|
|
| (2 intermediate revisions not shown.) |
| Line 1: |
Line 1: |
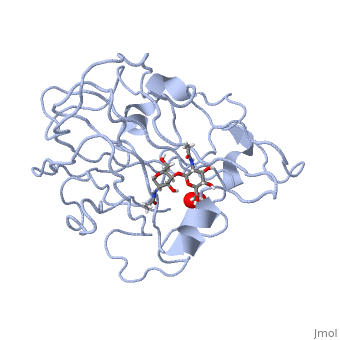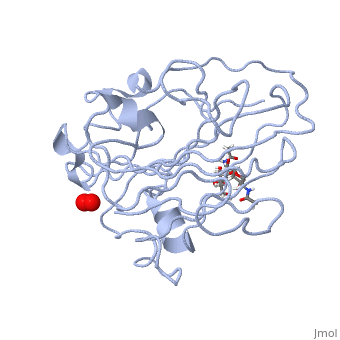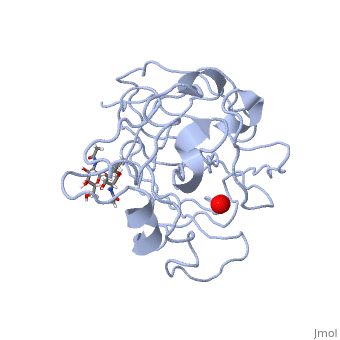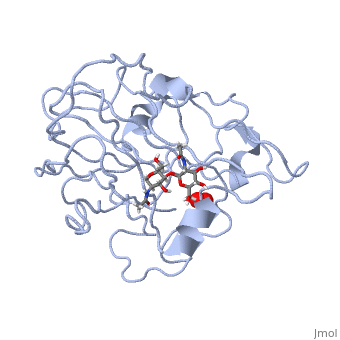
| | <StructureSection load='5tkh' size='350' side='right' scene='' caption='Glycosylated lytic polysaccharide monooxygenase complex with Cu(II) (orange) and peroxide (PDB code [[5tkh]])' > | | <StructureSection load='5tkh' size='350' side='right' scene='' caption='Glycosylated lytic polysaccharide monooxygenase complex with Cu(II) (orange) and peroxide (PDB code [[5tkh]])' > |
| - | | + | __TOC__ |
| | == Function == | | == Function == |
| | | | |
| Line 9: |
Line 9: |
| | In recent years there has been a significant interest in describing the interactions of copper-containing enzymes with O<sub>2</sub>/H<sub>2</sub>O<sub>2</sub>-derived species. The short-lived intermediates resulting from the activation of dioxygen are the key players in the mechanistic cycles in many metalloenzymes. In the enzyme <scene name='Journal:JBIC:17/Cv/3'>peptidylglycine alpha-hydroxylating monooxygenase (PHM)</scene> various reduced Cu/oxygen species have been proposed to act as catalytically competent intermediates, yet their exact nature and their role in the enzymatic reaction is still unknown. | | In recent years there has been a significant interest in describing the interactions of copper-containing enzymes with O<sub>2</sub>/H<sub>2</sub>O<sub>2</sub>-derived species. The short-lived intermediates resulting from the activation of dioxygen are the key players in the mechanistic cycles in many metalloenzymes. In the enzyme <scene name='Journal:JBIC:17/Cv/3'>peptidylglycine alpha-hydroxylating monooxygenase (PHM)</scene> various reduced Cu/oxygen species have been proposed to act as catalytically competent intermediates, yet their exact nature and their role in the enzymatic reaction is still unknown. |
| | Structural and other studies showed that peptidylglycine α-hydroxylating monooxygenase (PHM) contains <scene name='Journal:JBIC:17/Cv/4'>two non-equivalent copper sites (CuH and CuM)</scene>. CuM serves as an oxygen binding and hydrogen abstraction site, CuH is involved in electron transfer. In the structure of Cu(II)-PHM complexed with hydrogen peroxide determined to 1.98 Å resolution, <scene name='Journal:JBIC:17/Cv/7'>(hydro)peroxide binds exclusively to CuM in a slightly asymmetric side-on mode</scene>. The <scene name='Journal:JBIC:17/Cv/8'>interatomic O-O distance of the copper-bound ligand is 1.5, characteristic of peroxide/hydroperoxide species, and the copper-oxygen distances are 2.0 and 2.1</scene> Å. This Cu(II)-bound <scene name='Journal:JBIC:17/Cv/9'>peroxo