This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Wayne Decatur/kink-turn motif
From Proteopedia
m |
m |
||
| Line 21: | Line 21: | ||
<applet load='Image:Kt7.pdb.gz' size='400' frame='true' align='right' scene='User:Wayne_Decatur/kink-turn_motif/Kt7_halo/2' /> | <applet load='Image:Kt7.pdb.gz' size='400' frame='true' align='right' scene='User:Wayne_Decatur/kink-turn_motif/Kt7_halo/2' /> | ||
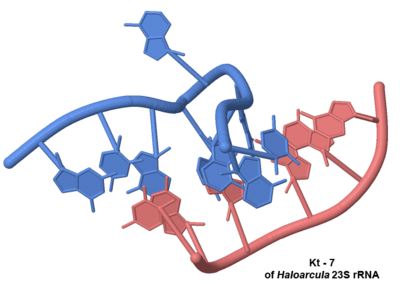
* [[Large Ribosomal Subunit of Haloarcula|The Large Ribosomal Subunit]] contains 8 identified kink-turns: <scene name='User:Wayne_Decatur/kink-turn_motif/Kt7_halo/2'>Kt-7</scene>,Kt-58 of [[3cc2]]. MAKE '''The location of all of them in the subunit.''' To see them in greater detail following the appropriate links on [http://www.dundee.ac.uk/biocentre/nasg/kturn/kturns_known.php this page at a structural database for k-turn motifs in RNA by the Lilley group]<ref name="lilleydatabase">PMID: 20562215</ref>. | * [[Large Ribosomal Subunit of Haloarcula|The Large Ribosomal Subunit]] contains 8 identified kink-turns: <scene name='User:Wayne_Decatur/kink-turn_motif/Kt7_halo/2'>Kt-7</scene>,Kt-58 of [[3cc2]]. MAKE '''The location of all of them in the subunit.''' To see them in greater detail following the appropriate links on [http://www.dundee.ac.uk/biocentre/nasg/kturn/kturns_known.php this page at a structural database for k-turn motifs in RNA by the Lilley group]<ref name="lilleydatabase">PMID: 20562215</ref>. | ||
| - | * | + | * The human spliceosomal and small nucleolar RNA-binding 15.5kD protein bound to a U4 spliceosomal RNA fragment ([[1e7k]])<ref>PMID: 11163207</ref> |
| - | * | + | * ''A. fulgidus'' small ribonucleoprotein particle box C/D RNA complexed with L7Ae ([[1rlg]])<ref>PMID: 15130473</ref> |
| - | * | + | * ''Pyrococcus furiosus'' small ribonucleoprotein particle H/ACA box RNA [[2hvy]])<ref>PMID: 16943774</ref> |
| - | * [[1u6b]], [[1zzn]], [[3bo2]], [[3bo3]], [[3bo4]], and [[3iin]] | + | * ''Azoarcus'' group I intron has a 'reverse' kink-turn'''PUT A SCENE HERE OF ALIGNMENT OF THIS WITH KT-7 with each Kt colored differently [[1u6b]], [[1zzn]], [[3bo2]], [[3bo3]], [[3bo4]], and [[3iin]]<ref>PMID: 15175762</ref><ref>PMID: 16141079</ref><ref> PMID:18408159</ref><ref>PMID:20145044</ref> |
| - | * | + | * ''S. cervisiae'' L30e bound to its pre-mRNA [[1t0k]]<ref>PMID: 15242593</ref> |
* [[1nkw]] – The Large Ribosomal Subunit From ''Deinococcus radiodurans''<ref> PMID:11733066</ref> | * [[1nkw]] – The Large Ribosomal Subunit From ''Deinococcus radiodurans''<ref> PMID:11733066</ref> | ||
* Small Ribosomal Subunit | * Small Ribosomal Subunit | ||
Revision as of 01:38, 2 August 2010
WHEN MADE INTO AN OFFICIAL PAGE:
- LINK FROM RNA motifs PAGE!!!!
- Fix link from topic pages and from table of contents
- add Redirect from K-turn motif
- add redirect from GA motif
- add redirect from kink-turn
- add redirect from k-turn
The kink-turn motif
A common RNA structural motif that consists of helix–internal loop–helix motif .
