This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hemeproteins
From Proteopedia
(Difference between revisions)
| (45 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
<StructureSection load='Cytc6.pdb' size='400' side='right' scene='82/828363/Cv/1' caption=''> | <StructureSection load='Cytc6.pdb' size='400' side='right' scene='82/828363/Cv/1' caption=''> | ||
| - | ==Cytochromes== | + | =[[Catalase]]= |
| - | ==Cytochrome b5== | + | =Cytochromes= |
| - | '''Cytochrome b5''' (CB) functions as an electron transport carrier for several membrane-bound oxygenases. | + | ==[[Complex III of Electron Transport Chain]] includes Cytochrome c1 and cytochrome b subunits== |
| + | ==Cytochrome b== | ||
| + | ===Cytochrome b5=== | ||
| + | '''Cytochrome b5''' (CB) functions as an electron transport carrier for several membrane-bound oxygenases. CB is heme-containing protein. The microsomal and mitochondrial CB are membrane-bound while bacterial and other animal tissue CB are soluble. '''Cytochrome b562''' is the the b-type cytochrome from ''E. coli''.<ref>PMID:12559387</ref> <scene name='49/490878/Cv/2'>Rat heme-containing cytochrome b5</scene> (PDB entry [[1b5m]]<ref>PMID:8973214</ref>) is shown. | ||
| + | |||
| + | ===Cytochrome bc1 complex=== | ||
| + | '''Cytochrome bc1''' (Cbc1) functions as the central pump which transfers protons across the cell membrane. The protons are used to power the rotation of ATP synthase. Cbc1 binds ubiquinol which carries hydrogen atoms. Cbc1 separates the protons and the electrons. The protons are released in the inner side of the membrane for use by ATP synthase and the electrons are transferred to cytochrome c or to the outer side of the membrane. Plants use '''cytochrome b6f''' in the same manner binding plastoquinol as a hydrogen carrier. Stigmatellin inhibits the Cbc1 electron transfer by binding to its quinone oxidation site. Antimycin inhibits Cbc1 by binding to its quinone reduction site.<ref>PMID:14977419</ref> | ||
| + | |||
| + | More details in [[Complex_III_of_Electron_Transport_Chain]]. | ||
| + | |||
| + | '''Structural highlights''' | ||
| + | |||
| + | Cbc1 is a <scene name='49/490879/Cv/11'>dimeric protein</scene> composed of 11 proteins and cofactors which include heme-carrying proteins like <scene name='49/490879/Cv/12'>cytochrome b (Cb)</scene> and <scene name='49/490879/Cv/13'>cytochrome c1 (Cc1)</scene> and iron-sulfur cluster proteins like <scene name='49/490879/Cv/14'>Rieske Fe-S protein (RISP)</scene>. The iron containing moieties are <scene name='49/490879/Cv/15'>heme</scene>, <scene name='49/490879/Cv/16'>heme C</scene> (where vinyl side chain of heme are replaced by thioether) and <scene name='49/490879/Cv/17'>Fe2S2</scene>. <ref>PMID:16034531</ref> | ||
| + | |||
==Cytochrome c== | ==Cytochrome c== | ||
===Structural and kinetic studies of imidazole binding to two members of the cytochrome c6 family reveal an important role for a conserved heme pocket residue<ref>DOI 10.1007/s00775-011-0758-y</ref>=== | ===Structural and kinetic studies of imidazole binding to two members of the cytochrome c6 family reveal an important role for a conserved heme pocket residue<ref>DOI 10.1007/s00775-011-0758-y</ref>=== | ||
| Line 26: | Line 39: | ||
* <scene name='69/696899/Cv/14'>Click here to see morph of this scene</scene>. | * <scene name='69/696899/Cv/14'>Click here to see morph of this scene</scene>. | ||
Only a proximal 5-coordinate NO adduct, confirmed by structural data, is observed with no detectable hexacoordinate distal NO adduct. | Only a proximal 5-coordinate NO adduct, confirmed by structural data, is observed with no detectable hexacoordinate distal NO adduct. | ||
| + | |||
| + | === ''Rhodothermus marinus'' cytochrome ''c'' === | ||
| + | |||
| + | '''Structure''' | ||
| + | |||
| + | <scene name='Sandbox_Reserved_335/Heme/1'>'Figure 1. The heme group of monoheme cytochrome ''c'' purified from ''Rhodothermus marinus''</scene> | ||
| + | |||