moiety interacts closely with a molecule of water</scene>, forming <scene name='Journal:JBIC:17/Cv/10'>hydrogen bonds that stabilize the structure</scene>. DFT and QM/MM calculations indicate that this species is a Cu-bound doubly deprotonated peroxidate and that its energy is similar to that of its isomer Cu(I)-bound superoxide. | | Structural and other studies showed that peptidylglycine α-hydroxylating monooxygenase (PHM) contains <scene name='Journal:JBIC:17/Cv/4'>two non-equivalent copper sites (CuH and CuM)</scene>. CuM serves as an oxygen binding and hydrogen abstraction site, CuH is involved in electron transfer. In the structure of Cu(II)-PHM complexed with hydrogen peroxide determined to 1.98 Å resolution, <scene name='Journal:JBIC:17/Cv/7'>(hydro)peroxide binds exclusively to CuM in a slightly asymmetric side-on mode</scene>. The <scene name='Journal:JBIC:17/Cv/8'>interatomic O-O distance of the copper-bound ligand is 1.5, characteristic of peroxide/hydroperoxide species, and the copper-oxygen distances are 2.0 and 2.1</scene> Å. This Cu(II)-bound <scene name='Journal:JBIC:17/Cv/9'>peroxo moiety interacts closely with a molecule of water</scene>, forming <scene name='Journal:JBIC:17/Cv/10'>hydrogen bonds that stabilize the structure</scene>. DFT and QM/MM calculations indicate that this species is a Cu-bound doubly deprotonated peroxidate and that its energy is similar to that of its isomer Cu(I)-bound superoxide. |
| - | </StructureSection> | |
| | | | |
| | ==3D structures of monooxygenase== | | ==3D structures of monooxygenase== |
| | + | [[Monooxygenase 3D structures]] |
| | | | |
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}}
| + | </StructureSection> |
| - | {{#tree:id=OrganizedByTopic|openlevels=0|
| + | |
| - | | + | |
| - | *'''Toluene 4-monooxygenase (TMO)'''
| + | |
| - | | + | |
| - | **[[3dhg]], [[3i5j]], [[2inc]] – PmTMO α+β+γ subunits + Fe – ''Pseudomonas mendocina''<br />
| + | |
| - | **[[2ind]] – PmTMO α+β+γ subunits + Mn <br />
| + | |
| - | **[[3rng]], [[3rnf]], [[3rne]], [[3rnc]], [[3rnb]], [[3rna]], [[3rn9]], [[3n20]], [[3n1z]], [[3n1y]], [[3n1x]], [[2rdb]] – PmTMO α+β+γ subunits (mutant) + Fe <br />
| + | |
| - | **[[1sjg]] – PmTMO protein C + Fe <br />
| + | |
| - | **[[1vm9]] – PmTMO protein C (mutant) + Fe <br />
| + | |
| - | **[[2bf5]], [[2bf3]], [[2bf2]] – PmTMO protein D <br />
| + | |
| - | **[[4wqm]] – PmTMO (mutant) + Ni <br />
| + | |
| - | **[[4p1c]], [[4p1b]] – PmTMO protein A+B+E+ferredoxin (mutant) + Fe <br />
| + | |
| - | **[[2q3w]] – PmTMO ferredoxin subunit (mutant) + Fe <br />
| + | |
| - | | + | |
| - | *TMO complexes
| + | |
| - | | + | |
