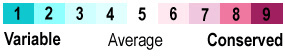This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1058
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
==Introduction== | ==Introduction== | ||
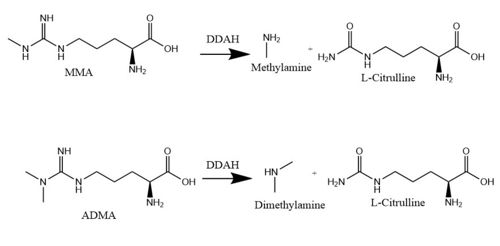
| - | Dimethylarginine | + | <scene name='75/752351/Ddah/2'>Dimethylarginine Dimethylaminohydrolase</scene> <span class="plainlinks">[http://www.chem.qmul.ac.uk/iubmb/enzyme/EC3/5/3/18.html EC 3.5.3.18]</span> (commonly known as DDAH) is a member of the <span class="plainlinks">[https://en.wikipedia.org/wiki/Hydrolase hydrolase]</span> family of enzymes which use water to break down molecules <ref name="palm">Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/17933965 17933965]</span> doi:<span class="plainlinks">[http://ajpheart.physiology.org/content/293/6/H3227 10.1152/ajpheart.00998.2007]</span></ref>. Additionally, DDAH is a <span class="plainlinks">[https://en.wikipedia.org/wiki/Nitric_oxide_synthase nitric oxide synthase (NOS)]</span> regulator. It metabolizes free arginine derivatives, namely <span class="plainlinks">[https://en.wikipedia.org/wiki/Asymmetric_dimethylarginine N<sup>Ѡ</sup>,N<sup>Ѡ</sup>-dimethyl-L-arginine (ADMA)]</span> and <span class="plainlinks">[https://en.wikipedia.org/wiki/Methylarginine N<sup>Ѡ</sup>-methyl-L-arginine (MMA)]</span>, which competitively inhibit NOS <ref name="tran">Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/14664901 14664901]</span> doi:<span class="plainlinks">[http://www.sciencedirect.com/science/article/pii/S1567568803000321 10.1016/S1567-5688(03)00032-1]</span></ref>. DDAH converts MMA or ADMA to two products: <span class="plainlinks">[https://en.wikipedia.org/wiki/Citrulline L-citrulline]</span> and an amine <ref name="frey">Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:<span class="plainlinks">[https://www.ncbi.nlm.nih.gov/pubmed/16698551]</span> doi:<span class="plainlinks">[http://www.sciencedirect.com/science/article/pii/S0969212606001717 10.1016/j.str.2006.03.006]</span></ref> (Figure 1). DDAH is expressed in the cytosol of cells in humans, mice, rats, sheep, cattle, and bacteria <ref name="palm" />. DDAH activity has been localized mainly to the brain, kidneys, pancreas, and liver in these organisms. Presented in this page is information from DDAH isoform 1 (DDAH-1); however, there are two different isoforms <ref name="frey" />. |
[[Image:DDAH mechanism.jpg|500 px|center|thumb|'''Figure 1.''' The normal DDAH mechanism]] | [[Image:DDAH mechanism.jpg|500 px|center|thumb|'''Figure 1.''' The normal DDAH mechanism]] | ||
| Line 17: | Line 17: | ||
====Lid Region Conservation==== | ====Lid Region Conservation==== | ||
| - | The specific residues in the lid region vary between organisms <ref name="frey" /> (Figure 2). | + | The specific residues in the lid region vary between organisms <ref name="frey" /> (Figure 2). Notable in this image is a <span class="plainlinks">[https://en.wikipedia.org/wiki/Conserved_sequence conserved]</span> leucine <scene name='75/752351/Hbond_leu29/9'>(Leu29)</scene> residue in this led that functions to hydrogen bond with the <span class="plainlinks">[https://en.wikipedia.org/wiki/Ligand ligand]</span> bound to the active site in DDAH-1 but not in DDAH-2 <ref name="rasheed" /> (Figure 2). Different <span class="plainlinks">[https://en.wikipedia.org/wiki/Protein_isoform isoforms]</span> from the same species can have differences in lid regions as well <ref name="frey" />. DDAH-2 has a negatively charged lid while DDAH-1 has a positively charged lid <ref name="frey" />. |
