We apologize for Proteopedia being slow to respond. For the past two years, a new implementation of Proteopedia has been being built. Soon, it will replace this 18-year old system. All existing content will be moved to the new system at a date that will be announced here.
1hyb
From Proteopedia
(Difference between revisions)
| (One intermediate revision not shown.) | |||
| Line 3: | Line 3: | ||
<StructureSection load='1hyb' size='340' side='right'caption='[[1hyb]], [[Resolution|resolution]] 2.00Å' scene=''> | <StructureSection load='1hyb' size='340' side='right'caption='[[1hyb]], [[Resolution|resolution]] 2.00Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[1hyb]] is a 1 chain structure with sequence from [ | + | <table><tr><td colspan='2'>[[1hyb]] is a 1 chain structure with sequence from [https://en.wikipedia.org/wiki/Methanothermobacter_thermautotrophicus Methanothermobacter thermautotrophicus]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=1HYB OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=1HYB FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 2Å</td></tr> |
| - | <tr id=' | + | <tr id='ligand'><td class="sblockLbl"><b>[[Ligand|Ligands:]]</b></td><td class="sblockDat" id="ligandDat"><scene name='pdbligand=NMN:BETA-NICOTINAMIDE+RIBOSE+MONOPHOSPHATE'>NMN</scene>, <scene name='pdbligand=SO4:SULFATE+ION'>SO4</scene></td></tr> |
| - | + | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=1hyb FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=1hyb OCA], [https://pdbe.org/1hyb PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=1hyb RCSB], [https://www.ebi.ac.uk/pdbsum/1hyb PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=1hyb ProSAT]</span></td></tr> | |
| - | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[ | + | |
</table> | </table> | ||
| + | == Function == | ||
| + | [https://www.uniprot.org/uniprot/NADM_METTH NADM_METTH] | ||
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 36: | Line 37: | ||
</StructureSection> | </StructureSection> | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Methanothermobacter thermautotrophicus]] |
| - | [[Category: Christendat | + | [[Category: Christendat D]] |
| - | [[Category: Edwards | + | [[Category: Edwards AM]] |
| - | [[Category: Kimber | + | [[Category: Kimber MS]] |
| - | [[Category: Pai | + | [[Category: Pai EF]] |
| - | [[Category: Saridakis | + | [[Category: Saridakis V]] |
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
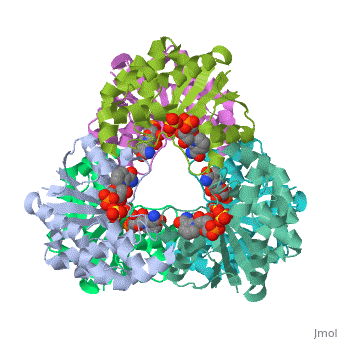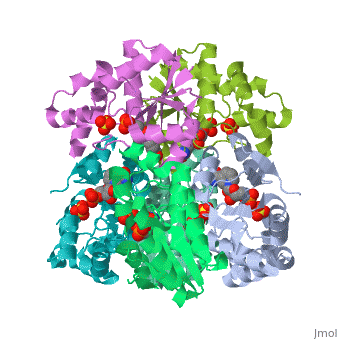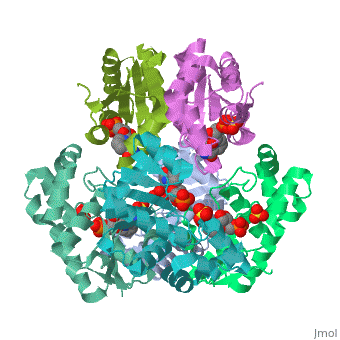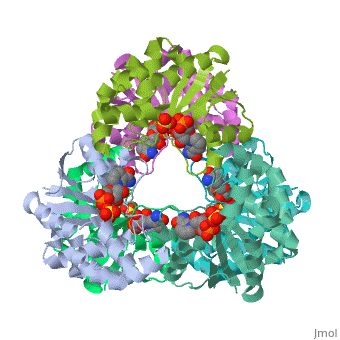
CRYSTAL STRUCTURE OF AN ACTIVE SITE MUTANT OF METHANOBACTERIUM THERMOAUTOTROPHICUM NICOTINAMIDE MONONUCLEOTIDE ADENYLYLTRANSFERASE
| |||||||||||