| - | **[[3dhh]], [[3q3n]], [[3q3o]] – PmTMO α+β+γ subunits + Fe + phenol derivative + TMO system effector protein<br />
| + | |
| - | **[[3rmk]], [[1t0s]], [[1t0q]] – PmTMO α+β+γ subunits + Fe + phenol derivative<br />
| + | |
| - | **[[3q14]] – PmTMO α+β+γ subunits + Fe + p-cresol + TMO system effector protein<br />
| + | |
| - | **[[3q2a]], [[3q3m]] – PmTMO α+β+γ subunits + Fe + benzoate inhibitor + TMO system effector protein<br />
| + | |
| - | **[[3dhi]] – PmTMO α+β+γ subunits + Fe + TMO system effector protein<br />
| + | |
| - | **[[3ge3]], [[3ge8]], [[3ri7]] – PmTMO α (mutant)+β+γ subunits + Fe + TMO system effector protein<br />
| + | |
| - | **[[3i63]] – PmTMO α+β+γ subunits + Fe + TMO system effector protein + H2O2<br />
| + | |
| - | **[[5tdv]] – PmTMO protein A+B+D+E + Fe + peroxide<br />
| + | |
| - | **[[5tdu]] – PmTMO protein A+B+D+E + Fe + cresol<br />
| + | |
| - | **[[5tdt]] – PmTMO protein A+B+D+E + Fe + peroxide + toluene<br />
| + | |
| - | **[[5tds]] – PmTMO protein A+B+D+E + Fe + toluene<br />
| + | |
| - | **[[1t0r]] – PsTMO + Fe + OH – ''Pseudomonas stutzeri''<br />
| + | |
| - | | + | |
| - | *'''Heme-degrading monooxygenase (IsdI)'''
| + | |
| - | | + | |
| - | **[[2zdp]] – SaIsdI + heme-Co – ''Staphylococcus aureus''<br />
| + | |
| - | **[[3lgm]], [[2zdo]] – SaIsdI + heme-Fe<br />
| + | |
| - | **[[4fnh]] – SaIsdI (mutant) + heme-Fe<br />
| + | |
| - | **[[3lgn]] – SaIsdI + heme-Fe + O2<br />
| + | |
| - | **[[3qgp]] – SaIsdI + heme-Fe + CN<br />
| + | |
| - | **[[4fni]] – SaIsdI (mutant) + heme-Fe + CN<br />
| + | |
| - | **[[4nl5]] – MtMO + heme + CN – ''Mycobacterium tuberculosis''<br />
| + | |
| - | **[[4oz5]] – MO + heme – ''Bacillus subtilis''<br />
| + | |
| - | | + | |
| - | *'''Kynurenine 3-monooxygenase (KMO)'''
| + | |
| - | | + | |
| - | **[[4j2w]], [[4j31]], [[4j33]], [[4j34]] – yKMO + FAD – yeast<br />
| + | |
| - | **[[4j36]] – yKMO + FAD + inhibitor<br />
| + | |
| - | **[[5na5]] – PfKMO (mutant) + FAD – ''Pseudomonas fluorescens''<br />
| + | |
| - | **[[5nak]] – PfKMO (mutant) + FAD + kynurenine <br />
| + | |
| - | **[[5nah]], [[5nag]], [[5nae]], [[5nab]], [[5n7t]], [[5mzk]], [[5mzi]], [[5mzc]], [[5fn0]] – PfKMO (mutant) + FAD + inhibitor<br />
| + | |
| - | | + | |
| - | *'''Phenol 2-monooxygenase (PMO)'''
| + | |
| - | | + | |
| - | **[[1foh]] – TcPMO + FAD – ''Trichosporon cutaneum''<br />
| + | |
| - | **[[1pn0]] – TcPMO + FAD + phenol<br />
| + | |
| - | | + | |
| - | *'''Phenylalanine 2-monooxygenase'''
| + | |
| - | | + | |
| - | **[[5pah]], [[6pah]]. – FMO catalytic domain + inhibitor – human<br />
| + | |
| - | | + | |
| - | *'''ActVA-Orf6 monooxygenase (AOMO)'''
| + | |
| - | | + | |
| - | **[[1lq9]] – ScAOMO – ''Streptomyces coelicolor''<br />
| + | |
