This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Amylase
From Proteopedia
(Difference between revisions)
| (2 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
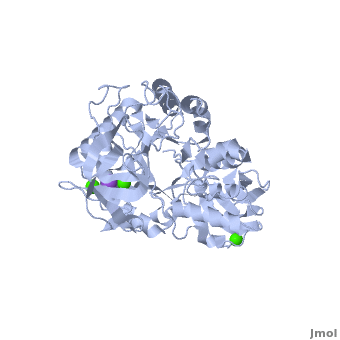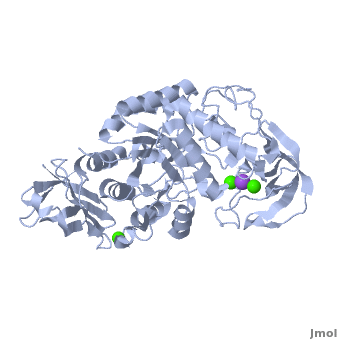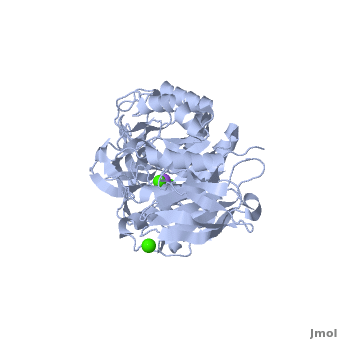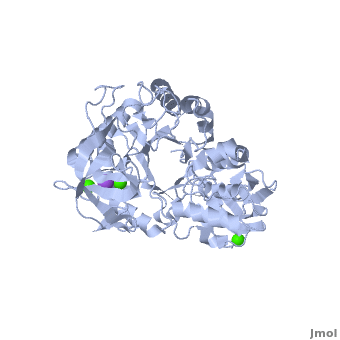
<StructureSection load='' size='350' side='right' scene='Sandbox_182/Alpha-amylase/1' caption='Amylase complex with Ca+2 (green) and Na+ (purple) ions (PDB code [[1hvx]])'> | <StructureSection load='' size='350' side='right' scene='Sandbox_182/Alpha-amylase/1' caption='Amylase complex with Ca+2 (green) and Na+ (purple) ions (PDB code [[1hvx]])'> | ||
=Introduction= | =Introduction= | ||
| - | Discovered and isolated by [http://en.wikipedia.org/wiki/Anselme_Payen Anselme Payen] in 1833, amylase was the first enzyme to be discovered<ref name="book">Yamamoto T.1988. Handbook of Amylases and Related Enzymes: Their Sources, Isolation Methods, Properties and Applications. Osaka Japan: Pergamon Press</ref>. Amylases are hydrolases, acting on α-1,4-glycosidic bonds<ref name="Path">PMID:9541387</ref>. They can be further subdivided into α,β and γ amylases<ref name="book"/>.'''α-Amylase''' (AAM) is an enzyme that acts as a catalyst for the hydrolysis of α-linked polysaccharides into α-anomeric products<ref name="Main">PMID:11226887</ref>. The enzyme can be derived from a variety of sources, each with different characteristics. α-Amylase found within the human body serves as the enzyme active in pancreatic juice and saliva<ref name="Path"/>. α-Amylase is not only essential in human physiology but has a number of important biotechnological functions in various processing industries. '''β/α amylase''' (BAAM) is a precursor protein which is cleaved to form the β-amylase and α-amylase after secretion. '''β amylase''' (BAM) acts at the non-reducing chain ends and liberate only β-maltose<ref>PMID:6168260</ref>. '''γ amylase''' (GAM) acts at the non-reducing chain ends of amylose and amylopectin and liberates glucose. '''Pullulanase''' hydrolyses the α-1,6 glucoside linkage in starch, amylopectin, pullulan and related oligosaccharides<ref>PMID:22991654</ref>.<br /> | + | Discovered and isolated by [http://en.wikipedia.org/wiki/Anselme_Payen Anselme Payen] in 1833, '''amylase''' was the first enzyme to be discovered<ref name="book">Yamamoto T.1988. Handbook of Amylases and Related Enzymes: Their Sources, Isolation Methods, Properties and Applications. Osaka Japan: Pergamon Press</ref>. Amylases are hydrolases, acting on α-1,4-glycosidic bonds<ref name="Path">PMID:9541387</ref>. They can be further subdivided into α,β and γ amylases<ref name="book"/>.'''