| + | All members in the C-type cytochrome superfamily contain a heme prosthetic group that is covalently attached to the protein via two thioether bonds to cysteine residues. Most cytochromes ''c'' occur in a where the histidine residue is one of the two axial ligands of the heme iron.<ref name=main>PMID:18855424</ref> In monoheme cytochromes ''c'', the other axial position may be left vacant or be occupied by histidine or methionine residues; however, it can sometimes be occupied by cysteine or lysine residues.<ref name=main />. In ''Rm''cyt''c'', XX represents a threonine (Thr46) and an alanine residue (Ala47) that help form the loop 2 structure. | ||
| + | |||
| + | The typical monoheme cyt ''c'' fold is formed by helices <scene name='Sandbox_Reserved_335/Helices/4'>A, C, and E</scene>. ''Rm''cyt''c'' contains seven α-helices that are folded around the heme, all connected by random coils.<ref name=main /> The heme group is axially coordinated by <scene name='Sandbox_Reserved_335/Axial/6'>His49 and Met100</scene>, and the disulfide linkages exist at <scene name='Sandbox_Reserved_335/Cys/1'>Cys45 and Cys48</scene>. The heme group in ''Rm''cyt''c'' is almost completely shielded from solvent due to it being in a mostly hydrophobic pocket. This pocket is formed in part by the seven helices surrounding the ring, but also by two structures that are uncommon in other cytochromes ''c''. First, a 21 amino acid extension of the N-terminal exists, forming <scene name='Sandbox_Reserved_335/Uncommon1/2'>α-helix A' and loop 1</scene>, which wraps around the back of the polypeptide.<ref name=main /> An extension resembling such has only been seen in ''Thermus thermophilus''; however, the extension occurs at the C-terminus rather than the N-terminus.<ref>doi:10.1006/jmbi.1997.1181</ref> A second rarity is that of <scene name='Sandbox_Reserved_335/Uncommon2/2'>helix B'</scene>, inserted between helix D and loop 3, that shields the bottom part of the heme from any solvent.<ref name=main /> In cytochrome ''c''<sub>2</sub> as well as mitochondrial cyt ''c'', a similar yet shorter helix was found, though this helix was present at a different place in the primary sequence. Also, instead of helix B', ''T. thermophilus'' contains a two-stranded [http://en.wikipedia.org/wiki/Beta_sheet β-sheet].<ref name=main /> One final note is the number of <scene name='Sandbox_Reserved_335/Met/1'>methionine</scene> residues that ''Rm''cyt''c'' contains. In general, cyt ''c'' contains about two methionines whereas ''Rm''cyt''c'' contains seven, located on the left of the heme.<ref name=main /> | ||
| + | |||
| + | As determined by X-ray crystallography, the ''Rm''cyt''c'' structure was found to contain a sulfate ion coordinated to Glu122 via hydrogen bonding to the protonated carboxylate oxygen. In the protein complex, this ion has been seen to mediate crystal contact between neighbouring protein molecules.<ref name=main /> | ||
| + | |||
| + | The observation of these structural motifs in other C-type cytochromes can support the divergent evolution of cytochromes ''c''.<ref name=main /> These motifs are present in a number of different bacteria and are seen in similar regions of the secondary structure; however, they exist in the primary sequence in places distinct to the phylum. For example, monoheme cytochromes ''c'' in the rest of the Bacteroidetes phylum have an N-terminus extension that is highly conserved to that of ''Rm''cyt''c'', and the regions in the primary structure that correspond to these secondary motifs are not observed in other bacterial phyla.<ref name=main /> Also, due to these motifs being absent from other phyla, the Bacteroidetes monoheme cyt ''c'' has been said to form a new subfamily of cyt ''c''. | ||
| + | |||
| + | '''Function''' | ||
| + | |||
| + | Monoheme cytochromes ''c'' are involved in electron transport chains in both prokaryotes and eukaryotic mitochondria.<ref name=main /> They mediate the transfer of electrons mainly from the ''bc''<sub>1</sub> complexes or their analogs to heme-copper oxygen reductases (HCOs) in the [http://en.wikipedia.org/wiki/Electron_transport_chain electron transport chain] of [http://en.wikipedia.org/wiki/Oxidative_phosphorylation oxidative phosphorylation]. Heme ''c'' containing domains are often found fused to other protein domains such as these HCOs, including the ''caa''<sub>3</sub> oxygen reductases<ref name=main /><ref>PMID:14691678</ref>; these enzymes are membrane-bound and catalyze the reduction of O<sub>2</sub> to water.<ref>PMID:11334784</ref> In addition to being involved in oxidative phosphorylation, monoheme cyt ''c'' has also been seen to participate in the electron transport chain of [http://en.wikipedia.org/wiki/Photosynthesis photosynthesis].<ref name=main /> Cytochrome ''c'' has also been determined to be a major signalling molecule in the apoptotic pathways. | ||