| - | **[[1n5q]] – ScAOMO + sancycline <br />
| + | |
| - | **[[1n5s]], [[1n5t]] – ScAOMO + acetyl dithranol<br />
| + | |
| - | **[[1n5v]] – ScAOMO + nanaomycine <br />
| + | |
| - | | + | |
| - | *'''Flavin-containing monooxygenase'''
| + | |
| - | | + | |
| - | **[[2gv8]] – fyMO + FAD + NADP – fission yeast<br />
| + | |
| - | **[[2gvc]] – fyMO + FAD + NADP + methimazole<br />
| + | |
| - | **[[4a9w]] – SmMO + FAD – ''Stenotrophomonas maltophilia''<br />
| + | |
| - | **[[4c5o]] – SmMO (mutant) + FAD <br />
| + | |
| - | **[[5wan]] – MO + FMN + uracil – Escherichia coli<br />
| + | |
| - | **[[5iq4]], [[5iq1]], [[5ipy]] – RnMO (mutant) + FAD + NAP – ''Roseovarius nubinhibens'' <br />
| + | |
| - | **[[5gsn]] – RnMO (mutant) + FAD + NAP + methimazole <br />
| + | |
| - | **[[4usr]] – PsMO + FAD <br />
| + | |
| - | **[[3rp6]] – KpMO + FAD – ''Klebsiella pneumoniae''<br />
| + | |
| - | **[[3rp8]] – KpMO (mutant) + FAD <br />
| + | |
| - | **[[3rp7]] – KpMO + FAD + uric acid<br />
| + | |
| - | **[[3c96]], [[2rgj]] – PaMO + FAD – ''Pseudomonas aeruginosa''<br />
| + | |
| - | **[[2xvf]], [[2xve]] – MaMO + FAD – ''Methylophaga aminisulfidivorans'' <br />
| + | |
| - | **[[2xvj]] – MaMO (mutant) + FAD + indole <br />
| + | |
| - | **[[2xvi]] – MaMO (mutant) + FAD + O2<br />
| + | |
| - | **[[2vqb]] – MaMO (mutant) + FAD + NADP + O2<br />
| + | |
| - | **[[2xvh]], [[2xlu]], [[2xlt]] – MaMO + FAD + NADP derivative<br />
| + | |
| - | **[[2xls]], [[2xlr]], [[2xlp]], [[2vq7]] – MaMO (mutant) + FAD + NADP <br />
| + | |
| - | | + | |
| - | *Pentachlorophenol 4-monooxygenase
| + | |
| - | | + | |
| - | **[[5kox]] – NfMO + FAD – ''Nocardia farcinica''<br />
| + | |
| - | **[[5kox]] – NfMO + FAD + rifampicin <br />
| + | |
| - | | + | |
| - | *2,4,6-trichlorophenol 4-monooxygenase
| + | |
| - | | + | |
| - | **[[4g5e]] – MO – ''Cupriavidus necator''<br />
| + | |
| - | | + | |
| - | *Chlorophenol 4-monooxygenase
| + | |
| - | | + | |
| - | **[[4oo2]] – MO – ''Streptomyces globisporus''<br />
| + | |
| - | **[[3k86]], [[3hwc]] – BcMO – ''Burkholderia cepacia'' <br />
| + | |
| - | **[[3k87]] – BcMO + FAD <br />
| + | |
| - | **[[3k88]] – BcMO + FAD + NAD <br />
| + | |
| - | | + | |
| - | *Phenylacetone monooxygenase
| + | |
| - | | + | |
| - | **[[4ovi]], [[4c74]] – TfMO + FAD + APADP – ''Thermobifida fusca''<br />
| + | |
| - | **[[1w4x]] – TfMO + FAD<br />
| + | |
| - | **[[2ylt]], [[2yls]], [[2ylr]] – TfMO + FAD + NADP <br />
| + | |
| - | **[[2ylz]] – TfMO (mutant) + FAD <br />
| + | |
| - | **[[4c77]] – TfMO (mutant) + FAD + APADP <br />
| + | |
| - | **[[4d04]] – TfMO (mutant) + FAD + NAP <br />
| + | |
| - | **[[4d03]], [[2ym2]], [[2ylx]], [[2ylw]] – TfMO (mutant) + FAD + NADP <br />
