Sandbox Reserved 1051
From Proteopedia
(Difference between revisions)
| (4 intermediate revisions not shown.) | |||
| Line 4: | Line 4: | ||
== Background == | == Background == | ||
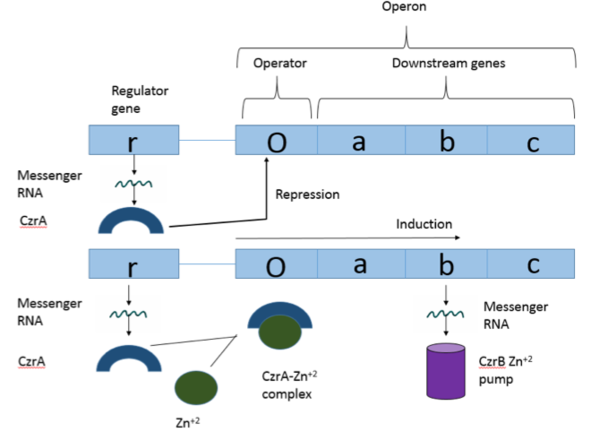
| - | [https://en.wikipedia.org/wiki/Operon Operons] are a critical genetic component of most prokaryotic cells. There are many different operons that are responsible for the production of proteins with a wide range of functions, the most well-known of which are the Lac and Trp operons, responsible for producing enzymes which metabolize lactose and tryptophan respectively. Despite many differences in each operon and the proteins that they encode, operons all function in the same general manner. Structurally, each operon contains a regulator, an operator, and one or more structural genes. The regulator protein is responsible for managing the expression level of the structural genes, the operator is similar to a promoter in a regular gene and is where transcription begins, and the structural genes code for proteins. The regulator protein (produced as a result of expression of the regulator gene) most often acts in a repressive manner, though this is not always the case. That is, the regulator protein will bind to the operator of the operon, inhibiting the binding and/or progression of RNA polymerase to the structural genes, thus inhibiting transcription of the genes into mRNA. If the regulator protein were to consistently be active, there could never be adequate expression of the structural genes, so there must be a way to inactive the regulator protein, thus enabling expression of the structural genes. This is achieved through the binding of an inhibitor to the regulator protein. Since regulator proteins are DNA binding proteins, often this inhibition is allosteric rather than competitive, that is the inhibitor is not something that mimics DNA and binds to the active site physically blocking DNA from binding. Rather, the inhibitor of the regulator binds to somewhere other than the active site of the protein, changing it in some way which decreases the proteins affinity or ability to bind DNA. The Chromosome Determined Zinc Responsible (Czr) operon acts in exactly this manner, Czr A specifically is the regulator protein. The role of Czr A in the Czr operon is described in further detail under biological function. | + | [https://en.wikipedia.org/wiki/Operon Operons] are a critical genetic component of most prokaryotic cells. There are many different operons that are responsible for the production of proteins with a wide range of functions, the most well-known of which are the Lac and Trp operons, responsible for producing enzymes which metabolize lactose and tryptophan respectively. Despite many differences in each operon and the proteins that they encode, operons all function in the same general manner. Structurally, each operon contains a regulator, an operator, and one or more structural genes. The regulator protein is responsible for managing the expression level of the structural genes, the operator is similar to a promoter in a regular gene and is where transcription begins, and the structural genes code for proteins. The regulator protein (produced as a result of expression of the regulator gene) most often acts in a repressive manner, though this is not always the case. That is, the regulator protein will bind to the operator of the operon, inhibiting the binding and/or progression of [http://proteopedia.org/wiki/index.php/RNA_polymerase RNA polymerase] to the structural genes, thus inhibiting transcription of the genes into mRNA. If the regulator protein were to consistently be active, there could never be adequate expression of the structural genes, so there must be a way to inactive the regulator protein, thus enabling expression of the structural genes. This is achieved through the binding of an inhibitor to the regulator protein. Since regulator proteins are DNA binding