| + | |||
| + | '''Electron transport chain''' | ||
| + | |||
| + | In the electron transport chain (ETC), cyt ''c'' shuttles electrons between the respiratory complexes III and IV; complex III is the cytochrome ''bc''<sub>1</sub> complex and IV is cyt ''c'' oxidase. Initially, the heme iron in cyt ''c'' is in the reduced, Fe<sup>3+</sup> state; this allows for the uptake of one electron, oxidizing the iron to the Fe<sup>2+</sup> state.<ref name='etc'>Karp, Gerald (2008). Cell and Molecular Biology (5th edition). Hoboken, NJ: John Wiley & Sons. ISBN 978-0470042175.</ref> The ETC in eukaryotes is quite simple compared to that of prokaryotes. | ||
| + | |||
| + | |||
| + | In prokaryotic systems, electrons can enter the ETC at a number of places and multiple donors can be in play; however, the underlying transport system remains the same. Electrons are ultimately transferred from donor to various redox complexes including the ''bc''<sub>1</sub> complex and cytochrome ''c'', and finally to a terminal electron acceptor such as molecular oxygen in eukaryotes.<ref name=etc /> | ||
| + | |||
| + | The cytochrome oxidase reaction accounts for nearly 90% of all oxygen uptake in most cells.<ref name=etc /> Due to the large role of cytochromes within the ETC, it would be highly detrimental to the cell if any inhibitors were to be present in the organism. Cyanide and azide bind tightly to the cytochrome oxidase complex, halting electron transport and reducing the overall ATP production.<ref name=etc /> | ||
| + | |||
| + | '''Apoptosis''' | ||
| + | |||
| + | In all organisms, cells undergo [http://en.wikipedia.org/wiki/Apoptosis apoptosis], or programmed cell death, by which there is an extrinsic and an intrinsic pathway. The extrinsic pathway involves an immune response by killer lymphocytes, and once the lymphocyte has been bound to the target cell, an apoptotic cascade occurs.<ref name=etc /> The intrinsic pathway includes cyt ''c'', present in the intermembrane space of mitochondria. In this pathway, the presence of an apoptotic stimulus causes cyt ''c'' to be released into the cytosol. Cytochrome ''c'' in the cytosol now can be recognized and bound to various apoptotic factors, activating them and forming the [http://en.wikipedia.org/wiki/Apoptosome apoptosome]. The apoptosome recruits [http://en.wikipedia.org/wiki/Caspase caspases], which are activated and result in a caspase cascade to proceed with apoptosis.<ref name=etc /> | ||
| + | |||
| + | Cytochrome ''c'' is required for the intrinsic apoptotic process to function properly. Such as with the electron transport chain, a mutation affecting cyt ''c'' or other structures in apoptosis could cause either an increase or a decrease in the rate of apoptosis. | ||
| + | |||
| + | ===[[Cytochrome c 7]]=== | ||
| + | |||
| + | ===[[Journal:Acta Cryst D:S2059798320003101|The crystal structure of heme ''d<sub>1</sub>'' biosynthesis-associated small c-type cytochrome NirC reveals mixed oligomeric states ''in crystallo'']]=== | ||
| + | |||
| + | ===For details on decaheme cyt see [[MtrF]]=== | ||
| + | |||
| + | ==Cytochrome f== | ||
| + | '''Cytochrome f''' (Cytf) is the largest subunit of the cytochrome b6f complex. This complex transfers electrons from plastocyanin to the two reaction center complexes of oxygenic photosynthetic membranes.<ref>PMID:7631417</ref> | ||
| + | |||
| + | The cytochrome b6f complex contains 4 subunits: Cytf, Cytb6, Rieske iron-sulfur protein and subunit IV. Cytf has an internal chain of water molecules conserved in all its 3D structures. The water chain is assumed to be a proton wire. | ||
| + | |||
| + | <scene name='48/484838/Cv/12'>Heme binding site</scene> in Turnip cytochrome f | ||
| + | |||
| + | <scene name='48/484838/Cv/13'>Covalent binding of heme to 2 Cys residues</scene> | ||
| + | |||
| + | <scene name='48/484838/Cv/14'>Fe coordination site</scene> (PDB entry [[1ctm]]<ref>PMID:8081747</ref>). | ||
| + | |||
| + | ==Cytochrome P450== | ||
