This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1cqj
From Proteopedia
(Difference between revisions)
| (2 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
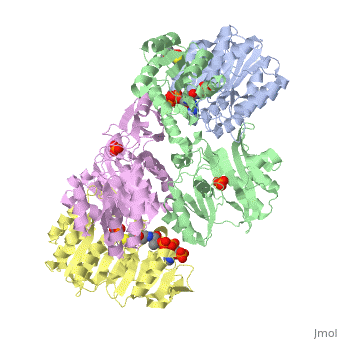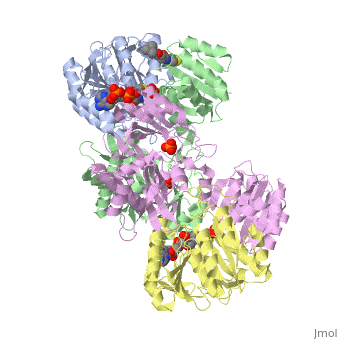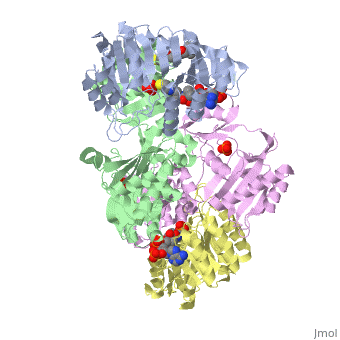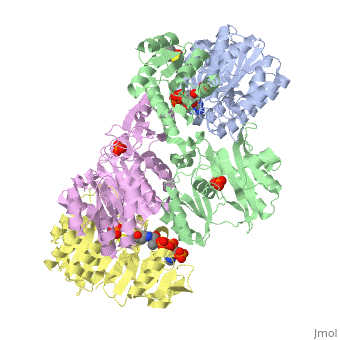
==CRYSTAL STRUCTURE OF DEPHOSPHORYLATED E. COLI SUCCINYL-COA SYNTHETASE== | ==CRYSTAL STRUCTURE OF DEPHOSPHORYLATED E. COLI SUCCINYL-COA SYNTHETASE== | ||
| - | <StructureSection load='1cqj' size='340' side='right' caption='[[1cqj]], [[Resolution|resolution]] 2.90Å' scene=''> | + | <StructureSection load='1cqj' size='340' side='right'caption='[[1cqj]], [[Resolution|resolution]] 2.90Å' scene=''> |
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1cqj]] is a 4 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1cqj]] is a 4 chain structure with sequence from [https://en.wikipedia.org/wiki/Escherichia_coli Escherichia coli]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1CQJ OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1CQJ FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2.9Å</td></tr> |
| - | <tr id=' | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=COA:COENZYME+A'>COA</scene>, <scene name='pdbligand=PO4:PHOSPHATE+ION'>PO4</scene></td></tr> |
| - | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1cqj FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1cqj OCA], [https://pdbe.org/1cqj PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1cqj RCSB], [https://www.ebi.ac.uk/pdbsum/1cqj PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1cqj ProSAT]</span></td></tr> | |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | |
</table> | </table> | ||
== Function == | == Function == | ||
| - | [ | + | [https://www.uniprot.org/uniprot/SUCD_ECOLI SUCD_ECOLI] During aerobic metabolism it functions in the citric acid cycle, coupling the hydrolysis of succinyl-CoA to the synthesis of ATP and thus represents an important site of substrate-level phosphorylation. It can also function in the other direction for anabolic purposes, and this may be particularly important for providing succinyl-CoA during anaerobic growth when the oxidative route from 2-oxoglutarate is severely repressed. The alpha-subunit binds CoA, as well as ATP and catalyzes phosphoryl transfer to one of its histidine residues. The complete active site is probably located in the region of alpha-beta contact. |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 21: | Line 20: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1cqj ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1cqj ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
| - | Succinyl-CoA synthetase (SCS) catalyzes the following reversible reaction via a phosphorylated histidine intermediate (His 246alpha): succinyl-CoA + P(i) + NDP <--> succinate + CoA + NTP (N denotes adenosine or guanosine). To determine the structure of the enzyme with nucleotide bound, crystals of phosphorylated Escherichia coli SCS were soaked in successive experiments adopting progressive strategies. In the first experiment, 1 mM ADP (>15 x K(d)) was added; Mg(2+) ions were omitted to preclude the formation of an insoluble precipitate with the phosphate and ammonium ions. X-ray crystallography revealed that the enzyme was dephosphorylated, but the nucleotide did not remain bound to the enzyme (R(working) = 17.2%, R(free) = 22.8% for data to 2.9 A resolution). Catalysis requires Mg(2+) ions; hence, the "true" nucleotide substrate is probably an ADP-Mg(2+) complex. In the successful experiment, the phosphate buffer was exchanged with MOPS, the concentration of sulfate ions was lowered, and the concentrations of ADP and Mg(2+) ions were increased to 10.5 and 50 mM, respectively. X-ray diffraction data revealed an ADP-Mg(2+) complex bound in the ATP-grasp fold of the N-terminal domain of each beta-subunit (R(working) = 19.1%, R(free) = 24.7% for data to 3.3 A resolution). We describe the specific interactions of the nucleotide-Mg(2+) complex with SCS, compare these results with those for other proteins containing the ATP-grasp fold, and present a hypothetical model of the histidine-containing loop in the "down" position where it can interact with the nucleotide approximately 35 A from where His 246alpha is seen in both phosphorylated and dephosphorylated SCS. | ||
| - | + | ==See Also== | |
| - | + | *[[Succinyl-CoA synthetase|Succinyl-CoA synthetase]] | |
| - | + | *[[Succinyl-CoA synthetase 3D structures|Succinyl-CoA synthetase 3D structures]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| - | [[Category: | + | [[Category: Escherichia coli]] |
| - | [[Category: | + | [[Category: Large Structures]] |
| - | [[Category: | + | [[Category: Bridger WA]] |
| - | [[Category: | + | [[Category: Fraser ME]] |
| - | [[Category: | + | [[Category: James MNG]] |
| - | [[Category: | + | [[Category: Joyce MA]] |
| - | [[Category: | + | [[Category: Wolodko WT]] |
| - | + | ||
| - | + | ||
Current revision
CRYSTAL STRUCTURE OF DEPHOSPHORYLATED E. COLI SUCCINYL-COA SYNTHETASE
| |||||||||||