This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Malate dehydrogenase
From Proteopedia
(Difference between revisions)
| (4 intermediate revisions not shown.) | |||
| Line 4: | Line 4: | ||
__TOC__ | __TOC__ | ||
==Function== | ==Function== | ||
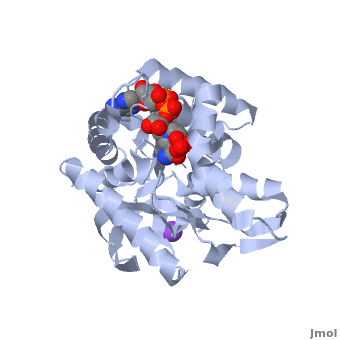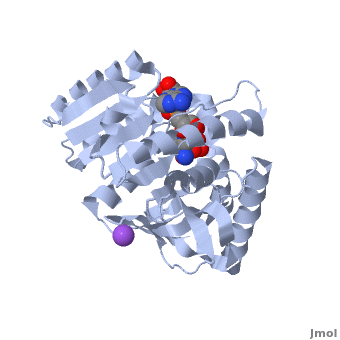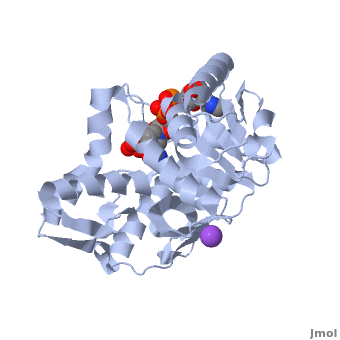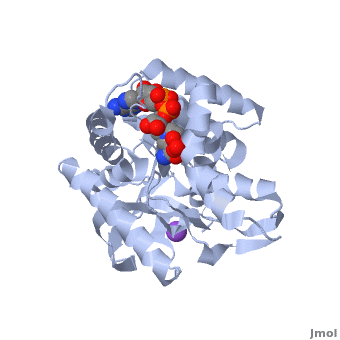
| - | [[Malate dehydrogenase|Malate Dehydrogenase]] (MDH | + | [[Malate dehydrogenase|Malate Dehydrogenase]] (MDH; PDB entry [http://www.pdb.org/pdb/explore/explore.do?structureId=2X0I 2x0i]) is most known for its role in the metabolic pathway of the [[tricarboxylic acid cycle]], also know as the Krebs cycle (after [http://en.wikipedia.org/wiki/Hans_Adolf_Krebs| Sir Hans Krebs]), which is critical to cellular respiration in cells [http://en.wikipedia.org/wiki/Citric_acid_cycle]; however, the enzyme is also involved on many other metabolic pathways such as glyoxylate bypass, amino acid synthesis, gluconeogenesis, and oxidation/reduction balance <ref>PMID:12537350</ref>. It is classified as an oxidoreductase[http://en.wikipedia.org/wiki/Oxidoreductase]. Malate dehydrogenase has been extensively studied due to its many isozymes <ref>PMID:20173310</ref>. The enzyme exists in two subcellular locations: mitochondria and cytoplasm. In the mitochondria, the enzyme catalyzes the reaction of malate to oxaloacetate; however, in the cytoplasm, the enzyme catalyzes oxaloacetate to malate to allow transport <ref>PMID:20173310</ref>. This conversion is an essential catalytic step in each different metabolic mechanism. The enzyme malate dehydrogenase is composed of either a dimer or tetramer depending on the location of the enzyme and the organism it is located in <ref>PMID: 9834842</ref>. During catalysis, the enzyme subunits are non-cooperative between active sites. The mitochondrial MDH suffers a complex allosteric control by citrate, but no other known metabolic regulation mechanisms have been discovered. Further, the exact mechanism of regulation has yet to be discovered <ref>PMID:7574693</ref>. The optimal pH is 7.6 for oxaloacetate conversion and 9.6 for malate conversion. The reported K<sub>m</sub> value for malate conversion is 215 µM and the V<sub>max</sub> value is 87.8 µM/min <ref>PMID:19277715</ref>. For halophilic MDH details, see [[Halophilic malate dehydrogenase]]. See also:<br /> |
*[[Krebs cycle carbons]] | *[[Krebs cycle carbons]] | ||
