This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1dmg
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 3: | Line 3: | ||
<StructureSection load='1dmg' size='340' side='right'caption='[[1dmg]], [[Resolution|resolution]] 1.70Å' scene=''> | <StructureSection load='1dmg' size='340' side='right'caption='[[1dmg]], [[Resolution|resolution]] 1.70Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1dmg]] is a 1 chain structure. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1DMG OCA]. For a <b>guided tour on the structure components</b> use [ | + | <table><tr><td colspan='2'>[[1dmg]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Thermotoga_maritima Thermotoga maritima]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1DMG OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1DMG FirstGlance]. <br> |
| - | </td></tr><tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat"><scene name='pdbligand=CIT:CITRIC+ACID'>CIT</scene></td></tr> | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 1.7Å</td></tr> |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=CIT:CITRIC+ACID'>CIT</scene></td></tr> |
| + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1dmg FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1dmg OCA], [https://pdbe.org/1dmg PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1dmg RCSB], [https://www.ebi.ac.uk/pdbsum/1dmg PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1dmg ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
| - | [ | + | [https://www.uniprot.org/uniprot/RL4_THEMA RL4_THEMA] One of the primary rRNA binding proteins, this protein initially binds near the 5'-end of the 23S rRNA. It is important during the early stages of 50S assembly. It makes multiple contacts with different domains of the 23S rRNA in the assembled 50S subunit and ribosome (By similarity).[HAMAP-Rule:MF_01328_B] This protein only weakly controls expression of the E.coli S10 operon. It is incorporated into E.coli ribosomes, however it is not as firmly associated as the endogenous protein.[HAMAP-Rule:MF_01328_B] Forms part of the polypeptide exit tunnel (By similarity).[HAMAP-Rule:MF_01328_B] |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 19: | Line 20: | ||
</jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1dmg ConSurf]. | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/main_output.php?pdb_ID=1dmg ConSurf]. | ||
<div style="clear:both"></div> | <div style="clear:both"></div> | ||
| - | <div style="background-color:#fffaf0;"> | ||
| - | == Publication Abstract from PubMed == | ||
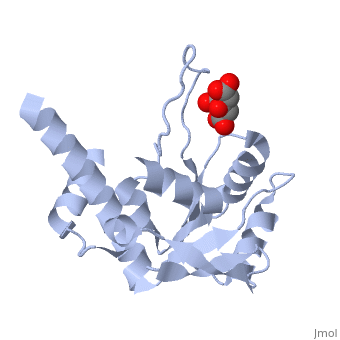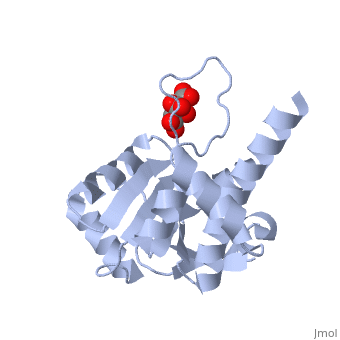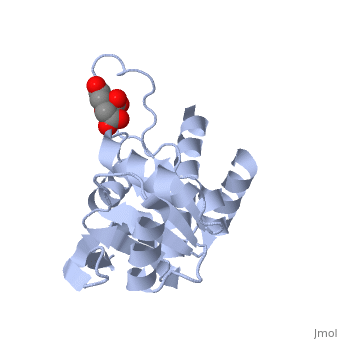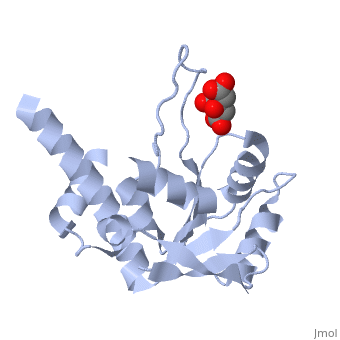
| - | Ribosomal protein L4 resides near the peptidyl transferase center of the bacterial ribosome and may, together with rRNA and proteins L2 and L3, actively participate in the catalysis of peptide bond formation. Escherichia coli L4 is also an autogenous feedback regulator of transcription and translation of the 11 gene S10 operon. The crystal structure of L4 from Thermotoga maritima at 1.7 A resolution shows the protein with an alternating alpha/beta fold and a large disordered loop region. Two separate binding sites for RNA are discernible. The N-terminal site, responsible for binding to rRNA, consists of the disordered loop with flanking alpha-helices. The C-terminal site, a prime candidate for the interaction with the leader sequence of the S10 mRNA, involves two non-consecutive alpha-helices. The structure also suggests a C-terminal protein-binding interface, through which L4 could be interacting with protein components of the transcriptional and/or translational machineries. | ||
| - | |||
| - | Crystal structure of ribosomal protein L4 shows RNA-binding sites for ribosome incorporation and feedback control of the S10 operon.,Worbs M, Huber R, Wahl MC EMBO J. 2000 Mar 1;19(5):807-18. PMID:10698923<ref>PMID:10698923</ref> | ||
| - | |||
| - | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | ||
| - | </div> | ||
| - | <div class="pdbe-citations 1dmg" style="background-color:#fffaf0;"></div> | ||
==See Also== | ==See Also== | ||
*[[Ribosomal protein L4|Ribosomal protein L4]] | *[[Ribosomal protein L4|Ribosomal protein L4]] | ||
| - | == References == | ||
| - | <references/> | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Thermotoga maritima]] |
| - | [[Category: | + | [[Category: Huber R]] |
| - | [[Category: | + | [[Category: Wahl MC]] |
| - | [[Category: | + | [[Category: Worbs M]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
CRYSTAL STRUCTURE OF RIBOSOMAL PROTEIN L4
| |||||||||||