This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Factor Xa
From Proteopedia
(→Low Barrier Hydrogen Bonds) |
(→Structure) |
||
| Line 6: | Line 6: | ||
==Structure== | ==Structure== | ||
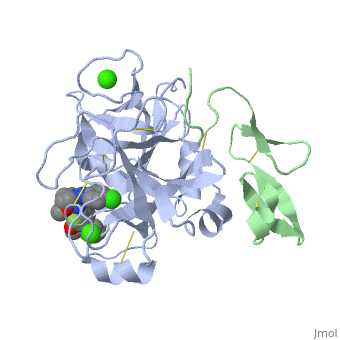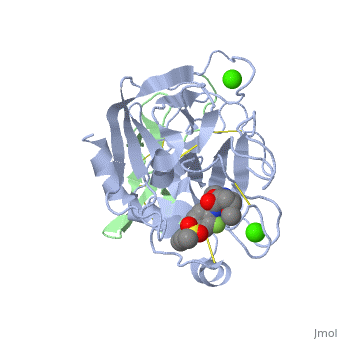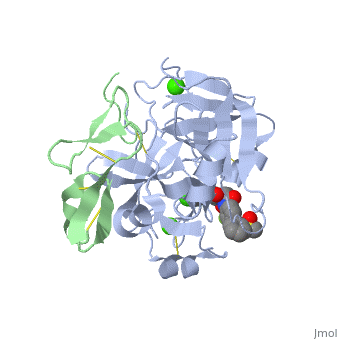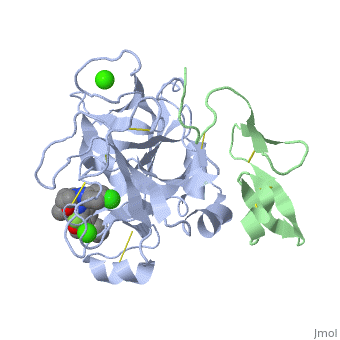
| - | <applet load=' | + | <applet load='2PR3' size='300' frame='true' align='right' caption='Structure of factor Xa PBD id: 2PR3' /> |
The factor Xa light chain contains a [http://en.wikipedia.org/wiki/Gla_domain γ-carboxyglutamic acid (Gla)] domain (11 gla residues) as well as two [http://en.wikipedia.org/wiki/Epidermal_growth_factor epidermal growth factor (EGF)]-like domains.<ref name="EGF">PMID:8355279</ref> The Gla domain is mediates calcium dependent binding of factor X to negatively charged phopholipid membrane surfaces. Recent crystal structures suggest that the N-terminal epidermal growth factor (EGF)-like domain is flexibly, while the second EGF domain maintains contacts with the catalytic domain. <ref name="EGF" /> | The factor Xa light chain contains a [http://en.wikipedia.org/wiki/Gla_domain γ-carboxyglutamic acid (Gla)] domain (11 gla residues) as well as two [http://en.wikipedia.org/wiki/Epidermal_growth_factor epidermal growth factor (EGF)]-like domains.<ref name="EGF">PMID:8355279</ref> The Gla domain is mediates calcium dependent binding of factor X to negatively charged phopholipid membrane surfaces. Recent crystal structures suggest that the N-terminal epidermal growth factor (EGF)-like domain is flexibly, while the second EGF domain maintains contacts with the catalytic domain. <ref name="EGF" /> | ||
Revision as of 16:58, 28 April 2010
Contents |
Introduction
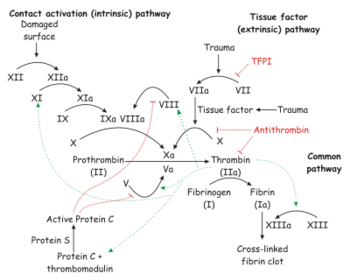
Factor X is a vitamin K-dependent glycoprotein that is synthesized in the liver. Zymogen factor X circulates in plasma as a 2 chain molecule composed of a disulfide linked light chain (Mr = 16500) and heavy chain (Mr = 42,000). Factor X is activated to factor Xa by cleavage of the activation peptide. This reaction is catalyzed by factor VIIa-tissue factor (extrinsic Xase complex) and factor IXa-factor VIIIa (intrinsic Xase complex).[1]
Factor Xa, along with factor Va, calcium, and a phospholipid membrane surface form the prothrombinase complex, to cleave prothrombin to its active form, thrombin.[1]
Structure
|
The factor Xa light chain contains a γ-carboxyglutamic acid (Gla) domain (11 gla residues) as well as two epidermal growth factor (EGF)-like domains.[2] The Gla domain is mediates calcium dependent binding of factor X to negatively charged phopholipid membrane surfaces. Recent crystal structures suggest that the N-terminal epidermal growth factor (EGF)-like domain is flexibly, while the second EGF domain maintains contacts with the catalytic domain. [2]
The factor Xa heavy chain contains the activation peptide and trypsin-like serine protease domain. [3]
Catalytic Triad
Serine proteases use a Ser-His-Asp catalytic triad, each playing an important role. The serine donates an OH group to act as a nucleophile and attack the carbonyl group of the peptide bond that will be broken within the substrate. Histidine coordinates the attack of the peptide bond by accepting the hydrogen from the serine –OH group with a pair of electrons on nitrogen. Aspartate contains a carboxyl group to hydrogen bond with the histidine, in order to properly position His, as well as to help with stabilization.
