This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1713
From Proteopedia
| Line 12: | Line 12: | ||
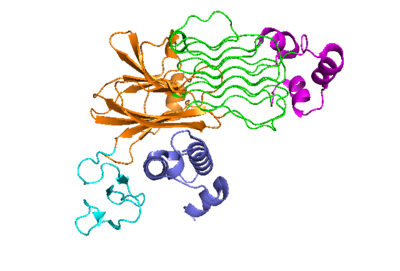
[[Image:Proteo_ALK-ALKAL_Monomer_White.png|400 px|right|thumb|Figure 2: ALK-ALKAL complex, showing the conformation change of ALK from the binding of ALKAL.]] | [[Image:Proteo_ALK-ALKAL_Monomer_White.png|400 px|right|thumb|Figure 2: ALK-ALKAL complex, showing the conformation change of ALK from the binding of ALKAL.]] | ||
===Conformational Change=== | ===Conformational Change=== | ||
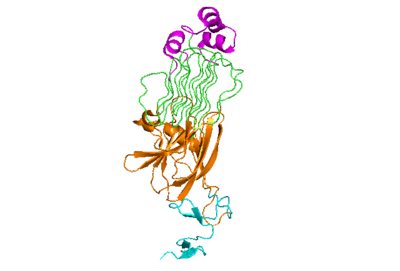
| - | The anaplastic lymphoma kinase activating ligand (ALKAL) is a triple alpha-helix polypeptide structure that signals for a conformational change of ALK. It <scene name='90/904318/ | + | The anaplastic lymphoma kinase activating ligand (ALKAL) is a triple alpha-helix polypeptide structure that signals for a conformational change of ALK. It <scene name='90/904318/Dimer_full_colored/1'>binds</scene> to ALKr at the TNFL domain, which has important negatively charged residues that form <scene name='90/904318/Binding_surface_with_residues/3'>ionic bonds</scene> with positively charged residues on ALKAL. These bonds initiate the conformational change, as these residues can only come into close proximity with each other if the conformational change occurs. The PXL and GlyR domains hinge forward when the change is initiated<ref>DOI: 10.1038/s41586-021-04140-8</ref> (Figure 2). Glu978, Glu974, Glu859, and Tyr966 are the residues of ALKr that form these bonds with Arg123, Arg133, Arg136, Arg140, and Arg117 of ALKAL. Once the ALK-ALKAL complex is formed, the <scene name='90/904317/Dimer_full_colored/3'>dimerization</scene> of two ALK-ALKAL complexes occurs. The main driving force of the interaction between two ALK-ALKAL complexes that dimerize are hydrophobic interactions of the PXL loop of one ALKr with the other complex's ALKAL and TNFL domain of ALKr. This dimer of two ALK-ALKAL complexes is the active form of ALK, and it is now able to perform its main function of phosphorylation. |
===Membrane Guidance of ALKAL to ALK=== | ===Membrane Guidance of ALKAL to ALK=== | ||
The negatively charged phosphate groups on the cell membrane interact with a highly conserved positively charged <scene name='90/904318/Alkalbindingsurfacewmembrane/1'>helix</scene> on ALKAL that faces the membrane. These <scene name='90/904318/Alkal1membraneinteraction/2'>residues that interact with the cell membrane</scene> guides ALKAL to ALK and correctly positions ALKAL for its <scene name='90/904318/Alk-alkal_binding_surface/4'>binding surface to face ALK's binding surface</scene>, which allows for a more favorable interaction. <ref>DOI: 10.1038/s41586-021-04140-8</ref> | The negatively charged phosphate groups on the cell membrane interact with a highly conserved positively charged <scene name='90/904318/Alkalbindingsurfacewmembrane/1'>helix</scene> on ALKAL that faces the membrane. These <scene name='90/904318/Alkal1membraneinteraction/2'>residues that interact with the cell membrane</scene> guides ALKAL to ALK and correctly positions ALKAL for its <scene name='90/904318/Alk-alkal_binding_surface/4'>binding surface to face ALK's binding surface</scene>, which allows for a more favorable interaction. <ref>DOI: 10.1038/s41586-021-04140-8</ref> | ||
Current revision
| This Sandbox is Reserved from February 28 through September 1, 2022 for use in the course CH462 Biochemistry II taught by R. Jeremy Johnson at the Butler University, Indianapolis, USA. This reservation includes Sandbox Reserved 1700 through Sandbox Reserved 1729. |
