||
<scene name='Sandbox_Reserved_335/Heme/1'>'Figure 1. The heme group of monoheme cytochrome ''c'' purified from ''Rhodothermus marinus''</scene> | <scene name='Sandbox_Reserved_335/Heme/1'>'Figure 1. The heme group of monoheme cytochrome ''c'' purified from ''Rhodothermus marinus''</scene> | ||
| - | All members in the C-type cytochrome superfamily contain a heme prosthetic group that is covalently attached to the protein via two thioether bonds to cysteine residues. Most cytochromes ''c'' occur in a | + | All members in the C-type cytochrome superfamily contain a heme prosthetic group that is covalently attached to the protein via two thioether bonds to cysteine residues. Most cytochromes ''c'' occur in a where the histidine residue is one of the two axial ligands of the heme iron.<ref name=main>PMID:18855424</ref><ref name=heme /> In monoheme cytochromes ''c'', the other axial position may be left vacant or be occupied by histidine or methionine residues; however, it can sometimes be occupied by cysteine or lysine residues.<ref name=main />. In ''Rm''cyt''c'', XX represents a threonine (Thr46) and an alanine residue (Ala47) that help form the loop 2 structure. |
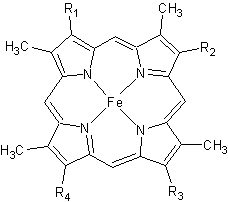
[[Image:heme.gif |frame|left| Figure 2. The tetrapyrrolic heme prosthetic group that can either be covalently attached to or closely associated with various proteins, such as cytochromes and other globin proteins. In ''Rm''cyt''c'', R2 is an ethyl group covalently attached to Cys 45, and R3 is a methyl group covalently attached to Cys48.]] | [[Image:heme.gif |frame|left| Figure 2. The tetrapyrrolic heme prosthetic group that can either be covalently attached to or closely associated with various proteins, such as cytochromes and other globin proteins. In ''Rm''cyt''c'', R2 is an ethyl group covalently attached to Cys 45, and R3 is a methyl group covalently attached to Cys48.]] | ||
| Line 64: | Line 65: | ||
* <scene name='69/696899/Cv/14'>Click here to see morph of this scene</scene>. | * <scene name='69/696899/Cv/14'>Click here to see morph of this scene</scene>. | ||
Only a proximal 5-coordinate NO adduct, confirmed by structural data, is observed with no detectable hexacoordinate distal NO adduct. | Only a proximal 5-coordinate NO adduct, confirmed by structural data, is observed with no detectable hexacoordinate distal NO adduct. | ||
| - | </StructureSection> | ||
==3D structures of cytochrome C== | ==3D structures of cytochrome C== | ||
| + | [[Cytochrome C 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | **[[3qvy]], [[3qw0]], [[3qw1]] – EcCyt + Zn<br /> | ||
| - | **[[3c62]], [[3c63]], [[3iq5]], [[3iq6]], [[3l1m]], [[3m15]], [[3nmi]], [[3nmk]] - EcCyt (mutant) + Zn<br /> | ||
| - | **[[3de8]], [[3m79]] - EcCyt (mutant) + Zn + Cu<br /> | ||
| - | **[[3de9]], [[3nmj]] - EcCyt (mutant) + Zn + Ni | ||
| - | }} | ||
== References == | == References == | ||
Current revision
| |||||||||||
References
- ↑ Gough J, Karplus K, Hughey R, Chothia C. Assignment of homology to genome sequences using a library of hidden Markov models that represent all proteins of known structure. J Mol Biol. 2001 Nov 2;313(4):903-19. PMID:11697912 doi:10.1006/jmbi.2001.5080
- ↑ 2.00 2.01 2.02 2.03 2.04 2.05 2.06 2.07 2.08 2.09 2.10 2.11 2.12 2.13 2.14 2.15 Stelter M, Melo AM, Pereira MM, Gomes CM, Hreggvidsson GO, Hjorleifsdottir S, Saraiva LM, Teixeira M, Archer M. A Novel Type of Monoheme Cytochrome c: Biochemical and Structural Characterization at 1.23 A Resolution of Rhodothermus marinus Cytochrome c. Biochemistry. 2008 Oct 15. PMID:18855424 doi:10.1021/bi800999g
- ↑ 3.0 3.1 3.2 Reedy CJ, Gibney BR. Heme protein assemblies. Chem Rev. 2004 Feb;104(2):617-49. PMID:14871137 doi:10.1021/cr0206115
- ↑ 4.0 4.1 4.2 Ambler RP. Sequence variability in bacterial cytochromes c. Biochim Biophys Acta. 1991 May 23;1058(1):42-7. PMID:1646017
- ↑ Cookson DJ, Moore GR, Pitt RC, Williams RJP, Campbell ID, Ambler RP, Bruschi M, Le Gall J. Structural homology of cytochromes c. Eur J Biochem. 1978 Feb;83(1):261-75.
- ↑ Than ME, Hof P, Huber R, Bourenkov GP, Bartunik HD, Buse G, Soulimane T. Thermus thermophilus cytochrome-c552: A new highly thermostable cytochrome-c structure obtained by MAD phasing. J Mol Biol. 1997 Aug 29;271(4):629-44. PMID:9281430 doi:10.1006/jmbi.1997.1181
- ↑ Soares CM, Baptista AM, Pereira MM, Teixeira M. Investigation of protonatable residues in Rhodothermus marinus caa3 haem-copper oxygen reductase: comparison with Paracoccus denitrificans aa3 haem-copper oxygen reductase. J Biol Inorg Chem. 2004 Mar;9(2):124-34. Epub 2003 Dec 23. PMID:14691678 doi:10.1007/s00775-003-0509-9
- ↑ Pereira MM, Santana M, Teixeira M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim Biophys Acta. 2001 Jun 1;1505(2-3):185-208. PMID:11334784
- ↑ 9.0 9.1 9.2 9.3 9.4 9.5 Karp, Gerald (2008). Cell and Molecular Biology (5th edition). Hoboken, NJ: John Wiley & Sons. ISBN 978-0470042175.
- ↑ Rajagopal BS, Wilson MT, Bendall DS, Howe CJ, Worrall JA. Structural and kinetic studies of imidazole binding to two members of the cytochrome c (6) family reveal an important role for a conserved heme pocket residue. J Biol Inorg Chem. 2011 Jan 26. PMID:21267610 doi:10.1007/s00775-011-0758-y
- ↑ Morelli X, Czjzek M, Hatchikian CE, Bornet O, Fontecilla-Camps JC, Palma NP, Moura JJ, Guerlesquin F. Structural model of the Fe-hydrogenase/cytochrome c553 complex combining transverse relaxation-optimized spectroscopy experiments and soft docking calculations. J Biol Chem. 2000 Jul 28;275(30):23204-10. PMID:10748163 doi:10.1074/jbc.M909835199
- ↑ Manole A, Kekilli D, Svistunenko DA, Wilson MT, Dobbin PS, Hough MA. Conformational control of the binding of diatomic gases to cytochrome c'. J Biol Inorg Chem. 2015 Mar 20. PMID:25792378 doi:http://dx.doi.org/10.1007/s00775-015-1253-7
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, David Canner, Joel L. Sussman, Melissa Morrison, Adis Hasic