*[[Krebs cycle importance]] | *[[Krebs cycle importance]] | ||
| Line 10: | Line 10: | ||
*[[Citric Acid Cycle]] | *[[Citric Acid Cycle]] | ||
*[[Krebs cycle step 8]] | *[[Krebs cycle step 8]] | ||
| + | *[[Glyoxylate cycle]] | ||
{{Clear}} | {{Clear}} | ||
| Line 27: | Line 28: | ||
</StructureSection> | </StructureSection> | ||
| - | |||
| - | == 3D Structures of Malate Dehydrogenase == | ||
| - | |||
| - | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} | ||
| - | {{#tree:id=OrganizedByTopic|openlevels=0| | ||
| - | <br /> | ||
| - | The '''holo-MDH''' contains NAD or its derivatives while the '''apo-MDH''' lacks it. | ||
| - | |||
| - | * Holo-MDH | ||
| - | |||
| - | **[[2dfd]], [[4wlv]] – hMDH2 +NAD – human<br /> | ||
| - | **[[4mdh]] – pMDH+NAD - pig<br /> | ||
| - | **[[2x0r]] – HmMDH (mutant)+NAD - ''Haloarcula marismortui''<br /> | ||
| - | **[[1o6z]] - HmMDH (mutant)+NADH<br /> | ||
| - | **[[1hlp]] – HmMDH+NAD<br /> | ||
| - | **[[1x0i]] – AfMDH+NADH – ''Archaeoglobus fulgidus''<br /> | ||
| - | **[[2x0j]] - AfMDH+etheno-NAD<br /> | ||
| - | **[[1hlp]] – HmMDH+NAD<br /> | ||
| - | **[[1x0i]] – AfMDH+NADH <br /> | ||
| - | **[[2x0j]] - AfMDH+etheno-NAD<br /> | ||
| - | **[[1ib6]], [[1ie3]] – EcMDH (mutant)+NAD - ''Escherichia coli''<br /> | ||
| - | **[[3i0p]] – MDH+NAD – ''Entamoeba histolytica''<br /> | ||
| - | **[[3gvh]] – BmMDH+NAD – ''Brucella melitensis''<br /> | ||
| - | **[[1wze]] – TfMDH (mutant)+NAD – ''Thermus flavus''<br /> | ||
| - | **[[1wzi]] - TfMDH (mutant)+NDP<br /> | ||
| - | **[[1bmd]] – TfMDH+NAD<br /> | ||
| - | **[[1y7t]] – TtMDH+NADPH – ''Thermus thermophilus''<br /> | ||
| - | **[[2cvq]] - TtMDH+NADP<br /> | ||
| - | **[[1v9n]] – MDH+NADPH – ''Pyrococcus horikoshii''<br /> | ||
| - | **[[1z2i]] – MDH+NAD – ''Agrobacterium tumefaciens''<br /> | ||
| - | **[[1uxg]], [[1uxh]], [[1uxi]], [[1uxj]], [[1uxk]], [[1ur5]] – ChaMDH (mutant)+NAD – ''Chloroflexus aurantiacus''<br /> | ||
| - | **[[1guz]], [[1guy]], [[1gv0]] – CvMDH+NAD – ''Chlorobium vibrioforme''<br /> | ||
| - | **[[1civ]] – MDH+NADP – ''Flaveria bidentis''<br /> | ||
| - | **[[1b8u]], [[1b8v]] – AaMDH+NAD - ''Aquaspirillum arcticum''<br /> | ||
| - | **[[4i1i]] – LmMd + NAD – ''Leishmania major''<br /> | ||
| - | **[[5ujk]] – MeMd + NAD - ''Methylobacterium extorquens'' <br /> | ||
| - | **[[5nuf]], [[5nue]] – Md1 + NAD – ''Arabidopsis thaliana''<br /> | ||
| - | **[[4uup]] – Md + NADH - ''Trichiminad'' <br /> | ||