| + | [[Cytochrome P450]] (P450) catalyzes the oxidation of organic substances like lipids. The P450 contains a heme cofactor. The protein is numbered by its gene.<ref>PMID:12369887</ref>.<br /> | ||
| + | * '''P450cam''', also known as camphor 5-monooxygenase, catalyzes the hydroxylation of camphor.<br /> | ||
| + | * '''P450nor''' is a Nitric Oxide Reductase P450.<br /> | ||
| + | * '''P450epoK''' converts epothilone D to epothilone B.<br /> | ||
| + | * '''P450cin''' monooxygenates 1,8-cineole which is similar to camphor.<br /> | ||
| + | * '''Bifunctional P450/NADPH P450 reductase''' (P450 BM3) is a fatty acid monooxygenase.<br /> | ||
| + | * '''P450 3A4''' see [[CYP3A4]].<br /> | ||
| + | * '''P450 19 family''' see [[Aromatase]].<br /> | ||
| + | Additional details on cytochrome P450 interactions with drugs see | ||
| + | *[[Drug Metabolism by CYP450 Enzymes]]<br /> | ||
| + | *[[Montelukast]] - anti rhinitis | ||
| + | *[[Noxafil]] - antifungal drug which binds to CYP51. | ||
| + | |||
| + | The <scene name='43/430105/Cv/6'>heme moiety is stabilized by several side chains</scene>. The <scene name='43/430105/Cv/7'>heme iron is pentacoordinated with Cys as one ligand</scene>.<ref>PMID:12861225</ref> | ||
| + | |||
| + | ==[[Flavocytochrome]]== | ||
| + | |||
| + | =Cytochrome c Nitrite Reductase= | ||
| + | == ''Shewanella oneidensis'' Cytochrome c Nitrite Reductase == | ||
| + | [[Journal:JBIC:16|Laue Crystal Structure of ''Shewanella oneidensis'' Cytochrome c Nitrite Reductase from a High-yield Expression System]]<ref name="Youngblut">doi 10.1007/s00775-012-0885-0</ref> | ||
| + | Cytochrome c nitrite reductase (ccNIR) is a central enzyme of the nitrogen cycle. It binds nitrite, and reduces it by transferring 6 electrons to form ammonia. This ammonia can then be utilized to synthesize nitrogen containing molecules such as amino acids or nucleic acids. However, ccNiR’s primary role is to help extract energy from the reduction; ammonia is simply a potentially useful byproduct. In general, heterotrophic organisms feed on electron-rich substances such as sugars or fatty acids. During the metabolism of these substances large numbers of electrons are produced. Many organisms use oxygen as the final acceptor of these electrons, in which case water is formed. However, some organisms can use alternative electron acceptors such as nitrite, which is where ccNiR comes in. | ||
| + | The ccNiR described here is produced by the ''Shewanella oneidensis'' bacterium, which is remarkable in its own right due to the large number of electron acceptors that it can utilize. ''Shewanella'' is a facultative anaerobe, which means that it will use oxygen if available, but in the absence of oxygen can get rid of its electrons by dumping them on a wide range of alternate acceptors, of which nitrite is only one example. To handle the electron flow ''Shewanella'' uses a large number of promiscuous <scene name='Journal:JBIC:16/Cv/8'>c-heme</scene> containing electron transfer proteins. Indeed, ''Shewanella'' is exceptionally adept at producing c-heme proteins under fast-growth conditions, which many bacteria commonly used for large-scale laboratory gene expression, such as ''E. coli'', are incapable of unless they are first extensively reprogrammed genetically. Since ''Shewanella'' can be easily grown in the lab, and can naturally and easily produce c-hemes, it is an ideal host for generating large quantities of c-heme proteins such as ccNiR. | ||
| + | The 2012 paper by Youngblut et al. <ref name="Youngblut">none yet</ref> describes a genetically modified ''Shewanella'' strain that can produce 20 – 40 times more ccNiR per liter of culture than the wild type bacterium. The ccNir so produced can be purified easily and in large amounts. This result is important because c-heme proteins have historically proved difficult to over-express in traditional vectors such as ''E. coli''. With large quantities of ''Shewanella'' ccNIR available, Youngblut et al <ref name="Youngblut">none yet</ref> were able to obtain the crystal structure ([[3ubr]]) and do a variety of experiments. The ccNIR consists of <scene name='Journal:JBIC:16/Cv/4'>two equal subunits</scene> (<font color='darkmagenta'><b>colored in darkmagenta</b></font> and <span style="color:lime;background-color:black;font-weight:bold;">in green</span>) with <scene name='Journal:JBIC:16/Cv/5'>five c-hemes each</scene>. In the oxidized ccNIR all central heme irons are Fe3+. They can be subsequently reduced to Fe2+ either by reducing agents or electrochemically. An important conclusion of the paper is that electrons added to ccNiR are likely <scene name='Journal:JBIC:16/Cv/6'>delocalized over several hemes</scene>, rather than localized on individual hemes. | ||