| + | |
| - | **[[2ym1]] – TfMO (mutant) + FAD + NADP + O2<br />
| + | |
| - | | + | |
| - | *6-hydroxynicotinate 3-monooxygenase
| + | |
| - | | + | |
| - | **[[5eow]] – PpMO + FAD – ''Pseudomonas putida''<br />
| + | |
| - | | + | |
| - | *Styrene monooxygenase
| + | |
| - | | + | |
| - | **[[3ihm]] – PpMO <br />
| + | |
| - | **[[4f07]] – PpMO (mutant) + FAD <br />
| + | |
| - | | + | |
| - | *Tryptophan 2-monooxygenase
| + | |
| - | | + | |
| - | **[[4iv9]] – MO + FAD – ''Pseudomonas savastanoi''<br />
| + | |
| - | | + | |
| - | *Tryptophan 5-monooxygenase
| + | |
| - | | + | |
| - | **[[1mlw]] – hMO + Fe + dihydrobiopterin <br />
| + | |
| - | | + | |
| - | *Tyrosine 3-monooxygenase
| + | |
| - | | + | |
| - | **[[2xsn]] – hMO + Zn<br />
| + | |
| - | **[[2toh]] – rMO + Fe + dihydrobiopterin + tyrosine - rat<br />
| + | |
| - | **[[2mda]] – rMO regulatory domain <br />
| + | |
| - | | + | |
| - | *Nitronate monooxygenase
| + | |
| - | | + | |
| - | **[[4q4k]] – PaMO + FMN <br />
| + | |
| - | **[[5lsm]] – MO + FMN – ''Shewanella oneidensis''<br />
| + | |
| - | | + | |
| - | *2-hydroxbiphenyl 3-monooxygenase
| + | |
| - | | + | |
| - | **[[4z2r]] – PnMO + FAD – ''Pseudomonas nitroreducens''<br />
| + | |
| - | **[[4cy6]] – PnMO (mutant) <br />
| + | |
| - | **[[4z2u]], [[4z2t]], [[4cy8]] – PnMO (mutant) + FAD <br />
| + | |
| - | **[[5brt]] – PnMO + FAD + hydroxybephenyl <br />
| + | |
| - | | + | |
| - | *4-hydroxyphenylacetate 3-monooxygenase
| + | |
| - | | + | |
| - | **[[4ira]] – BmMO + FAD – Brucella melitensis<br />
| + | |
| - | **[[3cb0]] – BmMO + FMN<br />
| + | |
| - | | + | |
| - | *Ornithine N(5)-monooxygenase
| + | |
| - | | + | |
| - | **[[5cku]] – MO + FAD + NADP + ornithine – ''Neosartorya fumigata''<br />
| + | |
| - | **[[4nzh]], [[4b69]] – AfMO + FAD + ornithine – ''Aspergillus fumigatus''<br />
| + | |
| - | **[[4b65]] – AfMO + FAD + NADP <br />
| + | |
| - | **[[4b68]], [[4b66]] – AfMO + FAD + NAP + arginine <br />
| + | |
| - | **[[4b67]], [[4b63]] – AfMO + FAD + NADP + ornithine <br />
| + | |
| - | **[[4b64]] – AfMO + FAD + NADP + lysine <br />
| + | |
| - | **[[3s61]] – PaMO + FAD + NADP + ornithine derivative<br />
| + | |
| - | | + | |
| - | *Steroid monooxygenase
| + | |
| - | | + | |
| - | **[[4ap1]], [[4aos]] – RrMO + FAD + NADP – ''Rhodococcus rhodochrous''<br />
| + | |
| - | **[[4aox]] – RrMO (mutant) + FAD <br />
| + | |
| - | **[[4ap3]] – RrMO (mutant) + FAD + NADP <br />
| + | |
| - | | + | |
| - | *Cyclohexanone monooxygenase
| + | |
| - | | + | |
| - | **[[5m10]] – TmMO + FAD + NAP + nicotinamide – ''Thermocripsum municipale''''Italic text''<br />
| + | |
| - | **[[5m0z]] – TmMO + FAD + NADP derivative <br />
| + | |
