This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 1786
From Proteopedia
(Difference between revisions)
| (3 intermediate revisions not shown.) | |||
| Line 19: | Line 19: | ||
===Transmembrane Region=== | ===Transmembrane Region=== | ||
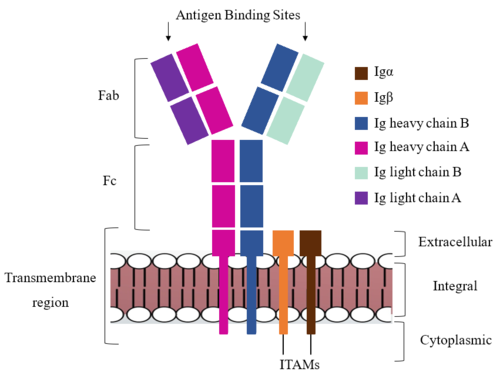
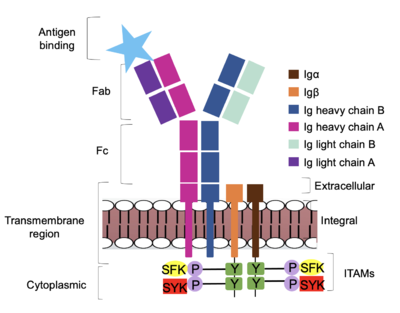
| - | <div style="text-align: left">The IgM BCR is anchored to [https://en.wikipedia.org/wiki/B_cell B-cell] membranes through the <scene name='95/952714/Integral_region/15'>transmembrane region</scene> which is broken up into both extracellular and integral domains which sit on top of or span through the membrane, respectively (Figure 1). The | + | <div style="text-align: left">The IgM BCR is anchored to [https://en.wikipedia.org/wiki/B_cell B-cell] membranes through the <scene name='95/952714/Integral_region/15'>transmembrane region</scene> which is broken up into both extracellular and integral domains which sit on top of or span through the membrane, respectively (Figure 1). The extracellular region is primarily composed of [https://proteopedia.org/wiki/index.php/Beta_sheet β-sheets]while the integral region is composed of [https://proteopedia.org/wiki/index.php/Alpha_helix#:~:text=An%20alpha%20helix%20is%20a,can%20be%20of%20arbitrary%20length. α-helices]. IgM BCR assembly requires dimerization of the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> subunits which embed within the B-cell membrane. <ref name="Tolar"/> The <scene name='95/952714/Ig_alpha_beta/5'>Igα and Igβ heterodimer</scene> dimerizes within the extracellular region via a <scene name='95/952714/Extracellular_disulfide_bridge/6'>disulfide bridge</scene>. Additional dimerization occurs within the integral region via a hydrogen bond; the residues involved have not been confirmed. Although the mechanism of disulfide bridge formation is still unknown, <scene name='95/952714/Extracellular_glycosylation/2'>glycosylation</scene> via [https://en.wikipedia.org/wiki/N-linked_glycosylation N-linked asparagine glycosyl groups] <b><span class="text-lightgreen">(NAGs)</span></b> in the extracellular region of both the <b><span class="text-brown">Igα</span></b> and and <b><span class="text-orange">Igβ</span></b> chains is hypothesized to facilitate this process. The NAG groups are believed to be essential for the recruitment of [https://en.wikipedia.org/wiki/Chaperone_(protein) Chaperone proteins] to optimize the folding process. <ref name="Daniels">PMID:12535523</ref> Past studies with human and viral proteins have shown that the presence of NAGs not only facilitate the rapid formation of disulfide bridges, but also ensure correct location. <ref name="Bakshi">PMID:35409101</ref>, <ref name="Mirazimi">PMID:9557673</ref> The recruited chaperone proteins will remain bound to the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> subunits until dimerization occurs. <ref name="Dylke"/> |
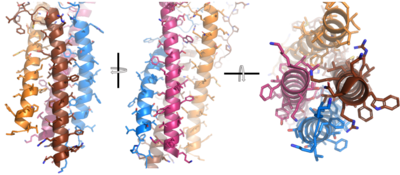
After <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> dimerization, the transmembrane helices of the heavy chains can embed within the B-cell membrane and intertwine with the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> chains. <ref name="Tolar"/> The side chains of this <scene name='95/952714/Integral_helices_2/2'>4-pass integral helix structure</scene>, made up by the <b><span class="text-brown">alpha</span></b>, <b><span class="text-orange">beta</span></b>, and heavy chains ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>), are primarily hydrophobic; this allows for interactions with the hydrophobic tails in the [https://en.wikipedia.org/wiki/Lipid_bilayer phospholipid bilayer]. The four helices (Figure 2) are intertwined and primarily held together through interactions between