| - | [[Image:Lid Region | + | [[Image:WebLogo for Lid Region.png|500 px|center|thumb|'''Figure 2.''' WebLogo for the lid region in DDAH-1 of eleven different organisms.]] |
===Active Site=== | ===Active Site=== | ||
Current revision
Dimethylarginine Dimethylaminohydrolase
| |||||||||||
References
- ↑ 1.0 1.1 Palm F, Onozato ML, Luo Z, Wilcox CS. Dimethylarginine dimethylaminohydrolase (DDAH): expression, regulation, and function in the cardiovascular and renal systems. American Journal of Physiology. 2007 Dec 1;293(6):3227-3245. PMID:17933965 doi:10.1152/ajpheart.00998.2007
- ↑ 2.0 2.1 2.2 Tran CTL, Leiper JM, Vallance P. The DDAH/ADMA/NOS pathway. Atherosclerosis Supplements. 2003 Dec;4(4):33-40. PMID:14664901 doi:10.1016/S1567-5688(03)00032-1
- ↑ 3.00 3.01 3.02 3.03 3.04 3.05 3.06 3.07 3.08 3.09 3.10 3.11 3.12 3.13 3.14 3.15 3.16 3.17 3.18 3.19 3.20 3.21 3.22 3.23 3.24 Frey D, Braun O, Briand C, Vasak M, Grutter MG. Structure of the mammalian NOS regulator dimethylarginine dimethylaminohydrolase: a basis for the design of specific inhibitors. Structure. 2006 May;14(5):901-911. PMID:[1] doi:10.1016/j.str.2006.03.006
- ↑ Humm A, Fritsche E, Mann K, Göhl M, Huber R. Recombinant expression and isolation of human L-arginine:glycine amidinotransferase and identification of its active-site cysteine residue. Biochemical Journal. 1997 March 15;322(3):771-776. PMID:9148748 doi:10.1042/bj3220771
- ↑ 5.0 5.1 5.2 Rasheed M, Richter C, Chisty LT, Kirkpatrick J, Blackledge M, Webb MR, Driscoll PC. Ligand-dependent dynamics of the active site lid in bacterial Dimethyarginine Dimethylaminohydrolase. Biochemistry. 2014 Feb 18;53:1092-1104. PMCID:PMC3945819 doi:10.1021/bi4015924
- ↑ 6.0 6.1 6.2 Stone EM, Costello AL, Tierney DL, Fast W. Substrate-assisted cysteine deprotonation in the mechanism of Dimethylargininase (DDAH) from Pseudomonas aeruginosa. Biochemistry. 2006 May 2;45(17):5618-5630. PMID:16634643 doi:10.1021/bi052595m
- ↑ 7.0 7.1 Pace NJ, Weerpana E. Zinc-binding cysteines: diverse functions and structural motifs. Biomolecules. 2014 June;4(2):419-434. PMCID:4101490 doi:10.3390/biom4020419
- ↑ Janssen W, Pullamsetti SS, Cooke J, Weissmann N, Guenther A, Schermuly RT. The role of dimethylarginine dimethylaminohydrolase (DDAH) in pulmonary fibrosis. The Journal of Pathology. 2012 Dec 12;229(2):242-249. Epub 2013 Jan. PMID:23097221 doi:10.1002/path.4127
- ↑ Colasanti M, Suzuki H. The dual personality of NO. ScienceDirect. 2000 Jul 1;21(7):249-252. PMID:10979862 doi:10.1016/S0165-6147(00)01499-1
- ↑ Rassaf T, Feelisch M, Kelm M. Circulating NO pool: assessment of nitrite and nitroso species in blood and tissues. Free Rad. Biol. Med. 2004 Feb 15;36(4):413-422. PMID:14975444 doi:10.1016/j.freeradbiomed.2003.11.011
- ↑ Tsao PS, Cooke JP. Endothelial alterations in hypercholesterolemia: more than simply vasodilator dysfunction. Journal of Cardiovascular Pharmacology. 1998;32(3):48-53. PMID:9883748
- ↑ Vallance P, Leiper J. Blocking NO synthesis: how, where and why? Nat. Rev. Drug Discov. 2002 Dec;1(12):939-950. PMID:12461516 doi:10.1038/nrd960
Student Contributors
- Natalie Van Ochten
- Kaitlyn Enderle
- Colton Junod
3D Structures of Dimethylarginine Dimethylaminohydrolase
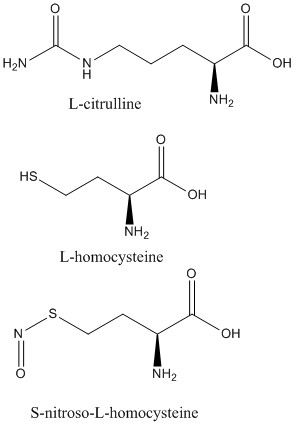
2C6Z L-citrulline bound to isoform 1
2CI1 S-nitroso-L-homocysteine bound to isoform 1
2CI3 crystal form 1
2CI4 crystal form 2
2CI5 L-homocysteine bound to isoform 1
2CI6 Zn (II) bound at low pH to isoform 1
2CI7 Zn (II) bound at high pH to isoform 1