This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Ubiquitin
From Proteopedia
(Difference between revisions)
| (3 intermediate revisions not shown.) | |||
| Line 1: | Line 1: | ||
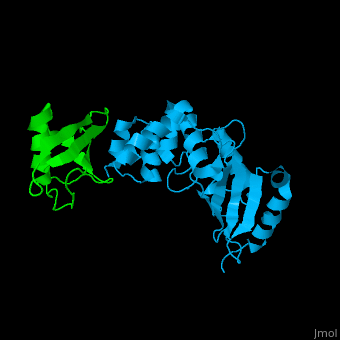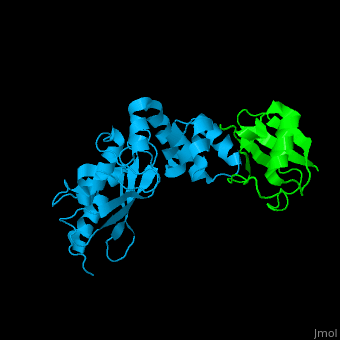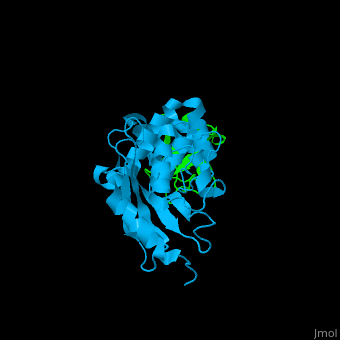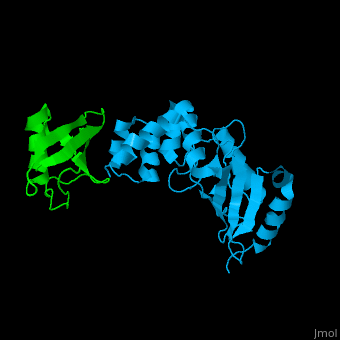
<StructureSection load='' size='340' side='right' caption='Human ubiquitin (green) complex with ubiquitin-conjugating enzyme E2 (deep sky blue), [[3k9p]]' scene='41/417541/Cv/3' > | <StructureSection load='' size='340' side='right' caption='Human ubiquitin (green) complex with ubiquitin-conjugating enzyme E2 (deep sky blue), [[3k9p]]' scene='41/417541/Cv/3' > | ||
== Function == | == Function == | ||
| - | [[Ubiquitin]] (UBB) is found in almost all cells. It binds to proteins tagging them for destruction in the proteasome. UBB is activated by the UBB-activating enzymes E1, E2 and E3. UBB+1 is a frameshifted mutant of UBB observed in several diseases. A dimer of UBB (DiUBB) is formed by linkage of K48 to the C-terminus of a second UBB molecule. '''Polyubiquitin''' (polyUBB) is a chain of ubiquitin bound by peptide bonds. At least 4 UBB molecules are needed to tag a protein for the proteasome<ref>PMID:9759494</ref>. <scene name='41/417541/Cv/4'>Human ubiquitin interactions with ubiquitin-conjugating enzyme E2</scene> ([[3k9p]]). For details see<br /> | + | [[Ubiquitin]] (UBB) is found in almost all cells. It binds to proteins tagging them for destruction in the proteasome. UBB is activated by the UBB-activating enzymes E1, E2 and E3. UBB+1 is a frameshifted mutant of UBB observed in several diseases. A dimer of UBB (DiUBB) is formed by linkage of K48 to the C-terminus of a second UBB molecule. '''Polyubiquitin''' (polyUBB) is a chain of ubiquitin molecules bound by peptide bonds. PolyUBB can be formed by lysine residues: K6, K11, K27, 29, K33, K48, and K63. Different lysine linkages convey different functions to polyUBB. Lys48- linked polyUBB and LYs11-linked polyUBB are associated with proteasome degradation; Lys63-linked and Lys-6 polyUBB are associated with non-proteolytic functions<ref>PMID:18438605</ref>. At least 4 UBB molecules are needed to tag a protein for the proteasome<ref>PMID:9759494</ref>. <scene name='41/417541/Cv/4'>Human ubiquitin interactions with ubiquitin-conjugating enzyme E2</scene> ([[3k9p]]). For details see<br /> |
* [[Ubiquitin Structure & Function]]<br /> | * [[Ubiquitin Structure & Function]]<br /> | ||
* [[Ubiquitin and Ubiquitination]]<br /> | * [[Ubiquitin and Ubiquitination]]<br /> | ||
Current revision
| |||||||||||
References
- ↑ Li W, Ye Y. Polyubiquitin chains: functions, structures, and mechanisms. Cell Mol Life Sci. 2008 Aug;65(15):2397-406. PMID:18438605 doi:10.1007/s00018-008-8090-6
- ↑ Hershko A, Ciechanover A. The ubiquitin system. Annu Rev Biochem. 1998;67:425-79. PMID:9759494 doi:http://dx.doi.org/10.1146/annurev.biochem.67.1.425
- ↑ Neefjes J, Groothuis TA, Dantuma NP. [The 2004 Nobel Prize in Chemistry for the discovery of ubiquitin-mediated protein degradation]. Ned Tijdschr Geneeskd. 2004 Dec 25;148(52):2579-82 PMID:15646859
- ↑ Paul S. Dysfunction of the ubiquitin-proteasome system in multiple disease conditions: therapeutic approaches. Bioessays. 2008 Nov;30(11-12):1172-84. doi: 10.1002/bies.20852. PMID:18937370 doi:http://dx.doi.org/10.1002/bies.20852
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Joel L. Sussman, David Canner, Jaime Prilusky