the <scene name='95/952714/Integral_helices_2/3'>hydrophobic side chains</scene>; however, a a few polar residues are included which allow for additional interactions with the polar residues on the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> chains. <ref name="Dylke"/> | After <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> dimerization, the transmembrane helices of the heavy chains can embed within the B-cell membrane and intertwine with the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> chains. <ref name="Tolar"/> The side chains of this <scene name='95/952714/Integral_helices_2/2'>4-pass integral helix structure</scene>, made up by the <b><span class="text-brown">alpha</span></b>, <b><span class="text-orange">beta</span></b>, and heavy chains ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>), are primarily hydrophobic; this allows for interactions with the hydrophobic tails in the [https://en.wikipedia.org/wiki/Lipid_bilayer phospholipid bilayer]. The four helices (Figure 2) are intertwined and primarily held together through interactions between the <scene name='95/952714/Integral_helices_2/3'>hydrophobic side chains</scene>; however, a a few polar residues are included which allow for additional interactions with the polar residues on the <b><span class="text-brown">Igα</span></b> and <b><span class="text-orange">Igβ</span></b> chains. <ref name="Dylke"/> | ||
| Line 34: | Line 34: | ||
===Fc Region=== | ===Fc Region=== | ||
| - | The constant region of IgM is made up of the two <scene name='95/952714/Heavy_chain/1'>heavy chains</scene>. These heavy chains form a bridge connecting the | + | The constant region of IgM is made up of the two <scene name='95/952714/Heavy_chain/1'>heavy chains</scene>. These heavy chains form a bridge connecting the Fab region or variable region to the transmembrane region (Figure 1). They also act as a wire that the variable region can send a signal through to the transmembrane region as a mechanical change. |
| - | <scene name='95/952715/Extracellular_transmembrane_v2/10'>Extracellular transmembrane interactions</scene> help hold the heavy chains and <b><span class="text-brown">Igα</span></b>/<b><span class="text-orange">Igβ</span></b> chains together in the extracellular portion of the transmembrane region. | + | <scene name='95/952715/Extracellular_transmembrane_v2/10'>Extracellular transmembrane interactions</scene> help hold the heavy chains and <b><span class="text-brown">Igα</span></b>/<b><span class="text-orange">Igβ</span></b> chains together in the extracellular portion of the transmembrane region. Because a conformational change occurs throughout the entirety of the IgM-BCR complex, the Fc region must be able to tolerate the contortion of the structure as the antigen binds. In constant region two, which is located at the start of the Fc region, '''{{Font color|violet|heavy chain A}}''' and <b><span class="text-blue">heavy chain B</span></b> make a <scene name='95/952713/Disulfides/5'>disulfide bridge</scene> to stabilize the IgM-BCR and drive downstream signaling. |
| - | + | To maximize the Fc region’s signal transduction efficiency and [https://en.wikipedia.org/wiki/Van_der_Waals_force#:~:text=If%20no%20other%20force%20is,between%20the%20atoms'%20electron%20clouds. Van der Waals contacts], constant region two of '''{{Font color|violet|heavy chain A}}''' makes an asymmetrical association with constant region three of <b><span class="text-blue">heavy chain B</span></b> to create a <scene name='95/952713/Trans_heavy/7'>heavy chain interface</scene>. More specifically, Arg243 and Arg251 residues from '''{{Font color|violet|heavy chain A}}''' donate three hydrogen bonds to Leu433, Thr431, and Asp376 residues on <b><span class="text-blue">heavy chain B</span></b>. Furthermore, Leu313 of heavy chain A accepts a hydrogen bond from Thr429 on heavy chain B. <ref name="Ma">PMID:35981028</ref> | |
