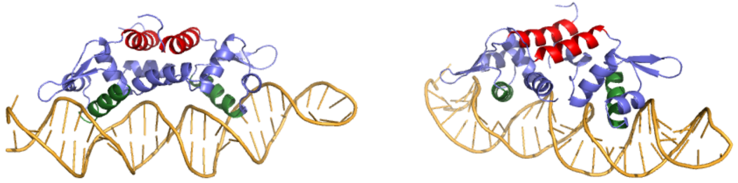This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1053
From Proteopedia
(Difference between revisions)
| Line 23: | Line 23: | ||
== Zinc Binding Site== | == Zinc Binding Site== | ||
| - | Many zinc-dependent proteins are transcriptional regulators<ref>DOI: 10.1128/MMBR.00015-06</ref>. Czr A fits into this category as an [https://en.wikipedia.org/wiki/Allosteric_regulation allosteric inhibitor] of the czr operon. Two [https://en.wikipedia.org/wiki/Zinc Zn<sup> +2</sup>] ions may bind to the dimer<ref name="critical"/>, at the location of the <scene name='69/694220/A5_helices__zn_binding/2'>α5 helix</scene> from each monomer. As zinc binds, the α5 helices <scene name='69/694218/2kjc_zinc_bound/1'>unalign</scene> to inhibit the DNA binding residues (Figure 2). Furthermore, CzrA must be in its dimer form for zinc to bind. The <scene name='69/694220/Spacefill_zinc_pockets/1'>zinc binding pockets</scene> are formed by two residues from each monomer, so Zn<sup>+2</sup> cannot bind to the monomer. The <scene name='69/694220/Zinc_binding_residues/7'>zinc binding site</scene> is formed by Asp84 and His | + | Many zinc-dependent proteins are transcriptional regulators<ref>DOI: 10.1128/MMBR.00015-06</ref>. Czr A fits into this category as an [https://en.wikipedia.org/wiki/Allosteric_regulation allosteric inhibitor] of the czr operon. Two [https://en.wikipedia.org/wiki/Zinc Zn<sup> +2</sup>] ions may bind to the dimer<ref name="critical"/>, at the location of the <scene name='69/694220/A5_helices__zn_binding/2'>α5 helix</scene> from each monomer. As zinc binds, the α5 helices <scene name='69/694218/2kjc_zinc_bound/1'>unalign</scene> to inhibit the DNA binding residues (Figure 2). Furthermore, CzrA must be in its dimer form for zinc to bind. The <scene name='69/694220/Spacefill_zinc_pockets/1'>zinc binding pockets</scene> are formed by two residues from each monomer, so Zn<sup>+2</sup> cannot bind to the monomer. The <scene name='69/694220/Zinc_binding_residues/7'>zinc binding site</scene> is formed by Asp84 and His 86 from one monomer, as well as His97 and His100 from the other monomer. Zinc ions were not present in the solution NMR structure<ref name="critical"/>, so a representation of a zinc ion in the binding pocket has been drawn in Figure 4. The large number of histidines used in the Czr A zinc pocket is a repetitive and commonly found feature in zinc-binding proteins <ref>Miller J, McLachlan AD, Klug A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985 Jun 4;4(6):1609-1614.</ref>. |
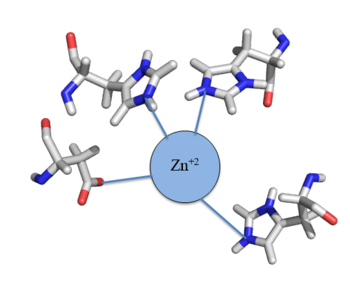
[[Image:Zinc tetrahedral complex.PNG|350px|thumb|center| Figure 4: Zn<sup>+2</sup> tetrahedral binding complex]] | [[Image:Zinc tetrahedral complex.PNG|350px|thumb|center| Figure 4: Zn<sup>+2</sup> tetrahedral binding complex]] | ||
| - | Zn<sup>+2</sup> binding is driven by a large [https://en.wikipedia.org/wiki/Entropy entropic] gain <ref>DOI:10.1021/ja906131b</ref>. Water molecules around the metal ion and Czr A protein are displaced, and gain greater freedom. This gain in entropy allows Zn<sup>+2</sup> to bind to Czr A with reasonable affinity and speed in vivo. The zinc<sup>+2</sup> ion forms a tetrahedral complex with four residues (Figure 4). Other metal ions that may form a tetrahedral complex will have some affinity for Czr A; however, the metal binding pocket of Czr A has been optimized to bind Zn<sup>+2</sup> with the highest affinity. | + | Zn<sup>+2</sup> binding is driven by a large [https://en.wikipedia.org/wiki/Entropy entropic] gain <ref>DOI:10.1021/ja906131b</ref>. Water molecules around the metal ion and Czr A protein are displaced, and gain greater freedom. This gain in entropy allows Zn<sup>+2</sup> to bind to Czr A with reasonable affinity and speed in vivo. The zinc<sup>+2</sup> ion forms a tetrahedral complex with the four residues (Figure 4). Other metal ions that may form a tetrahedral complex will have some affinity for Czr A; however, the metal binding pocket of Czr A has been optimized to bind Zn<sup>+2</sup> with the highest affinity. |
</StructureSection> | </StructureSection> | ||
| Line 34: | Line 34: | ||
== References == | == References == | ||
<references/> | <references/> | ||
| + | |||
| + | |||
| + | ==Student Contributors== | ||
| + | *Katelyn Baumer | ||
| + | *Jakob Jozwiakowski | ||
| + | *Catie Liggett | ||
Current revision
Zinc Dependent Transcriptional Repressor of the Czr operon (CzrA)
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Arunkumar A., Campanello G., Giedroc D. (2009). Solution Structure of a paradigm ArsR family zinc sensor in the DNA-bound state. PNAS 106:43 18177-18182.
- ↑ Chakravorty DK, Wang B, Lee CW, Giedroc DP, Merz KM Jr. Simulations of allosteric motions in the zinc sensor CzrA. J Am Chem Soc. 2012 Feb 22;134(7):3367-76. doi: 10.1021/ja208047b. Epub 2011 Nov , 14. PMID:22007899 doi:http://dx.doi.org/10.1021/ja208047b
- ↑ MacPherson S, Larochelle M, Turcotte B. A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiol Mol Biol Rev. 2006 Sep;70(3):583-604. PMID:16959962 doi:http://dx.doi.org/10.1128/MMBR.00015-06
- ↑ Miller J, McLachlan AD, Klug A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985 Jun 4;4(6):1609-1614.
- ↑ Grossoehme NE, Giedroc DP. Energetics of allosteric negative coupling in the zinc sensor S. aureus CzrA. J Am Chem Soc. 2009 Dec 16;131(49):17860-70. doi: 10.1021/ja906131b. PMID:19995076 doi:http://dx.doi.org/10.1021/ja906131b
Student Contributors
- Katelyn Baumer
- Jakob Jozwiakowski
- Catie Liggett