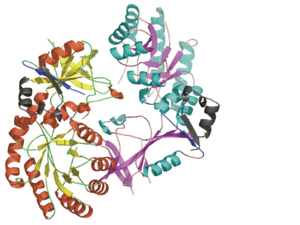
Antizyme Inhibitor
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 12: | Line 12: | ||
<scene name='Antizyme_Inhibitor/Binding_site/6'>Overlap</scene> of the AzI and mODC structures suggests that AzI does not bind [http://en.wikipedia.org/wiki/Pyridoxal_phosphate PLP]. <font color='black'><b>PLP</b></font> is in <font color='black'><b>yellow</b></font>, <font color='lime'><b>ODC residues D88, R154, R277, and Y389 are in lime</b></font>, and the corresponding <font color='magenta'><b>AzI residues A88, H154, S274, and D387 are in magenta</b></font>. Many of the residues participating in PLP-ODC binding are not conserved in AzI. These include D88A, R154H, R277S, D332E, and Y389D (ODC residue numbers follow the sequence of AzI). Notably, the absence of even one of these interactions, (''e.g.'' ODC R277A mutant) results in a 100-fold decrease in PLP binding, a 50% drop in K<sub>cat</sub>, and a 7-fold decrease in K<sub>M</sub>. | <scene name='Antizyme_Inhibitor/Binding_site/6'>Overlap</scene> of the AzI and mODC structures suggests that AzI does not bind [http://en.wikipedia.org/wiki/Pyridoxal_phosphate PLP]. <font color='black'><b>PLP</b></font> is in <font color='black'><b>yellow</b></font>, <font color='lime'><b>ODC residues D88, R154, R277, and Y389 are in lime</b></font>, and the corresponding <font color='magenta'><b>AzI residues A88, H154, S274, and D387 are in magenta</b></font>. Many of the residues participating in PLP-ODC binding are not conserved in AzI. These include D88A, R154H, R277S, D332E, and Y389D (ODC residue numbers follow the sequence of AzI). Notably, the absence of even one of these interactions, (''e.g.'' ODC R277A mutant) results in a 100-fold decrease in PLP binding, a 50% drop in K<sub>cat</sub>, and a 7-fold decrease in K<sub>M</sub>. | ||
| - | </StructureSection> | ||
==3D structures of Antizyme Inhibitor== | ==3D structures of Antizyme Inhibitor== | ||
| + | [[Antizyme inhibitor 3D structures]] | ||
| - | + | </StructureSection> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
==Additional Resources== | ==Additional Resources== | ||
Current revision
| |||||||||||
Additional Resources
For additional information, see: Cancer
Reference
Shira Albeck, Orly Dym, Tamar Unger, Zohar Snapir, Zippy Bercovich and Chaim Kahana. Crystallographic and biochemical studies revealing the structural basis for antizyme inhibitor function. Protein Sci. 2008 May; 17(5): 793-802. Epub 2008 Mar 27.
Proteopedia Page Contributors and Editors (what is this?)
Alexander Berchansky, Michal Harel, Joel L. Sussman, Dinesh Kulhary, David Canner, Jaime Prilusky, Orly Dym