|
|
| (2 intermediate revisions not shown.) |
| Line 7: |
Line 7: |
| | == Structural highlights == | | == Structural highlights == |
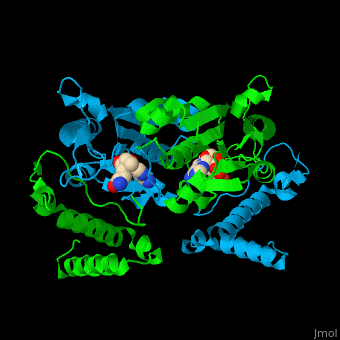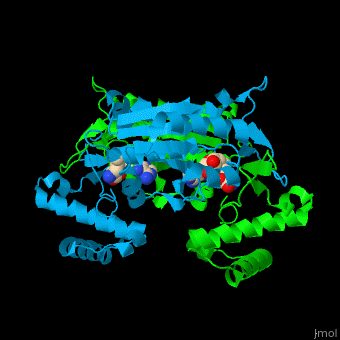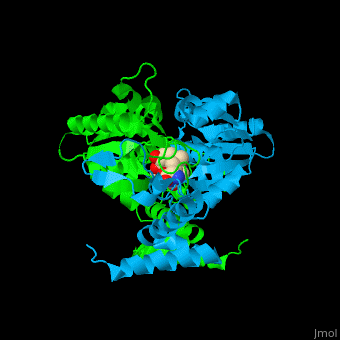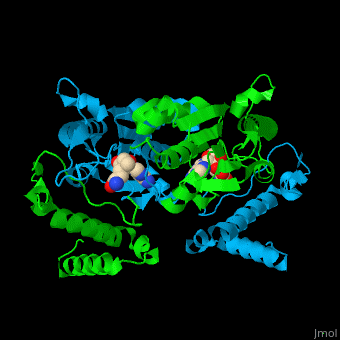
| | Trm <scene name='55/553109/Cv/4'>active site</scene> is located at the cleft at the subunit interface<ref>PMID:12773376</ref>. Water molecules are shown as red spheres. | | Trm <scene name='55/553109/Cv/4'>active site</scene> is located at the cleft at the subunit interface<ref>PMID:12773376</ref>. Water molecules are shown as red spheres. |
| - | </StructureSection> | |
| - | | |
| | ==3D structures of tRNA methyltransferase== | | ==3D structures of tRNA methyltransferase== |
| | + | [[tRNA methyltransferase 3D structures]] |
| | | | |
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}}
| + | </StructureSection> |
| - | {{#tree:id=OrganizedByTopic|openlevels=0|
| + | |
| - | | + | |
| - | *'''tRNA guanine 9 N1 methyltransferase'''
| + | |
| - | | + | |
| - | **[[4jwg]] – fyTrm – fission yeast<br />
| + | |
| - | **[[4jwf]], [[4jwh]] – fyTrm + SAH <br />
| + | |
| - | **[[4jwj]] – yTrm + SAH – yeast<br />
| + | |
| - | | + | |
| - | *'''tRNA guanine 37 N1 methyltransferase'''
| + | |
| - | | + | |
| - | **[[3knu]] – ApTrm – ''Anaplasma phagocytophilum'' <br />
| + | |
| - | **[[1oy5]] – AaTrm – ''Aquifex aeolicus'' <br />
| + | |
| - | **[[3ief]] – Trm – ''Bartonella henselae'' <br />
| + | |
| - | **[[1uaj]] – HiTrm – ''haemophilus influenzae''<br />
| + | |
| - | **[[3ky7]] – Trm – ''Staphylococcus aureus''<br />
| + | |
| - | **[[3quv]] – Trm – ''Mycobacterium abscessus''<br />
| + | |
| - | **[[5hjj]], [[5yac]] – PaTrm – ''Pyrococcus abyssi''<br />
| + | |
| - | | + | |
| - | *Complexes of tRNA guanine 37 N1 methyltransferase
| + | |
| - | | + | |
| - | **[[1uak]], [[4yvg]] – HiTrm + SAM <br />
| + | |
| - | **[[1ual]], [[1uam]] – HiTrm + SAH <br />
| + | |
| - | **[[3axz]] – HiTrm + adenosine<br />
| + | |
| - | **[[4mcb]], [[4mcc]], [[4mcd]] – HiTrm + inhibitor<br />
| + | |
| - | **[[4yvh]] – HiTrm + SFG <br />
| + | |
| - | **[[4yvi]], [[4yvj]], [[4yvk]] – HiTrm + SFG + tRNA<br />
| + | |
| - | **[[5d9f]], [[4ypw]], [[4ypx]], [[4ypy]], [[4ypz]], [[4yq0]], [[4yq1]], [[4yq2]], [[4yq3]], [[4yq4]], [[4yq5]], [[4yq6]], [[4yq7]], [[4yq8]], [[4yq9]], [[4yqa]], [[4yqb]], [[4yqc]], [[4yqg]], [[4yqi]], [[4yqj]], [[4yqk]], [[4yql]], [[4yqn]], [[4yqo]], [[4yqp]], [[4yqq]], [[4yqr]], [[4yqs]], [[4yqt]], [[4yqd]] – HiTrm + SAM competitive<br />
| + | |
| - | **[[1p9p]] – EcTrm + SAH – ''Escherichia coli'' <br />
| + | |
| - | **[[2yx1]] – MjTrm + SFG – ''Methanocalcococcus jannaschii''<br />
| + | |
| - | **[[3ay0]] – MjTrm + adenosine<br />
| + | |
| - | **[[4h3y]], [[4h3z]] – Trm + SAH – ''Burkholderia phymatum'' <br />
| + | |
| - | **[[4ig6]] – ApTrm + SAH <br />
| + | |
| - | **[[4gek]], [[4iwn]] – EcTrm + SAM derivative<br />
| + | |
| - | **[[5hr7]] – EcTrm + tRNA-Glu<br />
| + | |
| - | **[[5hji]], [[5hjm]] – PaTrm + adenosine derivative<br />
| + | |
| - | **[[5hjk]] – PaTrm + SAH <br />
| + | |
| - | | + | |