α-Amylase''' (AAM) is an enzyme that acts as a catalyst for the hydrolysis of α-linked polysaccharides into α-anomeric products<ref name="Main">PMID:11226887</ref>. The enzyme can be derived from a variety of sources, each with different characteristics. α-Amylase found within the human body serves as the enzyme active in pancreatic juice and saliva<ref name="Path"/>. α-Amylase is not only essential in human physiology but has a number of important biotechnological functions in various processing industries. '''β/α amylase''' (BAAM) is a precursor protein which is cleaved to form the β-amylase and α-amylase after secretion. '''β amylase''' (BAM) acts at the non-reducing chain ends and liberate only β-maltose<ref>PMID:6168260</ref>. '''γ amylase''' (GAM) acts at the non-reducing chain ends of amylose and amylopectin and liberates glucose. '''Pullulanase''' hydrolyses the α-1,6 glucoside linkage in starch, amylopectin, pullulan and related oligosaccharides<ref>PMID:22991654</ref>.<br /> |
| + | *'''Neopullulanase''' is involved in starch degrading<ref>PMID:8955399</ref>.<br /> | ||
For α-amylase see [[Raghad zoubi]]<br /> | For α-amylase see [[Raghad zoubi]]<br /> | ||
See also [[Amylase (Hebrew)]]. | See also [[Amylase (Hebrew)]]. | ||
| Line 25: | Line 26: | ||
β/α amylase (BAAM) is a precursor protein which is cleaved to form the β-amylase and α-amylase after secretion. | β/α amylase (BAAM) is a precursor protein which is cleaved to form the β-amylase and α-amylase after secretion. | ||
| - | = Structure of the AmyC GH13 alpha-amylase from Alicyclobacillus sp, reveals accommodation of starch branching points in the alpha-amylase family | + | = Structure of the AmyC GH13 alpha-amylase from Alicyclobacillus sp, reveals accommodation of starch branching points in the alpha-amylase family<ref>doi 10.1107/S2059798318014900</ref> = |
| - | + | ||
| - | + | ||
| - | + | ||
The enzymatic degradation of starch has a myriad industrial applications. However, the branched nature of the polysaccharides that compose it poses problems, as branches have to be accommodated within an active centre best suited to linear polysaccharides. Alpha-amylases are glycoside hydrolases that break the α-1,4 bonds in starch and related glycans. The present work provides a rare insight into branch-point acceptance in these industrial catalysts. | The enzymatic degradation of starch has a myriad industrial applications. However, the branched nature of the polysaccharides that compose it poses problems, as branches have to be accommodated within an active centre best suited to linear polysaccharides. Alpha-amylases are glycoside hydrolases that break the α-1,4 bonds in starch and related glycans. The present work provides a rare insight into branch-point acceptance in these industrial catalysts. | ||
Current revision
| |||||||||||
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Yamamoto T.1988. Handbook of Amylases and Related Enzymes: Their Sources, Isolation Methods, Properties and Applications. Osaka Japan: Pergamon Press
- ↑ 2.0 2.1 Aghajari N, Feller G, Gerday C, Haser R. Crystal structures of the psychrophilic alpha-amylase from Alteromonas haloplanctis in its native form and complexed with an inhibitor. Protein Sci. 1998 Mar;7(3):564-72. PMID:9541387
- ↑ 3.0 3.1 3.2 Suvd D, Fujimoto Z, Takase K, Matsumura M, Mizuno H. Crystal structure of Bacillus stearothermophilus alpha-amylase: possible factors determining the thermostability. J Biochem. 2001 Mar;129(3):461-8. PMID:11226887