| + | The <span style="color:yellow;background-color:black;font-weight:bold;">hemes 3-5 (colored in yellow)</span> and the <span style="color:seagreen;background-color:black;font-weight:bold;">hemes-2 (colored in seagreen)</span> are six coordinate and used for electron transport only, whereas the <font color='magenta'><b>two hemes-1 (colored in magenta)</b></font> are the active sites. Electrons are believed to enter via the <span style="color:seagreen;background-color:black;font-weight:bold;">hemes-2</span>, but can move between subunits. Though the physiological significance of this result is not yet known, it is possible that delocalizing the electrons keeps the active site redox-potential sufficiently high until enough electrons are accumulated that the reaction with nitrite can take place. That is, CcNIR acts like a capacitor that can store electrons until they are needed. The X-ray structure of the ccNIR reveals the architecture of this capacitor. To solve the structure a non-standard method, the Laue method, was used. This became necessary since attempts to collect a high resolution data set with monochromatic X-ray radiation were not successful. At room temperature the ccNIR crystals are susceptible to radiation damage. Freezing damaged the crystals because a suitable cryoprotectant could not be found. Single pulsed Laue crystallography with 100 ps highly intense polychromatic X-ray pulses provided a solution. A dataset was collected in a few minutes. The crystals were cooled slightly to 0 °C but not frozen. Crystal settings spanned a range of 180 °C and the crystals were orthorhombic. Therefore, a Laue dataset with very high multiplicity and good quality in terms of resolution and R<sub>merge</sub> could be collected. The structure of this ccNIR was then solved by molecular replacement using the ''E. coli'' ccNIR as a template. <scene name='Journal:JBIC:16/Cv/10'>An overlay</scene> of the ''S. oneidensis'' hemes within one monomer with the corresponding ''E. coli'' hemes reveals significant similarity. ''S. oneidensis'' <span style="color:yellow;background-color:black;font-weight:bold;">hemes 3-5</span>, <span style="color:seagreen;background-color:black;font-weight:bold;">hemes-2</span>, and <font color='magenta'><b>hemes-1</b></font> are colored in <span style="color:yellow;background-color:black;font-weight:bold;">yellow</span>, <span style="color:seagreen;background-color:black;font-weight:bold;">seagreen</span>, and <font color='magenta'><b>magenta</b></font>, respectively, whereas their corresponding ''E. coli'' hemes are in similar, but darker colors. The <scene name='Journal:JBIC:16/Cv/11'>overall structure</scene> of ''S. oneidensis'' ccNiR also is similar to that of ''E. coli'' ccNiR, except in the region where the enzyme interacts with its physiological electron donor (CymA in the case of ''S. oneidensis'' ccNiR, NrfB in the case of the ''E. coli'' protein) near heme 2. Subunits of ''S. oneidensis'' ccNiR <font color='darkmagenta'><b>colored in darkmagenta</b></font> and <span style="color:lime;background-color:black;font-weight:bold;">in green</span>; subunits of ''E. coli'' ccNiR <span style="color:hotpink;background-color:black;font-weight:bold;">colored in hot pink</span> and <span style="color:deepskyblue;background-color:black;font-weight:bold;">in deep-sky-blue</span>. | ||
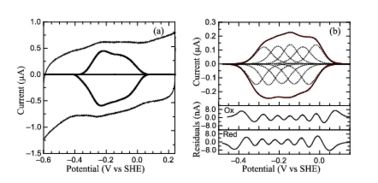
| + | Protein film voltammetry (PFV) experiments performed on ''S. oneidensis'' ccNiR films in the absence of substrate produced a broad envelope of reversible signals that span approximately 450 mV. At high pH values the envelope appears as a single peak, whereas at pH values below 7 the envelope appears to be composed of two large overlapping peaks. At pH values below 6 and at 0 °C, the envelope of signal can be better resolved and more than two peaks can be observed. This resulting envelope of signal can be deconvoluted as the sum of five one-electron peaks, each corresponding to one of the five hemes in a ccNiR monomer (see image below). | ||