| - | + | ||
| - | + | ||
---- | ---- | ||
| Line 46: | Line 44: | ||
===Fab Region=== | ===Fab Region=== | ||
| - | The Fab region of the antibody is where antigen recognition occurs upon binding (Figure 1). On each arm is one heavy ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>) and one light (<b><span class="text-purple">A</span></b>/<b><span class="text-cyan">B</span></b>) chain, both containing domains identical to their respective counterparts. Repeats of β-sandwiches form the [https://en.wikipedia.org/wiki/Antibody constant and variable domains] within the Fab region | + | The Fab region of the antibody is where antigen recognition occurs upon binding (Figure 1). On each arm is one heavy ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>) and one light (<b><span class="text-purple">A</span></b>/<b><span class="text-cyan">B</span></b>) chain, both containing domains identical to their respective counterparts. Repeats of β-sandwiches form the [https://en.wikipedia.org/wiki/Antibody constant and variable domains] within the Fab region. Antigen recognition occurs at the variable domain while the constant domain connects it to the rest of the IgM complex. Because the Fab region of IgM is poorly resolved, a structural analysis of an HIV neutralizing antibody called VCR01 was performed to approximate where an antigen would bind to at the <scene name='95/952714/Variable_region/1'>variable region</scene>. <ref name="Zhou">PMID:20616231</ref> |
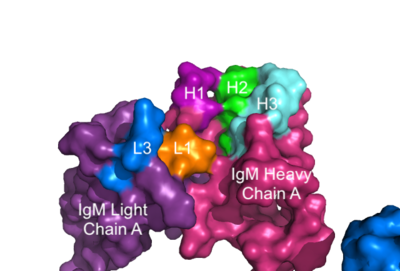
The IgM-BCR contains areas referred to as [https://en.wikipedia.org/wiki/Complementarity-determining_region complementary-determining regions](CDRs), which are where the antigen makes <scene name='95/952713/Antigen_antibody_contact/2'>contact</scene> with the antibody on the Fab domain. Figure 2 depicts this as a surface representation given that the specific residues within the antigen-binding motif are unknown. | The IgM-BCR contains areas referred to as [https://en.wikipedia.org/wiki/Complementarity-determining_region complementary-determining regions](CDRs), which are where the antigen makes <scene name='95/952713/Antigen_antibody_contact/2'>contact</scene> with the antibody on the Fab domain. Figure 2 depicts this as a surface representation given that the specific residues within the antigen-binding motif are unknown. | ||
| - | Due to the poor resolution of the Fab region, specific side chain interactions between the heavy ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>) and light (<b><span class="text-purple">A</span></b>/<b><span class="text-cyan">B</span></b>) chains have not been determined. | + | Due to the poor resolution of the Fab region, specific side chain interactions between the heavy ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>) and light (<b><span class="text-purple">A</span></b>/<b><span class="text-cyan">B</span></b>) chains have not been determined. Based on the general structure of an [https://en.wikipedia.org/wiki/Antibody antibody] it is estimated that each β-sandwich contains one disulfide bridge with additional hydrogen bonds. The <scene name='95/952713/Heavy-light_chain_interface/2'>heavy-light chain interface</scene> shows how the four heavy and light chain β-sandwiches fit together. The Fab region heavy chains attach to the Fc region heavy chains, before continuing down into the intracellular domain to interact with the <b><span class="text-brown">Igα</span></b>/<b><span class="text-orange">Igβ</span></b> subunits. The light chains (<b><span class="text-purple">A</span></b>/<b><span class="text-cyan">B</span></b>) however are only connected to the heavy chains ('''{{Font color|violet|A}}'''/<b><span class="text-blue">B</span></b>) within the Fab region, thus have no contact with the <b><span class="text-brown">Igα</span></b>/<b><span class="text-orange">Igβ</span></b> heterodimer. |