- ↑ French D. Amylases: enzymatic mechanisms. Basic Life Sci. 1981;18:151-82. PMID:6168260
- ↑ Hii SL, Tan JS, Ling TC, Ariff AB. Pullulanase: role in starch hydrolysis and potential industrial applications. Enzyme Res. 2012;2012:921362. doi: 10.1155/2012/921362. Epub 2012 Sep 6. PMID:22991654 doi:http://dx.doi.org/10.1155/2012/921362
- ↑ D'Elia JN, Salyers AA. Contribution of a neopullulanase, a pullulanase, and an alpha-glucosidase to growth of Bacteroides thetaiotaomicron on starch. J Bacteriol. 1996 Dec;178(24):7173-9. PMID:8955399
- ↑ 7.0 7.1 7.2 Aghajari N, Feller G, Gerday C, Haser R. Structural basis of alpha-amylase activation by chloride. Protein Sci. 2002 Jun;11(6):1435-41. PMID:12021442
- ↑ Maurus R, Begum A, Williams LK, Fredriksen JR, Zhang R, Withers SG, Brayer GD. Alternative catalytic anions differentially modulate human alpha-amylase activity and specificity(,). Biochemistry. 2008 Mar 18;47(11):3332-44. Epub 2008 Feb 20. PMID:18284212 doi:10.1021/bi701652t
- ↑ 9.0 9.1 Maurus R, Begum A, Williams LK, Fredriksen JR, Zhang R, Withers SG, Brayer GD. Alternative catalytic anions differentially modulate human alpha-amylase activity and specificity(,). Biochemistry. 2008 Mar 18;47(11):3332-44. Epub 2008 Feb 20. PMID:18284212 doi:10.1021/bi701652t
- ↑ 10.0 10.1 10.2 10.3 Kuriki T, Imanaka T. The concept of the alpha-amylase family: structural similarity and common catalytic mechanism. J Biosci Bioeng. 1999;87(5):557-65. PMID:16232518
- ↑ 11.0 11.1 PPMID: 17713601
- ↑ Franco OL, Rigden DJ, Melo FR, Grossi-De-Sa MF. Plant alpha-amylase inhibitors and their interaction with insect alpha-amylases. Eur J Biochem. 2002 Jan;269(2):397-412. PMID:11856298
- ↑ Yang RW, Shao ZX, Chen YY, Yin Z, Wang WJ. Lipase and pancreatic amylase activities in diagnosis of acute pancreatitis in patients with hyperamylasemia. Hepatobiliary Pancreat Dis Int. 2005 Nov;4(4):600-3. PMID:16286272
- ↑ Agirre J, Moroz O, Meier S, Brask J, Munch A, Hoff T, Andersen C, Wilson KS, Davies GJ. The structure of the AliC GH13 alpha-amylase from Alicyclobacillus sp. reveals the accommodation of starch branching points in the alpha-amylase family. Acta Crystallogr D Struct Biol. 2019 Jan 1;75(Pt 1):1-7. doi:, 10.1107/S2059798318014900. Epub 2019 Jan 4. PMID:30644839 doi:http://dx.doi.org/10.1107/S2059798318014900
- ↑ Machius M, Declerck N, Huber R, Wiegand G. Activation of Bacillus licheniformis alpha-amylase through a disorder-->order transition of the substrate-binding site mediated by a calcium-sodium-calcium metal triad. Structure. 1998 Mar 15;6(3):281-92. PMID:9551551
- ↑ Brzozowski AM, Lawson DM, Turkenburg JP, Bisgaard-Frantzen H, Svendsen A, Borchert TV, Dauter Z, Wilson KS, Davies GJ. Structural analysis of a chimeric bacterial alpha-amylase. High-resolution analysis of native and ligand complexes. Biochemistry. 2000 Aug 8;39(31):9099-107. PMID:10924103
Proteopedia Page Contributors and Editors (what is this?)
Shane Riley, Michal Harel, Joel L. Sussman, Randi Woodbeck, Jaime Prilusky, Alexander Berchansky, Ann Taylor, Andrea Gorrell, David Canner