| + | [[Image:figur5.jpg|left|378px|thumb|PFV of ''S. oneidensis'' ccNiR (a) Typical signal on a graphite electrode. (b) Baselinesubtracted non-turnover voltammogram]] | ||
| + | The Ca<sup>2+</sup> ion within <scene name='Journal:JBIC:16/Cv/14'>conserved site</scene> is coordinated in bidentate fashion by <scene name='Journal:JBIC:16/Cv/15'>Glu205</scene>, and in monodentate fashion by the <scene name='Journal:JBIC:16/Cv/16'>Tyr206 and Lys254</scene> backbone carbonyls, and the <scene name='Journal:JBIC:16/Cv/17'>Gln256</scene> side-chain carbonyl. In the ''S. oneidensis'' structure only <scene name='Journal:JBIC:16/Cv/18'>one water molecule</scene> is assigned to the Ca<sup>2+</sup> ion in subunit B. In subunit A the difference electron density that represents this water molecule is very close to the noise level, and it is difficult to identify even one water molecule there. The <scene name='Journal:JBIC:16/Cv/14'>carbonyl side chain of Asp242 and the hydroxyl of Tyr235</scene> come near to the open calcium coordination sites, but are not within bonding distance. Instead they interact with the water molecule that is weakly coordinated to the Ca<sup>2+</sup> ion. The ccNiR calcium ions appear to play a vital role in organizing the <scene name='Journal:JBIC:16/Cv/13'>active site</scene> (as was mentioned above <font color='magenta'><b>hemes-1</b></font> are the active sites). | ||
| + | |||
| + | ==[[Cytochrome c nitrite reductase]]== | ||
| + | |||
| + | =Cytochrome c oxidase= | ||
| + | '''Cytochrome c oxidase''' (CcO) is a transmembrane protein complex found in bacteria and mitochondria. CcO is the last enzyme in the electron transport chain. <ref>PMID:7652554</ref> | ||
| + | |||
| + | '''Disease''' | ||
| + | |||
| + | CcO deficiency is a genetic condition which affects muscle weakness, muscle tone upto severe brain dysfunction in severe cases. | ||
| + | |||
| + | '''Structural highlights''' | ||
| + | |||
| + | Mammalian CcO is composed of <scene name='46/466466/Cv/9'>13 subunits</scene>, <scene name='46/466466/Cv/10'>2 hemes, two copper centers</scene>, and cytochrome a and a3. The fully oxidized form of <scene name='46/466466/Cv/11'>CcO active site shows the heme, Cu+2 ion and an O2</scene> molecule. <scene name='46/466466/Cv/12'>Second heme binding site</scene>. <ref>PMID:9624044</ref> Bacterial CcO is composed of 2 subunits. | ||
| + | =Cytochrome c peroxidase= | ||
| + | '''Cytochrome c peroxidase''' (CcP) is a protoporphyrin-containing enzyme which reduces hydrogen peroxide to water.<ref>PMID:17027371</ref> | ||
| + | |||
| + | '''Structural highlights''' | ||
| + | |||
| + | The <scene name='46/466465/Cv/3'>O2 molecule in the active site coordinates with the heme</scene> (PDB entry [[1dcc]]).<ref>PMID:7664080</ref> Water molecules are shown as red spheres. | ||
| + | |||
| + | =Cytochrome P450 hydroxylase= | ||
| + | '''Cytochrome P450 hydroxylase''' (CPH) acts in the in-chain hydroxylation of lauric acid which is required for the development of the male organ in higher plants<ref>PMID:24798002</ref>. CPH hydroxylyzes the anti-cholesterol natural product herbosidiene<ref>PMID:25139567</ref>. | ||
| + | *<scene name='77/774654/Cv/6'>Lauric acid binding site</scene>. | ||
| + | *<scene name='77/774654/Cv/7'>Heme group binding site</scene>. | ||
| + | *<scene name='77/774654/Cv/8'>Fe coordination site</scene>. | ||
| + | *<scene name='77/774654/Cv/9'>Distances between Lauric acid oxygens and heme group Fe</scene> in Cytochrome P450 hydroxylase from ''Steptomyces avermitilis'' (PDB code [[5cwe]]). | ||
| + | =[[Heme oxygenase]]= | ||
| + | =[[Hemoglobin]]= | ||
| + | =[[Myoglobin]]= | ||
| + | *[[Extremophile|Elephant ''vs'' whale myoglobin]] | ||
| + | =Ascorbate peroxidase= | ||
| + | Function | ||
| + | |||
| + | '''Ascorbate peroxidase''' (APX) catalyzes the conversion of ascorbate and hydrogen peroxide to dehydroascorbate and water thus detoxifying hydrogen peroxide. APX is involved in the glutathione-ascorbate cycle. APX is accumulated in plants in response to heat and drought stress. APX contains a heme group. | ||