| - | *'''tRNA guanine 46 N7 methyltransferase'''
| + | |
| - | | + | |
| - | **[[1yzh]] – Trm – ''Streptococcus pneumoniae'' <br />
| + | |
| - | **[[2fca]] – Trm – ''Bacillus subtilis'' <br />
| + | |
| - | **[[2vdu]], [[2vdv]] – yTrm <br />
| + | |
| - | **[[3dxx]] – EcTrm <br />
| + | |
| - | | + | |
| - | *Complexes of tRNA guanine 46 N7 methyltransferase
| + | |
| - | | + | |
| - | **[[3ckk]] – hTrm + SAM - human<br />
| + | |
| - | **[[3dxy]] – EcTrm + SAM <br />
| + | |
| - | **[[3dxz]] – EcTrm + SAH <br />
| + | |
| - | | + | |
| - | *Trm to O2 of cytidine 32
| + | |
| - | | + | |
| - | **[[4cnd]] – EcTrm (mutant)<br />
| + | |
| - | **[[4xbo]] – EcTrm + SAH<br />
| + | |
| - | **[[4cne]] – EcTrm (mutant) + SAH<br />
| + | |
| - | | + | |
| - | *'''tRNA cytidine 34 O2 methyltransferase'''
| + | |
| - | | + | |
| - | **[[3e5y]] – BpTrm – ''Burkholderia pseudomalllei'' <br />
| + | |
| - | **[[4jak]], [[4kdz]] – EcTrm <br />
| + | |
| - | | + | |
| - | *Complexes of tRNA cytidine 34 O2 methyltransferase
| + | |
| - | | + | |
| - | **[[4kgn]] – BpTrm + SAH <br />
| + | |
| - | **[[3n4k]], [[3n4j]] – Trm + SAH – ''Yersinia pestis'' <br />
| + | |
| - | **[[1mxi]], [[1im8]] – HiTrm + SAH <br />
| + | |
| - | **[[4jal]] – EcTrm + SAH<br />
| + | |
| - | | + | |
| - | *'''tRNA guanine 6 N2 methyltransferase'''
| + | |
| - | | + | |
| - | **[[3tm5]] – PfTrm + SFG – ''Pyrococcvus furiosus''<br />
| + | |
| - | **[[3tlj]] – PfTrm + SAH<br />
| + | |
| - | **[[3tm4]] – PfTrm + SAM<br />
| + | |
| - | | + | |
| - | *Trm M2G/M22G10
| + | |
| - | | + | |
| - | **[[5e71]] – TkTrm – ''Thermococcvus kodakarensis''<br />
| + | |
| - | **[[5e72]] – TkTrm + SAM<br />
| + | |
| - | | + | |
| - | *Trm to N1 of adenine 9
| + | |
| - | | + | |
| - | **[[5a7t]], [[5a7z]] – SaTrm – ''Sulfolobus acidocaldarius''<br />
| + | |
| - | **[[4cnf]] – SaTrm (mutant)<br />
| + | |
| - | **[[5a7y]] – SaTrm + SAH<br />
| + | |
| - | **[[4cng]] – SaTrm (mutant) + SAH<br />
| + | |
| - | | + | |
| - | *'''tRNA adenine 58 N1 methyltransferase'''
| + | |
| - | | + | |
| - | **[[2pwy]] – TtTrm – ''Thermus thermophilus''<br />
| + | |
| - | **[[5c0o]], [[5c1i]] – TtTrm (mutant) + SAM<br />
| + | |
| - | **[[5eqj]] – yTrm <br />
| + | |
| - | **[[5erg]] – yTrm + SAM<br />
| + | |
| - | **[[5ccb]], [[5cd1]] – hTrm + SAH + tRNA-Lys<br />
| + | |
| - | **[[5ccx]] – hTrm + tRNA-Lys<br />
| + | |
| - | | + | |
| - | *Trm to uracil 34
| + | |
| - | | + | |
| - | **[[4qnx]] – EcTrm <br />
| + | |
| - | **[[4qnu]], [[4qnv]] – EcTrm + SAM derivative<br />
| + | |
| - | | + | |
| - | *'''tRNA uracil 54 methyltransferase'''
| + | |
| - | | + | |
| - | **[[3bt7]] – yTrm + RNA<br />
| + | |
| - | | + | |
| - | *Trm9-TRM112
| + | |
| - | | + | |
| - | **[[5cm2]] – Trm – ''Yarrowia lipolytica''<br />
| + | |
| - | | + | |
| - | *Trm dual specificity
| + | |
| | | | |
| - | **[[4pl1]], [[4pl2]] – EcTrm (mutant) + SAM<br /> | |
| - | **[[6ems]], [[6emt]] – TkTrm <br /> | |
| - | **[[6emu]] – TkTrm + SAM<br /> | |
| - | **[[6emv]] –TkTrm + SAH<br /> | |
| - | }} | |
| | == References == | | == References == |
| | <references/> | | <references/> |
| | [[Category:Topic Page]] | | [[Category:Topic Page]] |