Substrate Recognition Sites
The determines binding selectivity for factor Xa. The natural substrate of factor Xa is prothromin, which is cleaved after the arginine in the sequence: Ile12-Asp13-Gly14-Arg15-Ile16- Val17-Glu18-Gly19. Arg15 binds in the S1 pocket, Gly14 binds the S2 pocket. The S1 pocket is formed by loops in residues 214-220 and 189-195 that are linked by a Cys220-Cys191 disulfide bond. Residues 225-228 form the lower portion of the pocket.[4] The is formed by the backbone amides of Gly193 and Ser195.[5] The oxyanion hole uses its main chain amide groups to stabilize the tetrahedral intermediate.[6]
The of factor Xa is formed by the 90s loop which is positioned adjacent to His57. Consistent with glycine as the P2 element in prothrombin, S2 is a small, shallow pocket.
S4 pocket is formed between the 90s and 170s loops and binds an Ile. This region contains 3 ligand binding domains. The is located at the entrance to S4 and contains Phe174, Tyr99 and Trp215, which form a deep aryl-binding pocket. The is formed by the backbone carbonyl and side chain of Glu97 (green) and the backbone carbonyl (red) of Lys96. The is composed of the hydrophobic side chains of Thr98, Ile175 and Thr177 and traps a water molecule. [3]
Enzyme Mechanism
General Serine Protease Mechanism
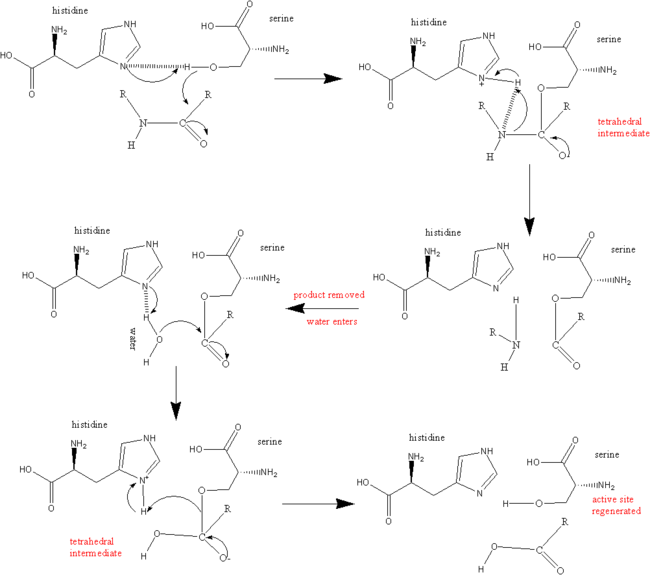
During the acylation half of the reaction His57 acts as a general base to remove a proton from Ser195, allowing it to attack the carbonyl of the peptide bond to be broken within the substrate, to yield the first tetrahedral intermediate. The negative oxygen ion of the tetrahedral intermediate is stabilized through hydrogen bonding with the oxyanion hole (Gly192 and Ser195). Asp102 stabilizes the protonated His57 through hydrogen bonding. His57 protonates the amine of the scissile bond, promoting formation of the acylenzyme and release of the C-terminal portion of the substrate.
The deacylation portion repeats the same sequence. A water molecule is deprotonated by His57 and attacks the acyl enzyme, to yielding a second tetrahedral intermediate. Again, the tetrahedral intermediate is stabilized by the oxyanion hole. Upon collapse of the tetrahedral intermediate, the N-terminal portion of the protein is released.[6]Controversial Mechanisms
His Flip Mechanisms
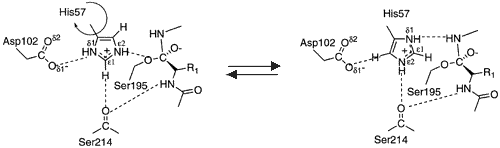
However, there are several arguments against the His flip mechanism. Flipping of His57 would require breaking and reforming many hydrogen bonds while the short lived tetrahedral intermediate is present. Also, His57 is sterically hindered by the P2 and P1’ residues of the peptide substrates. [10]These observations disfavor the His flip mechanism.
Other observations have suggested that Ser195 must move at least 1Å in order to form the tetrahedral intermediate. This conformation change would cause Ser195 and His57 to be oriented away from each other upon formation of the tetrahedral intermediate, preventing reprotonation. [6]
Low Barrier Hydrogen Bonds
| |||||||||||
Proteopedia Page Contributors and Editors (what is this?)
Jacqueline Gertz, Michal Harel, Alexander Berchansky, David Canner, Jaime Prilusky