| + | |||
| + | Relevance | ||
| + | |||
| + | APX is part of plants antioxidant defense. APX converts H2O2 to water using ascorbate as an electron donor. APX is used in electron microscopy as a genetic tag which can be stained independently of light activation. | ||
| + | |||
| + | Structural highlights | ||
| + | |||
| + | The <scene name='48/486478/Cv/4'>heme-containing active site</scene> of APX contains a <scene name='48/486478/Cv/5'>His residue (H163 in soybean) which coordinates with the heme</scene> and confers stability to the Fe state in the heme. <ref>PMID:12640445</ref> | ||
| + | |||
| + | =[[Journal:Acta Cryst D:S2059798320013510|Influence of the presence of the heme cofactor on the JK-loop structure in indoleamine-2,3-dioxygenase-1]]= | ||
</StructureSection> | </StructureSection> | ||
<b>References</b><br> | <b>References</b><br> | ||
| Line 31: | Line 167: | ||
[[Category:Topic Page]] | [[Category:Topic Page]] | ||
| - | [[Category:Hemoproteins]] | ||
Current revision
| |||||||||||
References
- ↑ Schenkman JB, Jansson I. The many roles of cytochrome b5. Pharmacol Ther. 2003 Feb;97(2):139-52. PMID:12559387
- ↑ Rodriguez-Maranon MJ, Qiu F, Stark RE, White SP, Zhang X, Foundling SI, Rodriguez V, Schilling CL 3rd, Bunce RA, Rivera M. 13C NMR spectroscopic and X-ray crystallographic study of the role played by mitochondrial cytochrome b5 heme propionates in the electrostatic binding to cytochrome c. Biochemistry. 1996 Dec 17;35(50):16378-90. PMID:8973214 doi:10.1021/bi961895o
- ↑ Crofts AR. The cytochrome bc1 complex: function in the context of structure. Annu Rev Physiol. 2004;66:689-733. PMID:14977419 doi:http://dx.doi.org/10.1146/annurev.physiol.66.032102.150251
- ↑ Berry EA, Huang LS, Saechao LK, Pon NG, Valkova-Valchanova M, Daldal F. X-Ray Structure of Rhodobacter Capsulatus Cytochrome bc (1): Comparison with its Mitochondrial and Chloroplast Counterparts. Photosynth Res. 2004;81(3):251-75. PMID:16034531 doi:http://dx.doi.org/10.1023/B:PRES.0000036888.18223.0e
- ↑ Rajagopal BS, Wilson MT, Bendall DS, Howe CJ, Worrall JA. Structural and kinetic studies of imidazole binding to two members of the cytochrome c (6) family reveal an important role for a conserved heme pocket residue. J Biol Inorg Chem. 2011 Jan 26. PMID:21267610 doi:10.1007/s00775-011-0758-y
- ↑ Morelli X, Czjzek M, Hatchikian CE, Bornet O, Fontecilla-Camps JC, Palma NP, Moura JJ, Guerlesquin F. Structural model of the Fe-hydrogenase/cytochrome c553 complex combining transverse relaxation-optimized spectroscopy experiments and soft docking calculations. J Biol Chem. 2000 Jul 28;275(30):23204-10. PMID:10748163 doi:10.1074/jbc.M909835199
- ↑ Manole A, Kekilli D, Svistunenko DA, Wilson MT, Dobbin PS, Hough MA. Conformational control of the binding of diatomic gases to cytochrome c'. J Biol Inorg Chem. 2015 Mar 20. PMID:25792378 doi:http://dx.doi.org/10.1007/s00775-015-1253-7
- ↑ 8.00 8.01 8.02 8.03 8.04 8.05 8.06 8.07 8.08 8.09 8.10 8.11 8.12 Stelter M, Melo AM, Pereira MM, Gomes CM, Hreggvidsson GO, Hjorleifsdottir S, Saraiva LM, Teixeira M, Archer M. A Novel Type of Monoheme Cytochrome c: Biochemical and Structural Characterization at 1.23 A Resolution of Rhodothermus marinus Cytochrome c. Biochemistry. 2008 Oct 15. PMID:18855424 doi:10.1021/bi800999g
- ↑ Than ME, Hof P, Huber R, Bourenkov GP, Bartunik HD, Buse G, Soulimane T. Thermus thermophilus cytochrome-c552: A new highly thermostable cytochrome-c structure obtained by MAD phasing. J Mol Biol. 1997 Aug 29;271(4):629-44. PMID:9281430 doi:10.1006/jmbi.1997.1181
- ↑ Soares CM, Baptista AM, Pereira MM, Teixeira M. Investigation of protonatable residues in Rhodothermus marinus caa3 haem-copper oxygen reductase: comparison with Paracoccus denitrificans aa3 haem-copper oxygen reductase. J Biol Inorg Chem. 2004 Mar;9(2):124-34. Epub 2003 Dec 23. PMID:14691678 doi:10.1007/s00775-003-0509-9
- ↑ Pereira MM, Santana M, Teixeira M. A novel scenario for the evolution of haem-copper oxygen reductases. Biochim Biophys Acta. 2001 Jun 1;1505(2-3):185-208. PMID:11334784
- ↑ 12.0 12.1 12.2 12.3 12.4 12.5 Karp, Gerald (2008). Cell and Molecular Biology (5th edition). Hoboken, NJ: John Wiley & Sons. ISBN 978-0470042175.