proteins, often this inhibition is allosteric rather than competitive, that is the inhibitor is not something that mimics DNA and binds to the active site physically blocking DNA from binding. Rather, the inhibitor of the regulator binds to somewhere other than the active site of the protein, changing it in some way which decreases the proteins affinity or ability to bind DNA. The Chromosome Determined Zinc Responsible (Czr) operon acts in exactly this manner, Czr A specifically is the regulator protein. The role of Czr A in the Czr operon is described in further detail under biological function. |
In addition to being a component of an operon, Czr A is also considered to be a metal sensor protein. While the immediate function of Czr A is gene regulation, this serves the larger purpose of acting to maintain an appropriate concentration of Zn <sup>2+</sup> in the cell. | In addition to being a component of an operon, Czr A is also considered to be a metal sensor protein. While the immediate function of Czr A is gene regulation, this serves the larger purpose of acting to maintain an appropriate concentration of Zn <sup>2+</sup> in the cell. | ||
| + | |||
| + | [[Image:operon.png|600px|thumb|center|Visual Overview of Czr Operon]] | ||
== Biological Function == | == Biological Function == | ||
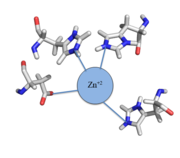
| - | Czr A is a transcriptional repressor protein responsible for the regulation of the Czr operon. The Czr operon contains genes for the proteins Czr A and | + | Czr A is a transcriptional repressor protein responsible for the regulation of the Czr operon <ref>DOI: 10.1073/pnas.0905558106</ref>. The Czr operon contains genes for the proteins Czr A and [http://proteopedia.org/wiki/index.php/3byr CzrB]. Czr B is a Zinc transport protein which moves Zn<sup>2+</sup> out of a cell while Czr A regulates this process by controlling expression level of Czr B. When relatively low amounts of zinc are present in the cell Czr A will bind to DNA, preventing the progression of RNA polymerase and thus inhibiting expression of Czr B<ref>DOI: 10.1073/pnas.0905558106</ref>. Decreased expression of Czr B results in the ability of the cell to retain Zn<sup>2+</sup> more readily. Because Czr A and Czr B are transcribed as part of the same operon, an inhibitor of Czr A must be readily available to allow full transcription of Czr B when necessary. Czr A is noncompetitively inhibited by the binding of two Zn<sup>2+</sup> ions<ref>DOI: 10.1073/pnas.0905558106</ref>, which is ideal in that this allows for expression of Czr B, a Zn<sup>2+</sup> transporter to be dependent on the relative amount of Zn<sup>2+</sup> in the cell. Czr A displays two different conformations; the first typically binds DNA and has relatively low affinity for Zn<sup>2+</sup>, in this conformation the <scene name='69/694220/A5_helices__dna_binding/1'>a5 helices are open</scene>. The <scene name='69/694220/A5_helices_dna_binding/1'>a5 helices swing down</scene> to achieve the other conformation which binds two Zn<sup>2+</sup> ions and has relatively low affinity for DNA. |
===DNA Binding === | ===DNA Binding === | ||
Czr A performs it's primary function when bound to DNA. Each monomeric subunit of the protein binds DNA individually, coming together once attached to the DNA. While bound, Czr A prevents the transcription of the DNA in the Czr operon, acting as a repressor protein and effectively turning off the operon. As was briefly mentioned above, the Czr operon contains the gene responsible for producing Czr B, a metal transport protein which regulates the concentration of zinc in the cell. So, by extension, Czr A is responsible for retaining Zn<sup>2+</sup> inside the cell by inhibiting the production of the protein responsible for transporting zinc out of the cell. | Czr A performs it's primary function when bound to DNA. Each monomeric subunit of the protein binds DNA individually, coming together once attached to the DNA. While bound, Czr A prevents the transcription of the DNA in the Czr operon, acting as a repressor protein and effectively turning off the operon. As was briefly mentioned above, the Czr operon contains the gene responsible for producing Czr B, a metal transport protein which regulates the concentration of zinc in the cell. So, by extension, Czr A is responsible for retaining Zn<sup>2+</sup> inside the cell by inhibiting the production of the protein responsible for transporting zinc out of the cell. | ||
