This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Test160122
From Proteopedia
(Difference between revisions)
| (59 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
| - | <StructureSection load='3i3f' size=' | + | <StructureSection load='3i3f' size='350' side='right' scene='90/901530/Cv/14' caption=''> |
| - | + | ||
| - | + | ||
| - | + | '''23/01/22''' | |
| - | == | + | *<scene name='90/901530/Cv/24'>Tusual, color is chosen same as atom after clicking on atom</scene> - after that, the label color is remaining white |
| + | *Everything is OK: <scene name='90/901530/Cv/30'>Similar to Tusual, color is chosen same as atom before clicking on atom</scene> | ||
| - | == | + | *<scene name='90/901530/Cv/26'>Trot</scene> |
| + | *<scene name='90/901530/Cv/27'>Trock</scene> | ||
| + | *<scene name='90/901530/Cv/28'>Twobble</scene> | ||
| - | == Structural highlights == | ||
| - | + | *<scene name='90/901530/Cv1/1'>Surfacenew</scene> | |
| + | *<scene name='90/901530/Cv1/2'>Surfacenew1</scene> | ||
| + | *<scene name='90/901530/Cv1/3'>Morph_Loop_6models</scene><jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | *<scene name='90/901530/Cv1/4'>Morph_Palindrom_2models</scene><jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | |||
| + | |||
| + | *<scene name='90/901530/L22s/2'>Mutation L22S (copy saved in present page): </scene> | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame 1 1 play; animation off; frame 1; select [LEU]22;color selectionHalos green;selectionHalos on; | ||
| + | </script> | ||
| + | <text>Wild type </text> | ||
| + | </jmolLink> | ||
| + | </jmol> | ||
| + | conformation and | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame 2 2 play; animation off; frame 2; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>mutation. | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> | ||
| + | Click here to see the | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation off; frame all; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>mutated and wild type | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> residues together, and an | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame play; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>animation | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> between the two states. Putative new hydrogen bonds were added. | ||
| + | <jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | |||
| + | *<scene name='90/901530/Cv1/5'>From uploaded file</scene> | ||
| + | |||
| + | |||
| + | |||
| + | --------- | ||
| + | *<scene name='90/901530/Cv/4'>160122was_with_labels/now180122_without_labels</scene> | ||
| + | |||
| + | *<scene name='90/901530/Cv/16'>180122saved_today_with_labels/in_text_without_labels</scene> | ||
| + | *<scene name='90/901530/Cv/17'>180122saved_today/should_be_simple_copy_of_previous_scene_without_rotation</scene> | ||
| + | *<scene name='90/901530/Cv/11'>Surface</scene> | ||
| + | *<scene name='90/901530/Cv/19'>Te</scene> | ||
| + | *<scene name='90/901530/Cv/23'>Terot</scene> | ||
| + | *<scene name='90/901530/Cv/22'>Tewobble</scene> | ||
| + | *<scene name='90/901530/Cv/3'>Test1</scene> | ||
| + | |||
| + | *<scene name='90/901530/Cv/5'>Te3</scene> | ||
| + | *<scene name='90/901530/Cv/10'>T8</scene> <jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | *<scene name='90/901530/Cv/12'>T7</scene> <jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | *<scene name='90/901530/Cv/8'>T7a</scene> <jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | *<scene name='75/751104/L22s/7'>Mutation L22S: </scene> | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame 1 1 play; animation off; frame 1; select [LEU]22;color selectionHalos green;selectionHalos on; | ||
| + | </script> | ||
| + | <text>Wild type </text> | ||
| + | </jmolLink> | ||
| + | </jmol> | ||
| + | conformation and | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame 2 2 play; animation off; frame 2; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>mutation. | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> | ||
| + | Click here to see the | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation off; frame all; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>mutated and wild type | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> residues together, and an | ||
| + | <jmol> | ||
| + | <jmolLink> | ||
| + | <script> | ||
| + | animation on; animation mode loop; frame play; select [SER]22;color selectionHalos red;selectionHalos on; | ||
| + | </script> | ||
| + | <text>animation | ||
| + | </text> | ||
| + | </jmolLink> | ||
| + | </jmol> between the two states. Putative new hydrogen bonds were added. | ||
| + | <jmol><jmolButton> | ||
| + | <script>if (_animating); anim pause;set echo bottom left; color echo white; font echo 20 sansserif;echo Animation Paused; else; anim resume; set echo off;endif;</script> | ||
| + | <text>Stop Animation</text> | ||
| + | </jmolButton></jmol> | ||
| + | |||
| + | ------------------------------ | ||
| + | [[Image:Figure3newt.png|left|450px|thumb|NEW: Despite the low sequence similarity the hypothetical protein is a prototypical YjgF/YER057c/UK114 endoribonuclease]] | ||
| + | {{Clear}} | ||
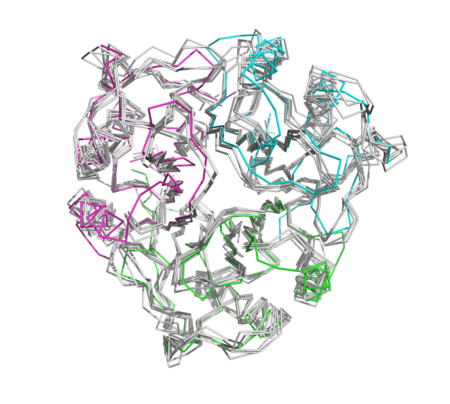
| + | [[Image:Trimercomparison1.png|left|450px|thumb|NEW]] | ||
| + | {{Clear}} | ||
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
<references/> | <references/> | ||
Current revision
| |||||||||||