This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
FirstGlance/How To Measure A Virus Capsid
From Proteopedia
(Difference between revisions)
| (23 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
<StructureSection load='' size='350' side='right' caption='' scene=''> | <StructureSection load='' size='350' side='right' caption='' scene=''> | ||
| - | + | {{Template:EEEV}} | |
| - | + | '''Quick Start''': [http://firstglance.jmol.org/fg.htm?mol=6mx4 Analyze the EEEV capsid 6mx4 in FirstGlance]. | |
| - | + | [[FirstGlance in Jmol]] automatically constructs the capsid, and simplifies it to a subset of alpha carbon atoms small enough (not more than 25,000) to be analyzed efficiently in FirstGlance and [[JSmol]], both of which run in the Javascript of the web browser. FirstGlance offers a number of useful color schemes, including distance from center, colors that distinguish sequence-identical groups of chains, and [[Help:Color_Keys#Rainbows:_N_to_C.2C_5.27_to_3.27|amino-to-carboxy rainbow]]. When the structure is small enough that all alpha carbons can be displayed (not more than 250,000 alpha carbons; EEEV has 242,340), it can also be colored by charge, hydrophobic vs. polar, or [[evolutionary conservation]]. | |
| - | + | ==Slab of the Capsid== | |
| + | In order to estimate the dimensions of the capsid, it is useful to "cut out a slab". | ||
| + | {{Template:EEEV-slab}} | ||
| + | |||
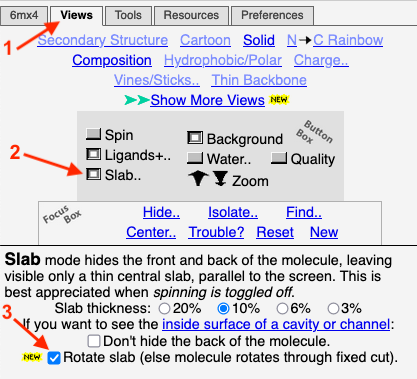
| + | To obtain this slab in FirstGlance, you simply depress a ''Slab'' button. By default, the thickness of the slab is 10% of the diameter of the capsid (but this is adjustable). Here is a '''snapshot of the ''Views'' tab of FirstGlance'''. After the initial view of the capsid appears, getting this slab takes 3 clicks indicated by the red arrows: Click on the ''Views'' tab, depress the ''Slab'' button, and check ''Rotate slab''. | ||
| + | [[Image:Firstglance-slab-controls.png]] | ||
| + | |||
| + | ==Measuring the Slab== | ||
<jmol> | <jmol> | ||
<jmolLink> | <jmolLink> | ||
<script> | <script> | ||
script /wiki/images/e/ee/Echo-loading.spt; | script /wiki/images/e/ee/Echo-loading.spt; | ||
| - | script /wiki/images/5/ | + | script /wiki/images/5/52/6mx4-nohet-slab-distance-diameters2.spt; |
spin on; | spin on; | ||
</script> | </script> | ||
| - | <text> | + | <text>The outside diameter of the capsid is about 644 Angstroms.</text> |
</jmolLink> | </jmolLink> | ||
</jmol> | </jmol> | ||
| - | ( | + | In FirstGlance (and JSmol), distances are measured by double-clicking on first one, then the second of the two relevant atoms. The distance between the centers of two of the outermost blue atoms is '''644 Å''', while the distance between two of the innermost red atoms is '''315 Å'''. The length of the transmembrane spikes/posts is about '''35 Å''', consistent with the thickness of a lipid bilayer. |
| - | + | ==Three Protein Sequences== | |
| - | + | ||
| - | == | + | |
| - | + | ||
<jmol> | <jmol> | ||
<jmolLink> | <jmolLink> | ||
<script> | <script> | ||
script /wiki/images/e/ee/Echo-loading.spt; | script /wiki/images/e/ee/Echo-loading.spt; | ||
| - | script /wiki/images/ | + | script /wiki/images/2/26/6mx4-nohet-slab-seqid.spt; |
spin on; | spin on; | ||
</script> | </script> | ||
| - | <text> | + | <text>The capsid is made up of 240 copies of each of 3 protein sequences |
| + | </text> | ||
</jmolLink> | </jmolLink> | ||
</jmol> | </jmol> | ||
| - | + | (total 720 protein chains). | |
| - | + | For more about the EEEV capsid, see [[FirstGlance/Virus_Capsids_and_Other_Large_Assemblies#Eastern_Equine_Encephalitis_Virus|Virus Capsids and Other Large Assemblies]]. | |
| - | [[ | + | |
</StructureSection> | </StructureSection> | ||
| + | |||
| + | ==Reference== | ||
| + | <references /> | ||
Current revision
| |||||||||||
Reference
- ↑ Hasan SS, Sun C, Kim AS, Watanabe Y, Chen CL, Klose T, Buda G, Crispin M, Diamond MS, Klimstra WB, Rossmann MG. Cryo-EM Structures of Eastern Equine Encephalitis Virus Reveal Mechanisms of Virus Disassembly and Antibody Neutralization. Cell Rep. 2018 Dec 11;25(11):3136-3147.e5. doi: 10.1016/j.celrep.2018.11.067. PMID:30540945 doi:http://dx.doi.org/10.1016/j.celrep.2018.11.067