To get started:
More help: Help:Editing |
Anaplastic Lymphoma Kinase
Background
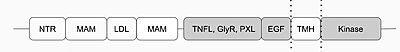
The anaplastic lymphoma kinase (ALK) was first discovered in 1994 as a tyrosine kinase in anaplastic large-cell lymphoma (ALCL) cells.[1] The specific type of tyrosine kinase ALK is classified as is a receptor tyrosine kinase (RTK) and like other RTKs, it's an integral protein with extracellular and intracellular domains and is involved in transmembrane signaling and communication within the cell. ALK is commonly expressed in the development of the nervous system. Anaplastic lymphoma kinase receptor (ALKr) is the extracellular portion of the RTK that includes a binding surface for a ligand to bind. When the ALK activating ligand (ALKAL) binds to ALKr, this causes a conformational change of ALK, allowing two ALK-ALKAL complexes to interact with each other, which will then allow intracellular kinase domain of ALK to phosphorylate a tyrosine residue on a downstream enzyme, which will activate this enzyme and activate a signaling cascade. Abnormal forms of ALK are closely related to the formation of several cancers. [2]
| |||||||||||
References
- ↑ Huang H. Anaplastic Lymphoma Kinase (ALK) Receptor Tyrosine Kinase: A Catalytic Receptor with Many Faces. Int J Mol Sci. 2018 Nov 2;19(11). pii: ijms19113448. doi: 10.3390/ijms19113448. PMID:30400214 doi:http://dx.doi.org/10.3390/ijms19113448
- ↑ Huang H. Anaplastic Lymphoma Kinase (ALK) Receptor Tyrosine Kinase: A Catalytic Receptor with Many Faces. Int J Mol Sci. 2018 Nov 2;19(11). pii: ijms19113448. doi: 10.3390/ijms19113448. PMID:30400214 doi:http://dx.doi.org/10.3390/ijms19113448
- ↑ Murray PB, Lax I, Reshetnyak A, Ligon GF, Lillquist JS, Natoli EJ Jr, Shi X, Folta-Stogniew E, Gunel M, Alvarado D, Schlessinger J. Heparin is an activating ligand of the orphan receptor tyrosine kinase ALK. Sci Signal. 2015 Jan 20;8(360):ra6. doi: 10.1126/scisignal.2005916. PMID:25605972 doi:http://dx.doi.org/10.1126/scisignal.2005916
- ↑ Reshetnyak AV, Rossi P, Myasnikov AG, Sowaileh M, Mohanty J, Nourse A, Miller DJ, Lax I, Schlessinger J, Kalodimos CG. Mechanism for the activation of the anaplastic lymphoma kinase receptor. Nature. 2021 Dec;600(7887):153-157. doi: 10.1038/s41586-021-04140-8. Epub 2021, Nov 24. PMID:34819673 doi:http://dx.doi.org/10.1038/s41586-021-04140-8
- ↑ Reshetnyak AV, Rossi P, Myasnikov AG, Sowaileh M, Mohanty J, Nourse A, Miller DJ, Lax I, Schlessinger J, Kalodimos CG. Mechanism for the activation of the anaplastic lymphoma kinase receptor. Nature. 2021 Dec;600(7887):153-157. doi: 10.1038/s41586-021-04140-8. Epub 2021, Nov 24. PMID:34819673 doi:http://dx.doi.org/10.1038/s41586-021-04140-8
- ↑ De Munck S, Provost M, Kurikawa M, Omori I, Mukohyama J, Felix J, Bloch Y, Abdel-Wahab O, Bazan JF, Yoshimi A, Savvides SN. Structural basis of cytokine-mediated activation of ALK family receptors. Nature. 2021 Oct 13. pii: 10.1038/s41586-021-03959-5. doi:, 10.1038/s41586-021-03959-5. PMID:34646012 doi:http://dx.doi.org/10.1038/s41586-021-03959-5
- ↑ Li T, Stayrook SE, Tsutsui Y, Zhang J, Wang Y, Li H, Proffitt A, Krimmer SG, Ahmed M, Belliveau O, Walker IX, Mudumbi KC, Suzuki Y, Lax I, Alvarado D, Lemmon MA, Schlessinger J, Klein DE. Structural basis for ligand reception by anaplastic lymphoma kinase. Nature. 2021 Dec;600(7887):148-152. doi: 10.1038/s41586-021-04141-7. Epub 2021, Nov 24. PMID:34819665 doi:http://dx.doi.org/10.1038/s41586-021-04141-7