| Line 18: | Line 20: | ||
== Binding of DNA == | == Binding of DNA == | ||
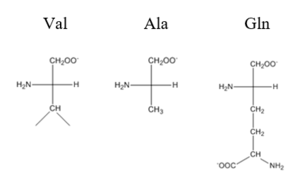
| - | The <scene name='69/694219/Serandhisresidues/2'>main DNA interactions</scene> have been found to occur at the Ser 54 and 57 along with His 58 residues. These residues are likely to interact with the 5'-TGAA sequence found in the half-site of the DNA. These residues are found in the N terminal of the R helix. The residues involved in the <scene name='69/694219/Dna_binding_pocket/1'>DNA binding pocket</scene> are Val 42 and Gln 53. This was experimentally determined by mutating the Gln and Val with Ala residues and measuring the binding capacity; In a previously published article <ref>DOI: 10. | + | The <scene name='69/694219/Serandhisresidues/2'>main DNA interactions</scene> have been found to occur at the Ser 54 and 57 along with His 58 residues. These residues are likely to interact with the 5'-TGAA sequence found in the half-site of the DNA. These residues are found in the N terminal of the R helix. The residues involved in the <scene name='69/694219/Dna_binding_pocket/1'>DNA binding pocket</scene> are Val 42 and Gln 53. This was experimentally determined by mutating the Gln and Val with Ala residues and measuring the binding capacity; In a previously published article <ref>DOI: 10.1073/pnas.0905558106</ref>, the DNA bound state of CzrA was tested by using the known critical residues for DNA interactions. Critical residues, Gln53, Val42, Ser54, Ser57, and His58, were replaced with Ala and then compared to the kinetics of the wild type protein. Replacing only the Q53 and V42 residues resulted in an 11-fold and 160-fold decrease in K<sub>a</sub>, respectively. Other residues such as S54, S57, and H58 were also replaced with Ala residues, and it was found that these mutations caused binding similar to the fully inhibited Zn<sup>2+</sup> bound state. Table 1 in this same article shows the different K<sub>observed</sub>, and the measured decrease in K<sub>observed</sub> for each mutation. The bind between the DNA and the protein can be attributed to losing certain intermolecular forces such as possible hydrogen bonding when changing from Gln and Ala, and a loss of London Dispersion forces in the Val to Ala change. [[Image:Capture01.PNG|300px|center|thumb| Comparison of Val, Ala, and Gln residues]] |
The differences in binding favorability can also be seen when comparing the ΔG for the Apo-state vs. the DNA bound state and the Zinc vs. the Zinc and DNA bound state. These ΔGs were found to be -15.2kcal/mol and -9kcal/mol respectively<ref>DOI: 10.1021/ja208047b</ref>. This agrees with previously published data showing the Zinc binding inhibits the affinity the protein has to DNA. | The differences in binding favorability can also be seen when comparing the ΔG for the Apo-state vs. the DNA bound state and the Zinc vs. the Zinc and DNA bound state. These ΔGs were found to be -15.2kcal/mol and -9kcal/mol respectively<ref>DOI: 10.1021/ja208047b</ref>. This agrees with previously published data showing the Zinc binding inhibits the affinity the protein has to DNA. | ||
Current revision
| This Sandbox is Reserved from 02/09/2015, through 05/31/2016 for use in the course "CH462: Biochemistry 2" taught by Geoffrey C. Hoops at the Butler University. This reservation includes Sandbox Reserved 1051 through Sandbox Reserved 1080. |
To get started:
More help: Help:Editing |
CzrA
| |||||||||||