This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2yxr
From Proteopedia
(Difference between revisions)
| Line 3: | Line 3: | ||
<StructureSection load='2yxr' size='340' side='right'caption='[[2yxr]], [[Resolution|resolution]] 3.60Å' scene=''> | <StructureSection load='2yxr' size='340' side='right'caption='[[2yxr]], [[Resolution|resolution]] 3.60Å' scene=''> | ||
== Structural highlights == | == Structural highlights == | ||
| - | <table><tr><td colspan='2'>[[2yxr]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/ | + | <table><tr><td colspan='2'>[[2yxr]] is a 3 chain structure with sequence from [https://en.wikipedia.org/wiki/Methanocaldococcus_jannaschii Methanocaldococcus jannaschii]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2YXR OCA]. For a <b>guided tour on the structure components</b> use [https://proteopedia.org/fgij/fg.htm?mol=2YXR FirstGlance]. <br> |
| - | </td></tr><tr id=' | + | </td></tr><tr id='method'><td class="sblockLbl"><b>[[Empirical_models|Method:]]</b></td><td class="sblockDat" id="methodDat">X-ray diffraction, [[Resolution|Resolution]] 3.6Å</td></tr> |
| - | + | ||
<tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2yxr FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2yxr OCA], [https://pdbe.org/2yxr PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2yxr RCSB], [https://www.ebi.ac.uk/pdbsum/2yxr PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2yxr ProSAT]</span></td></tr> | <tr id='resources'><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[https://proteopedia.org/fgij/fg.htm?mol=2yxr FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2yxr OCA], [https://pdbe.org/2yxr PDBe], [https://www.rcsb.org/pdb/explore.do?structureId=2yxr RCSB], [https://www.ebi.ac.uk/pdbsum/2yxr PDBsum], [https://prosat.h-its.org/prosat/prosatexe?pdbcode=2yxr ProSAT]</span></td></tr> | ||
</table> | </table> | ||
== Function == | == Function == | ||
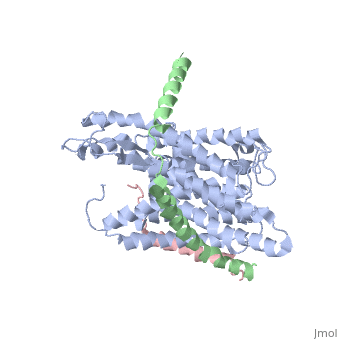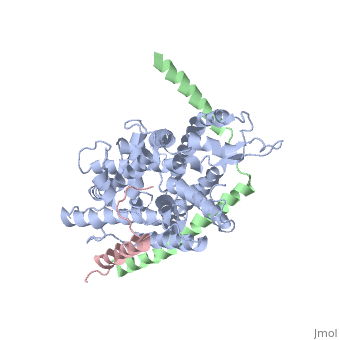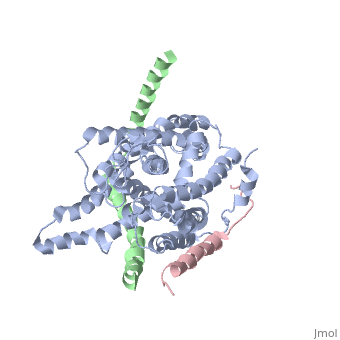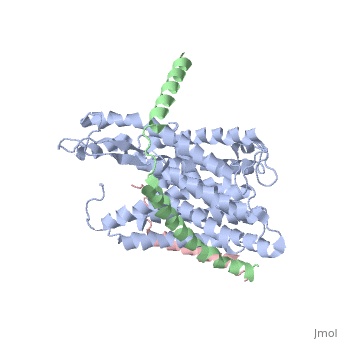
| - | + | [https://www.uniprot.org/uniprot/SECY_METJA SECY_METJA] The central subunit of the protein translocation channel SecYEG. Consists of two halves formed by TMs 1-5 and 6-10. These two domains form a lateral gate at the front which open onto the bilayer between TMs 2 and 7, and are clamped together by SecE at the back. The channel is closed by both a pore ring composed of hydrophobic SecY resides and a short helix (helix 2A) on the extracellular side of the membrane which forms a plug. The plug probably moves laterally to allow the channel to open. The ring and the pore may move independently. | |
== Evolutionary Conservation == | == Evolutionary Conservation == | ||
[[Image:Consurf_key_small.gif|200px|right]] | [[Image:Consurf_key_small.gif|200px|right]] | ||
| Line 23: | Line 22: | ||
==See Also== | ==See Also== | ||
*[[Preprotein translocase|Preprotein translocase]] | *[[Preprotein translocase|Preprotein translocase]] | ||
| + | *[[Preprotein translocase 3D structures|Preprotein translocase 3D structures]] | ||
__TOC__ | __TOC__ | ||
</StructureSection> | </StructureSection> | ||
| - | [[Category: Atcc 43067]] | ||
[[Category: Large Structures]] | [[Category: Large Structures]] | ||
| - | [[Category: | + | [[Category: Methanocaldococcus jannaschii]] |
| - | [[Category: | + | [[Category: Li W]] |
| - | [[Category: | + | [[Category: Schulman S]] |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Current revision
The plug domain of the SecY protein stablizes the closed state of the translocation channel and maintains a membrane seal
| |||||||||||