This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Nitroreductase
From Proteopedia
(Difference between revisions)
| (7 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
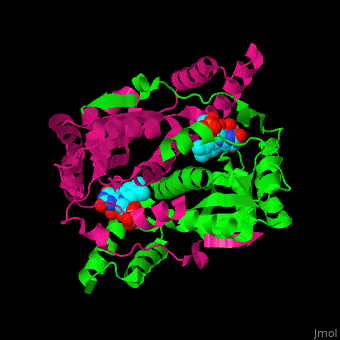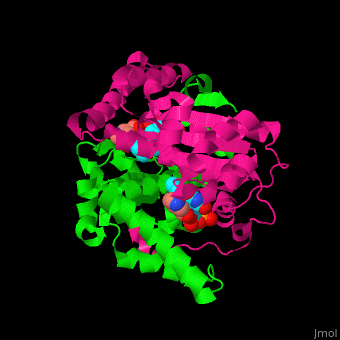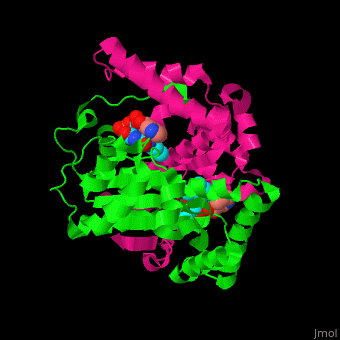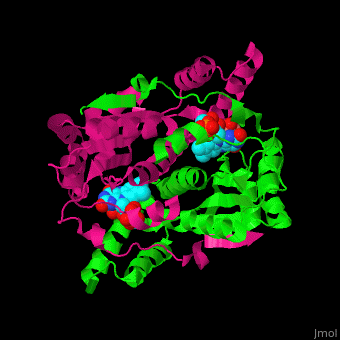
| - | <StructureSection load=' | + | <StructureSection load='' size='350' side='right' caption='E. coli nitroreductase dimer containing FMN and nicotinic acid (PDB entry [[1icr]])' scene='48/489319/Cv/1'> |
== Function == | == Function == | ||
| - | '''Nitroreductase''' (NR) is involved in the reduction of nitrogen-containing compounds including those containing NO<sub>2</sub> group. The NO<sub>2</sub> group is reduced to hydroxylamine<ref>PMID:11491290</ref>. NR uses FMN as cofactor. Among NRs are the oxygen-insensitive NAD(P)H NR and NADH dehydrogenase. | + | '''Nitroreductase''' or '''oxygen-insensitive NAD(P)H nitroreductase''' (NR) is involved in the reduction of nitrogen-containing compounds including those containing NO<sub>2</sub> group. The NO<sub>2</sub> group is reduced to hydroxylamine<ref>PMID:11491290</ref>. NR uses FMN as cofactor. Among NRs are the oxygen-insensitive NAD(P)H NR and NADH dehydrogenase. |
== Relevance == | == Relevance == | ||
| Line 8: | Line 8: | ||
== Structural highlights == | == Structural highlights == | ||
| - | NR <scene name='48/489319/Cv/ | + | NR <scene name='48/489319/Cv/4'>cofactor FMN is bound directly to one subunit</scene> while the <scene name='48/489319/Cv/5'>ligand - nicotinic acid - interacts with both subunits</scene>. Water molecules are shown as red spheres. |
| - | + | ==3D structures of nitroreductase== | |
| + | [[Nitroreductase 3D structures]] | ||
</StructureSection> | </StructureSection> | ||
| - | ==3D structures of nitroreductase== | ||
| - | |||
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| - | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| - | |||
| - | *Nitroreductase | ||
| - | |||
| - | **[[1f5v ]], [[1oo5]] – EcNR – ''Escherichia coli''<br /> | ||
| - | **[[3x21]], [[3x22]] – EcNR (mutant)<br /> | ||
| - | **[[3bm1]], [[1ds7]] – EcNR minimal<br /> | ||
| - | **[[3bm2]] – EcNR minimal -FMN<br /> | ||
| - | **[[1nec]], [[1kqd]] – EclNR – ''Enterobacter cloacae''<br /> | ||
| - | **[[1ywq]], [[2i7h]] – NR – ''Bacillus cereus''<br /> | ||
| - | **[[2b67]] – NR – ''Streptococcus pneumonia''<br /> | ||
| - | **[[3ek3]], [[3hoi]] – NR – ''Bacterioides fragilis''<br /> | ||
| - | **[[3g14]] – NR – ''Clostridium novyi''<br /> | ||