[[Image:IgM_Surface2.png|400 px|left|thumb|'''Figure 3. Surface Representation of IgM Antibody Binding Pocket.''' On one arm of the IgM antibody, the antigen makes contact with light chain A at the L1 and L3 complementary-determining regions. Furthermore, it makes contact with heavy chain A at the H1, H2, and H3 complementary-determining regions. The location of the complementary-determining regions were approximated using the structure of the VCR01 variable region and were visualized using PyMOL.]] | [[Image:IgM_Surface2.png|400 px|left|thumb|'''Figure 3. Surface Representation of IgM Antibody Binding Pocket.''' On one arm of the IgM antibody, the antigen makes contact with light chain A at the L1 and L3 complementary-determining regions. Furthermore, it makes contact with heavy chain A at the H1, H2, and H3 complementary-determining regions. The location of the complementary-determining regions were approximated using the structure of the VCR01 variable region and were visualized using PyMOL.]] | ||
Current revision
Human B-cell Antigen Receptor: IgM BCR
| |||||||||||
References
- ↑ Sathe A, Cusick JK. Biochemistry, Immunoglobulin M. 2022 Dec 19. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023 Jan–. PMID: 32310455. https://pubmed.ncbi.nlm.nih.gov/32310455/
- ↑ 2.0 2.1 Su Q, Chen M, Shi Y, Zhang X, Huang G, Huang B, Liu D, Liu Z, Shi Y. Cryo-EM structure of the human IgM B cell receptor. Science. 2022 Aug 19;377(6608):875-880. doi: 10.1126/science.abo3923. Epub 2022, Aug 18. PMID:35981043 doi:http://dx.doi.org/10.1126/science.abo3923
- ↑ 3.0 3.1 3.2 3.3 Ma X, Zhu Y, Dong, Chen Y, Wang S, Yang D, Ma Z, Zhang A, Zhang F, Guo C, Huang Z. Cryo-EM structures of two human B cell receptor isotypes. Science. 2022 Aug 19;377(6608):880-885. doi: 10.1126/science.abo3828. Epub 2022, Aug 18. PMID:35981028 doi:http://dx.doi.org/10.1126/science.abo3828
- ↑ 4.0 4.1 4.2 Tolar P, Pierce SK. Unveiling the B cell receptor structure. Science. 2022 Aug 19;377(6608):819-820. doi: 10.1126/science.add8065. Epub 2022 Aug 18.[http://dx.doi.org/10.1126/science.add8065 DOI:10.1126/science.add8065
- ↑ 5.0 5.1 5.2 Dylke J, Lopes J, Dang-Lawson M, Machtaler S, Matsuuchi L. Role of the extracellular and transmembrane domain of Ig-alpha/beta in assembly of the B cell antigen receptor (BCR). Immunol Lett. 2007 Sep 15;112(1):47-57. doi: 10.1016/j.imlet.2007.06.005. Epub 2007 Jul 23. [http://dx.doi.org/10.1016/j.imlet.2007.06.005 DOI:10.1016/j.imlet.2007.06.005
- ↑ Daniels R, Kurowski B, Johnson AE, Hebert DN. N-linked glycans direct the cotranslational folding pathway of influenza hemagglutinin. Mol Cell. 2003 Jan;11(1):79-90. PMID:12535523 doi:10.1016/s1097-2765(02)00821-3
- ↑ Bakshi T, Pham D, Kaur R, Sun B. Hidden Relationships between N-Glycosylation and Disulfide Bonds in Individual Proteins. Int J Mol Sci. 2022 Mar 29;23(7):3742. PMID:35409101 doi:10.3390/ijms23073742
- ↑ Mirazimi A, Svensson L. Carbohydrates facilitate correct disulfide bond formation and folding of rotavirus VP7. J Virol. 1998 May;72(5):3887-92. PMID:9557673 doi:10.1128/JVI.72.5.3887-3892.1998
- ↑ Zhou T, Georgiev I, Wu X, Yang ZY, Dai K, Finzi A, Do Kwon Y, Scheid JF, Shi W, Xu L, Yang Y, Zhu J, Nussenzweig MC, Sodroski J, Shapiro L, Nabel GJ, Mascola JR, Kwong PD. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science. 2010 Aug 13;329(5993):811-7. Epub 2010 Jul 8. PMID:20616231 doi:10.1126/science.1192819
Student Contributors
DeTonyeá Dickson, Allison Goss, Jackson Payton