- ↑ Prince RC, George GN. Cytochrome f revealed. Trends Biochem Sci. 1995 Jun;20(6):217-8. PMID:7631417
- ↑ Martinez SE, Huang D, Szczepaniak A, Cramer WA, Smith JL. Crystal structure of chloroplast cytochrome f reveals a novel cytochrome fold and unexpected heme ligation. Structure. 1994 Feb 15;2(2):95-105. PMID:8081747
- ↑ Danielson PB. The cytochrome P450 superfamily: biochemistry, evolution and drug metabolism in humans. Curr Drug Metab. 2002 Dec;3(6):561-97. PMID:12369887
- ↑ Williams PA, Cosme J, Ward A, Angove HC, Matak Vinkovic D, Jhoti H. Crystal structure of human cytochrome P450 2C9 with bound warfarin. Nature. 2003 Jul 24;424(6947):464-8. Epub 2003 Jul 13. PMID:12861225 doi:http://dx.doi.org/10.1038/nature01862
- ↑ 17.0 17.1 17.2 Youngblut M, Judd ET, Srajer V, Sayyed B, Goelzer T, Elliott SJ, Schmidt M, Pacheco AA. Laue crystal structure of Shewanella oneidensis cytochrome c nitrite reductase from a high-yield expression system. J Biol Inorg Chem. 2012 Mar 2. PMID:22382353 doi:10.1007/s00775-012-0885-0
- ↑ Tsukihara T, Aoyama H, Yamashita E, Tomizaki T, Yamaguchi H, Shinzawa-Itoh K, Nakashima R, Yaono R, Yoshikawa S. Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 A. Science. 1995 Aug 25;269(5227):1069-74. PMID:7652554
- ↑ Yoshikawa S, Shinzawa-Itoh K, Nakashima R, Yaono R, Yamashita E, Inoue N, Yao M, Fei MJ, Libeu CP, Mizushima T, Yamaguchi H, Tomizaki T, Tsukihara T. Redox-coupled crystal structural changes in bovine heart cytochrome c oxidase. Science. 1998 Jun 12;280(5370):1723-9. PMID:9624044
- ↑ Atack JM, Kelly DJ. Structure, mechanism and physiological roles of bacterial cytochrome c peroxidases. Adv Microb Physiol. 2007;52:73-106. PMID:17027371 doi:http://dx.doi.org/10.1016/S0065-2911(06)52002-8
- ↑ Miller MA, Shaw A, Kraut J. 2.2 A structure of oxy-peroxidase as a model for the transient enzyme: peroxide complex. Nat Struct Biol. 1994 Aug;1(8):524-31. PMID:7664080
- ↑ Yang X, Wu D, Shi J, He Y, Pinot F, Grausem B, Yin C, Zhu L, Chen M, Luo Z, Liang W, Zhang D. Rice CYP703A3, a cytochrome P450 hydroxylase, is essential for development of anther cuticle and pollen exine. J Integr Plant Biol. 2014 Oct;56(10):979-94. doi: 10.1111/jipb.12212. Epub 2014, Jul 15. PMID:24798002 doi:http://dx.doi.org/10.1111/jipb.12212
- ↑ Yu D, Xu F, Shao L, Zhan J. A specific cytochrome P450 hydroxylase in herboxidiene biosynthesis. Bioorg Med Chem Lett. 2014 Sep 15;24(18):4511-4514. doi:, 10.1016/j.bmcl.2014.07.078. Epub 2014 Aug 6. PMID:25139567 doi:http://dx.doi.org/10.1016/j.bmcl.2014.07.078
- ↑ Sharp KH, Mewies M, Moody PC, Raven EL. Crystal structure of the ascorbate peroxidase-ascorbate complex. Nat Struct Biol. 2003 Apr;10(4):303-7. PMID:12640445 doi:http://dx.doi.org/10.1038/nsb913