| - | **[[3h4o]], [[3koq]] - NR – ''Clostridium difficile''<br /> | ||
| - | **[[3gr3]] – NR – ''Bartonella henselae''<br /> | ||
| - | **[[3hzn]] – NR – ''Salmonella typhimurium''<br /> | ||
| - | **[[3k6h]] – NR – ''Agrobacterium tumefaciens''<br /> | ||
| - | **[[2wzv]], [[2ymv]] – MsNR – ''Mycobacterium smegmatis''<br /> | ||
| - | **[[3of4]] – NR FMN/FAD – ''Idiomarina loihiensis''<br /> | ||
| - | **[[3n2s]] – NR – ''Bacillus subtilis''<br /> | ||
| - | **[[3pxv]] – NR – ''Desulfitobacterium hafniense''<br /> | ||
| - | **[[3qdl]] – NR – ''Helicobacter pylori''<br /> | ||
| - | **[[3r5l]], [[3r5p]] - MtNR – ''Mycobacterium tuberculosis''<br /> | ||
| - | **[[3r5r]], [[3r5w]] – MtNR F420<br /> | ||
| - | **[[3ge6]] – NR – ''Exiguobacterium sibiricum''<br /> | ||
| - | **[[4dn2]] – NR – ''Geobacter metallireducens''<br /> | ||
| - | **[[5ko7]], [[5ko8]], [[5krd]] – NR – ''Haliscomenobacter hydrossis''<br /> | ||
| - | |||
| - | *Copper-induced nitroreductase | ||
| - | |||
| - | **[[2wqf]] – LlNR – ''Lactococcus lactis''<br /> | ||
| - | **[[4bn6]] – LlNR + chloramphenicol<br /> | ||
| - | **[[4bn7]], [[4bn8]] – LlNR + phenol derivative<br /> | ||
| - | **[[4bn9]] – LlNR + nicotinic acid<br /> | ||
| - | **[[4bnb]] – LlNR + nitroquinoline<br /> | ||
| - | |||
| - | *Nitroreductase binary complex | ||
| - | **[[1kqb]], [[5j8g]]– EclNR + benzoate derivative<br /> | ||
| - | **[[1kqc]] – EclNR + acetate<br /> | ||
| - | **[[1ylr]], [[1ylu]] - EcNR + acetate<br /> | ||
| - | **[[1icr]], [[1icu]], [[1icv]], [[5j8d]] – EcNR + NAD derivative<br /> | ||
| - | **[[1ooq]] – EcNR + dicoumarol<br /> | ||
| - | **[[1oo6]], [[1oon]], [[1idt]] – EcNR + benzamide derivative<br /> | ||
| - | **[[1yki]] – EcNR + antibiotic<br /> | ||
| - | **[[2wzw]] - MsNR + NADPH | ||
| - | }} | ||
== References == | == References == | ||
<references/> | <references/> | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
Current revision
| |||||||||||
References
- ↑ Lovering AL, Hyde EI, Searle PF, White SA. The structure of Escherichia coli nitroreductase complexed with nicotinic acid: three crystal forms at 1.7 A, 1.8 A and 2.4 A resolution. J Mol Biol. 2001 May 25;309(1):203-13. PMID:11491290 doi:10.1006/jmbi.2001.4653
- ↑ Williams EM, Little RF, Mowday AM, Rich MH, Chan-Hyams JV, Copp JN, Smaill JB, Patterson AV, Ackerley DF. Nitroreductase gene-directed enzyme prodrug therapy: insights and advances toward clinical utility. Biochem J. 2015 Oct 15;471(2):131-53. doi: 10.1042/BJ20150650. PMID:26431849 doi:http://dx.doi.org/10.1042/BJ20150650