| - | **[[4rg4]], [[4rg3]], [[3gwf]], [[3gwd]] – RhMO + FAD + NAP + caprolactone – ''Rhodococcus''<br />
| + | |
| - | **[[3ucl]] – RhMO + FAD + NADP + cyclohexanone <br />
| + | |
| - | | + | |
| - | *Lysine 6-monooxygenase
| + | |
| - | | + | |
| - | **[[5cqf]] – MO – ''Pseudomonas syringae''<br />
| + | |
| - | **[[4d7e]] – MO (mutant) + FAD – ''Nocardia farcinica''<br />
| + | |
| - | | + | |
| - | *EDTA monooxygenase
| + | |
| - | | + | |
| - | **[[5dqp]] – MO – ''Chelativorans''<br />
| + | |
| - | | + | |
| - | *3,6-diketocamphane 1,6 monooxygenase
| + | |
| - | | + | |
| - | **[[5aec]] – PpMO <br />
| + | |
| - | **[[4uwm]] – PpMO + FMN<br />
| + | |
| - | | + | |
| - | *Lytic polysaccharide monooxygenase
| + | |
| - | | + | |
| - | **[[5tki]], [[5foh]], [[4qi8]] – NcMO-2 + Cu – ''Neurospora crassa''''Italic text''<br />
| + | |
| - | **[[5tkh]], [[5tkg]], [[5tkf]], [[4eir]] – NcMO-2 + Cu + O2 <br />
| + | |
| - | **[[4eis]] – NcMO-3 + Cu + peroxide <br />
| + | |
| - | **[[5iju]] – MO + Cu – ''Bacillus amyloliquefaciens''<br />
| + | |
| - | **[[5acj]], [[5aci]], [[5ach]], [[5acg]], [[5acf]] – MO + Cu – ''Lentinus similis''<br />
| + | |
| - | **[[4mai]] – moMO + Cu – mold<br />
| + | |
| - | **[[4mah]] – moMO + Zn<br />
| + | |
| - | | + | |
| - | *Peptidyl-glycine alpha-amidating monooxygenase
| + | |
| - | | + | |
| - | **[[3mib]], [[3mic]], [[3mid]], [[3mie]], [[3mif]], [[3mig]], [[3mih]], [[3mlj]], [[3mlk]], [[3mll]], [[1opm]] – rMO + Cu + Ni <br />
| + | |
| - | **[[1sdw]] – rMO + Cu + Ni + O2 + threonine derivative<br />
| + | |
| - | **[[1yjl]] – rMO (mutant)<br />
| + | |
| - | **[[1yjk]], [[1yip]], [[1phm]] – rMO + Cu<br />
| + | |
| - | **[[1yi9]] – rMO (mutant) + Cu<br />
| + | |
| - | **[[3fw0]] – rMO + Hg<br />
| + | |
| - | **[[3fvz]] – rMO + Zn + Fe<br />
| + | |
| - | | + | |
| - | *Antibiotic biosynthesis monooxygenase
| + | |
| - | | + | |
| - | **[[4hl9]] – MO – ''Rhodospirillum rubrum''<br />
| + | |
| - | **[[4dn9]] – MO – ''Chloroflexus aurantiacus''<br />
| + | |
| - | **[[2ril]] – MO – ''Shewanella loihica''<br />
| + | |
| - | | + | |
| - | *Monooxygenase
| + | |
| - | | + | |
| - | **[[2i7g]] – MO – ''Agrobacterium tumefaciens''<br />
| + | |
| - | **[[5uq4]] – MtMO <br />
| + | |
| - | **[[5f5l]] – MiMO – ''Micromonospora'' <br />
| + | |
| - | **[[5f5n]] – MiMO + NAD + substrate <br />
| + | |
| - | | + | |
| - | '''Methane monooxygenase''' See [[Methane monooxygenase]]
| + | |
| - | | + | |
| - | '''Camphor 5-monooxygenase''' See [[Cytochrome P450]]
| + | |
| - | | + | |
| - | '''Luciferin 4-monooxygenase and Alkanal monooxygenase''' See [[Luciferase]]
| + | |
| - | }}
| + | |
| | | | |
| | == References == | | == References == |