This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Triose Phosphate Isomerase Structure & Mechanism
From Proteopedia
| Line 1: | Line 1: | ||
==Triose Phosphate Isomerase (TIM)== | ==Triose Phosphate Isomerase (TIM)== | ||
| + | ===General Information=== | ||
Triose phosphate isomerase (TIM)<ref>PMID:16511037</ref><ref>PMID:8061610</ref> (PDB [[1wyi]] and [[1hti]]) is a crucial enzyme in the glycolytic pathway. <scene name='Christian_Krenk_Sandbox/Nc_rainbow/1'>TIM</scene> reversibly converts the aldose Glyceraldehyde-3-phosphate (GAP) to the ketose Dihydroxyacetone phosphate (DHAP). The interconversion proceeds by an enediol intermediate. Triose phosphate isomerase is not directly regulated, but the enzyme two steps before it in the glycolytic pathway, phosphofructokinase, is a heavily regulated, irreversible enzyme. | Triose phosphate isomerase (TIM)<ref>PMID:16511037</ref><ref>PMID:8061610</ref> (PDB [[1wyi]] and [[1hti]]) is a crucial enzyme in the glycolytic pathway. <scene name='Christian_Krenk_Sandbox/Nc_rainbow/1'>TIM</scene> reversibly converts the aldose Glyceraldehyde-3-phosphate (GAP) to the ketose Dihydroxyacetone phosphate (DHAP). The interconversion proceeds by an enediol intermediate. Triose phosphate isomerase is not directly regulated, but the enzyme two steps before it in the glycolytic pathway, phosphofructokinase, is a heavily regulated, irreversible enzyme. | ||
| - | ===Structural Characteristics | + | ===Structural Characteristics=== |
The secondary structure consists of 14 alpha helices and 8 beta sheets per monomer, making it fall in the SCOP category of alpha and beta proteins. The tertiary structure is a <scene name='Christian_Krenk_Sandbox/Alpha_beta_barrel/2'>alpha-beta barrel.</scene> | The secondary structure consists of 14 alpha helices and 8 beta sheets per monomer, making it fall in the SCOP category of alpha and beta proteins. The tertiary structure is a <scene name='Christian_Krenk_Sandbox/Alpha_beta_barrel/2'>alpha-beta barrel.</scene> | ||
The quaternary structure is a homodimer. The molecular weight of the enzyme is estimated at 57,400 Da.<ref>PMID:752201</ref> | The quaternary structure is a homodimer. The molecular weight of the enzyme is estimated at 57,400 Da.<ref>PMID:752201</ref> | ||
| Line 9: | Line 10: | ||
| - | ===Mechanism | + | ===Mechanism=== |
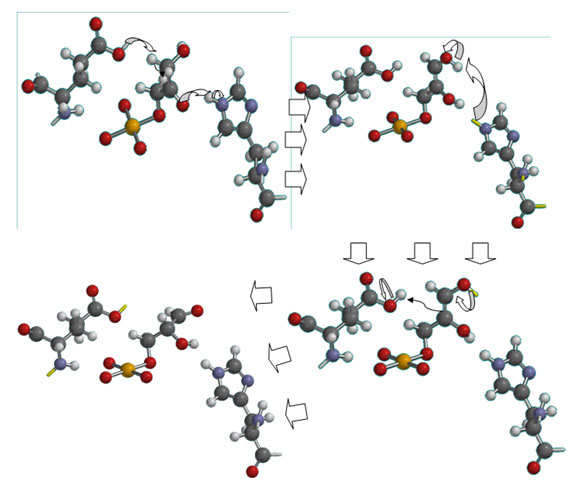
The enzyme aids in catalysis by binding tightly to the enediol transition state. To convert GAP to the enediol intermediate, a proton is abstracted from C2 by a base and the carbonyl oxygen atom is protonated by an acid.<ref>Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 495. Print.</ref> | The enzyme aids in catalysis by binding tightly to the enediol transition state. To convert GAP to the enediol intermediate, a proton is abstracted from C2 by a base and the carbonyl oxygen atom is protonated by an acid.<ref>Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 495. Print.</ref> | ||
Revision as of 15:14, 30 March 2010
Contents |
Triose Phosphate Isomerase (TIM)
General Information
Triose phosphate isomerase (TIM)[1][2] (PDB 1wyi and 1hti) is a crucial enzyme in the glycolytic pathway. reversibly converts the aldose Glyceraldehyde-3-phosphate (GAP) to the ketose Dihydroxyacetone phosphate (DHAP). The interconversion proceeds by an enediol intermediate. Triose phosphate isomerase is not directly regulated, but the enzyme two steps before it in the glycolytic pathway, phosphofructokinase, is a heavily regulated, irreversible enzyme.