| - | **[[5kvv]] – MtMd + NADH – ''Mycobacterium tuberculosis'' <br /> | ||
| - | **[[6ihd]] – MsMd + NAD – ''Metallosphaera sedula''<br /> | ||
| - | |||
| - | * Md binary complex | ||
| - | |||
| - | **[[4wle]] – hMd2 + citrate <br /> | ||
| - | **[[4wlf]] – hMd2 + malate <br /> | ||
| - | **[[5kka]] – EcMd + inhibitor <br /> | ||
| - | **[[5z3w]] – EcMd + Ag<br /> | ||
| - | **[[2cmd]] - EcMd + citrate<br /> | ||
| - | **[[3gvi]] - BmMDH+ADP<br /> | ||
| - | **[[2hjr]] – MDH+adenosine diphosphoribose – ''Cryptosporidium parvum''<br /> | ||
| - | **[[1bdm]] - TfMDH (mutant)+beta-6-hydroxy-1,4,5,6-tetrahydronicotinamide adenine dinucleotide<br /> | ||
| - | **[[6bal]] – HiMd + malate - ''Haemophilus influenzae''<br /> | ||
| - | **[[6qss]] – Md + Tb derivative – ''Ignicoccus islandicus''<br /> | ||
| - | |||
| - | * Md ternary complex | ||
| - | |||
| - | **[[4wlo]] – hMd2 + NADH + oxaloacetate <br /> | ||
| - | **[[4wlu]] – hMd2 + NADH + malate <br /> | ||
| - | **[[5mdh]] – pMd+NAD+alpha-ketomalonic acid <br /> | ||
| - | **[[4plh]], [[4plt]] – ApMd + NAD + oxamate – ''Apicomplexa''<br /> | ||
| - | **[[4plv]], [[4plw]] – ApMd + NAD + lactate <br /> | ||
| - | **[[4ply]] – ApMd + NAD + malate <br /> | ||
| - | **[[4ros]] – MeMd + oxaloacetate + ADPR <br /> | ||
| - | **[[1emd]] – EcMd+NAD+citrate<br /> | ||
| - | **[[6ihe]] – MsMd + NAD + malate <br /> | ||
| - | **[[5zi2]] – yMd + NAD + ADP - yeast<br /> | ||
| - | **[[5zi4]] – yMd + NAD + oxalacetate <br /> | ||
| - | |||
| - | * apo-MDH | ||
| - | |||
| - | **[[4wln]] – hMd2 <br /> | ||
| - | **[[1mld]] – pMDH<br /> | ||
| - | **[[5zi3]] – yMd <br /> | ||
| - | **[[2j5r]], [[2j5k]], [[2j5q]], [[1d3a]], [[4jco]] – HmMDH <br /> | ||
| - | **[[2hlp]] – HmMDH (mutant)<br /> | ||
| - | **[[3hhp]], [[2pwz]] – EcMDH <br /> | ||
| - | **[[3fi9]] – MDH – ''Porphyromonas gingivalis'' <br /> | ||
| - | **[[3d5t]] - MDH – ''Burkholderia pseudomallei''<br /> | ||
| - | **[[2d4a]] – MDH – ''Aeropyrum pernix''<br /> | ||
| - | **[[1iz9]], [[4kde]], [[4kdf]] - TtMDH<br /> | ||
| - | **[[1sev]], [[1smk]] – MDH – ''Citrullus lanatus''<br /> | ||
| - | **[[1gv1]] – CvMDH <br /> | ||
| - | **[[1b8p]] – AaMDH <br /> | ||
| - | **[[7mdh]] – MDH – Sorgum bicolor<br /> | ||
| - | **[[3nep]] – Md – ''Salinibacter ruber''<br /> | ||
| - | **[[3p7m]] – Md – ''Francisella tularensis''<br /> | ||
| - | **[[3tl2]] – Md – ''Bacillus anthracis''<br /> | ||
| - | **[[4e0b]] – Md – ''Vibrio vulnificus''<br /> | ||