Structural Characteristics
The secondary structure consists of 14 alpha helices and 8 beta sheets per monomer, making it fall in the SCOP category of alpha and beta proteins. The tertiary structure is a The quaternary structure is a homodimer. The molecular weight of the enzyme is estimated at 57,400 Da.[3]
Mechanism
The enzyme aids in catalysis by binding tightly to the enediol transition state. To convert GAP to the enediol intermediate, a proton is abstracted from C2 by a base and the carbonyl oxygen atom is protonated by an acid.[4] The enediol intermediate is negatively charged, but is somewhat [5] Mutation of Lys 12 to Arg increases Km by a factor of 22 and decreases Vmax by a factor of 180.[6] To convert the enediol intermediate to DHAP, C1 is protonated by Glu 165, with His 95 removing a proton from C2’s OH group. As a result, the catalytic groups are back to their original states, and catalysis is complete. With GAP as a substrate, Km for the reaction is .34 mM and Vmax is 7200 units/mg protein at 25 degrees C and pH 7.5.[7]
An interesting part of the enzyme is the that stabilizes the enediol-like transition state. The flexible loop (residues 167-176)[8] closes over the active site like a hinged lid when substrate is bound, thus preventing phosphate from leaving. A four-residue segment of the loop H-bonds with the phosphate group of the substrate.[9] Without the loop, the enediol intermediate would eliminate phosphate, with the end products being inorganic phosphate and toxic methylglyoxal.[10]
| |||||||||
| 1wyi, resolution 2.20Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Activity: | Triose-phosphate isomerase, with EC number 5.3.1.1 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
| |||||||||
| 1hti, resolution 2.80Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Activity: | Triose-phosphate isomerase, with EC number 5.3.1.1 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
References
- ↑ Kinoshita T, Maruki R, Warizaya M, Nakajima H, Nishimura S. Structure of a high-resolution crystal form of human triosephosphate isomerase: improvement of crystals using the gel-tube method. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2005 Apr 1;61(Pt, 4):346-9. Epub 2005 Mar 24. PMID:16511037 doi:10.1107/S1744309105008341
- ↑ Mande SC, Mainfroid V, Kalk KH, Goraj K, Martial JA, Hol WG. Crystal structure of recombinant human triosephosphate isomerase at 2.8 A resolution. Triosephosphate isomerase-related human genetic disorders and comparison with the trypanosomal enzyme. Protein Sci. 1994 May;3(5):810-21. PMID:8061610
- ↑ Dabrowska A, Kamrowska I, Baranowski T. Purification, crystallization and properties of triosephosphate isomerase from human skeletal muscle. Acta Biochim Pol. 1978;25(3):247-56. PMID:752201
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 495. Print.
- ↑ Lodi PJ, Chang LC, Knowles JR, Komives EA. Triosephosphate isomerase requires a positively charged active site: the role of lysine-12. Biochemistry. 1994 Mar 15;33(10):2809-14. PMID:8130193
- ↑ Lodi PJ, Chang LC, Knowles JR, Komives EA. Triosephosphate isomerase requires a positively charged active site: the role of lysine-12. Biochemistry. 1994 Mar 15;33(10):2809-14. PMID:8130193
- ↑ Dabrowska A, Kamrowska I, Baranowski T. Purification, crystallization and properties of triosephosphate isomerase from human skeletal muscle. Acta Biochim Pol. 1978;25(3):247-56. PMID:752201
- ↑ Lolis E, Petsko GA. Crystallographic analysis of the complex between triosephosphate isomerase and 2-phosphoglycolate at 2.5-A resolution: implications for catalysis. Biochemistry. 1990 Jul 17;29(28):6619-25. PMID:2204418
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 495. Print.
- ↑ Voet, Donald, Judith G. Voet, and Charlotte W. Pratt. Fundamentals of Biochemistry Life at the Molecular Level. New York: John Wiley & Sons, 2008. p. 495. Print.
Proteopedia Page Contributors and Editors (what is this?)
Christian Krenk, Alexander Berchansky, Diamond B. Reese, Michal Harel, Jane S. Richardson, David Canner