| - | **[[4h7p]] - LmMd<br /> | ||
| - | **[[4cl3]] – ChaMd <br /> | ||
| - | **[[4bgu]] – Md – ''Haloferax volcanii''<br /> | ||
| - | **[[4bgv]] – Md – ''Picrophilus torridus''<br /> | ||
| - | **[[4ror]] – MeMd <br /> | ||
| - | **[[4tvo]] – Md – ''Mycobacterium tuberculosis'' <br /> | ||
| - | **[[6aoo]] – HiMd <br /> | ||
| - | **[[5ulv]] – MeMd <br /> | ||
| - | **[[5nfr]] – Md – ''Plasmodium falciparum''<br /> | ||
| - | **[[4uuo]] – Md – ''Trichomonas vaginalis''<br /> | ||
| - | |||
| - | }} | ||
==Additional Resources== | ==Additional Resources== | ||
* [[Carbohydrate Metabolism]] | * [[Carbohydrate Metabolism]] | ||
| - | + | ||
<br /> | <br /> | ||
Current revision
| |||||||||||
Additional Resources
References
- ↑ Minarik P, Tomaskova N, Kollarova M, Antalik M. Malate dehydrogenases--structure and function. Gen Physiol Biophys. 2002 Sep;21(3):257-65. PMID:12537350
- ↑ Matsuda T, Takahashi-Yanaga F, Yoshihara T, Maenaka K, Watanabe Y, Miwa Y, Morimoto S, Kubohara Y, Hirata M, Sasaguri T. Dictyostelium Differentiation-Inducing Factor-1 Binds to Mitochondrial Malate Dehydrogenase and Inhibits Its Activity. J Pharmacol Sci. 2010 Feb 20. PMID:20173310
- ↑ Matsuda T, Takahashi-Yanaga F, Yoshihara T, Maenaka K, Watanabe Y, Miwa Y, Morimoto S, Kubohara Y, Hirata M, Sasaguri T. Dictyostelium Differentiation-Inducing Factor-1 Binds to Mitochondrial Malate Dehydrogenase and Inhibits Its Activity. J Pharmacol Sci. 2010 Feb 20. PMID:20173310
- ↑ Musrati RA, Kollarova M, Mernik N, Mikulasova D. Malate dehydrogenase: distribution, function and properties. Gen Physiol Biophys. 1998 Sep;17(3):193-210. PMID:9834842
- ↑ Boernke WE, Millard CS, Stevens PW, Kakar SN, Stevens FJ, Donnelly MI. Stringency of substrate specificity of Escherichia coli malate dehydrogenase. Arch Biochem Biophys. 1995 Sep 10;322(1):43-52. PMID:7574693 doi:http://dx.doi.org/10.1006/abbi.1995.1434
- ↑ Plancarte A, Nava G, Mendoza-Hernandez G. Purification, properties, and kinetic studies of cytoplasmic malate dehydrogenase from Taenia solium cysticerci. Parasitol Res. 2009 Jul;105(1):175-83. Epub 2009 Mar 10. PMID:19277715 doi:10.1007/s00436-009-1380-6
- ↑ Goward CR, Nicholls DJ. Malate dehydrogenase: a model for structure, evolution, and catalysis. Protein Sci. 1994 Oct;3(10):1883-8. PMID:7849603 doi:http://dx.doi.org/10.1002/pro.5560031027
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Jake Ezell, Joel L. Sussman, Joshua Johnson, Angel Herraez, Jaime Prilusky, David Canner