Contents |
Introduction
Originally identified in the course of analyzing the large ribosomal subunit[1], the motif was identified in other RNAs. Particular instances are referred to as the the GA motif [2] A motif that includes the A-minor motif. Many K-turns bind proteins; however, they also mediate RNA tertiary structure interactions.
Structures Containing the Motif
|
- The Large Ribosomal Subunit contains 8 identified kink-turns: ,Kt-58 of 3cc2. MAKE The location of all of them in the subunit. To see them in greater detail following the appropriate links on this page at a structural database for k-turn motifs in RNA by the Lilley group[3].
- The human spliceosomal and small nucleolar RNA-binding 15.5kD protein bound to a U4 spliceosomal RNA fragment (1e7k)[4]
- A. fulgidus small ribonucleoprotein particle box C/D RNA complexed with L7Ae (1rlg)[5]
- Pyrococcus furiosus small ribonucleoprotein particle H/ACA box RNA 2hvy)[6]
- Azoarcus group I intron has a 'reverse' kink-turnPUT A SCENE HERE OF ALIGNMENT OF THIS WITH KT-7 with each Kt colored differently 1u6b, 1zzn, 3bo2, 3bo3, 3bo4, and 3iin[7][8][9][10]
- S. cervisiae L30e bound to its pre-mRNA 1t0k[11]
- 1nkw – The Large Ribosomal Subunit From Deinococcus radiodurans[12]
- Small Ribosomal Subunit
See Also
- Kink-turns in RNA - a structural database for k-turn motifs in RNA by the Lilley group[3]
- Ribosome
- 1go1, 1go0, 1w3e and 1h7m – the Thermococcus celer ribosomal protein L30[13][14]
- A-minor motif
- The adenosine wedge motif[15]
- The G-ribo motif[16]
- The lonepair triloop motif[17]
- RNA ribose zipper[18]
References
- ↑ Klein DJ, Schmeing TM, Moore PB, Steitz TA. The kink-turn: a new RNA secondary structure motif. EMBO J. 2001 Aug 1;20(15):4214-21. PMID:11483524 doi:http://dx.doi.org/10.1093/emboj/20.15.4214
- ↑ Winkler WC, Grundy FJ, Murphy BA, Henkin TM. The GA motif: an RNA element common to bacterial antitermination systems, rRNA, and eukaryotic RNAs. RNA. 2001 Aug;7(8):1165-72. PMID:11497434
- ↑ 3.0 3.1 Schroeder KT, McPhee SA, Ouellet J, Lilley DM. A structural database for k-turn motifs in RNA. RNA. 2010 Aug;16(8):1463-8. Epub 2010 Jun 18. PMID:20562215 doi:10.1261/rna.2207910
- ↑ Vidovic I, Nottrott S, Hartmuth K, Luhrmann R, Ficner R. Crystal structure of the spliceosomal 15.5kD protein bound to a U4 snRNA fragment. Mol Cell. 2000 Dec;6(6):1331-42. PMID:11163207
- ↑ Moore T, Zhang Y, Fenley MO, Li H. Molecular basis of box C/D RNA-protein interactions; cocrystal structure of archaeal L7Ae and a box C/D RNA. Structure. 2004 May;12(5):807-18. PMID:15130473 doi:http://dx.doi.org/10.1016/j.str.2004.02.033
- ↑ Li L, Ye K. Crystal structure of an H/ACA box ribonucleoprotein particle. Nature. 2006 Sep 21;443(7109):302-7. Epub 2006 Aug 30. PMID:16943774 doi:http://dx.doi.org/10.1038/nature05151
- ↑ Adams PL, Stahley MR, Kosek AB, Wang J, Strobel SA. Crystal structure of a self-splicing group I intron with both exons. Nature. 2004 Jul 1;430(6995):45-50. Epub 2004 Jun 2. PMID:15175762 doi:10.1038/nature02642
- ↑ Stahley MR, Strobel SA. Structural evidence for a two-metal-ion mechanism of group I intron splicing. Science. 2005 Sep 2;309(5740):1587-90. PMID:16141079 doi:309/5740/1587
- ↑ Lipchock SV, Strobel SA. A relaxed active site after exon ligation by the group I intron. Proc Natl Acad Sci U S A. 2008 Apr 15;105(15):5699-704. Epub 2008 Apr 11. PMID:18408159
- ↑ Antonioli AH, Cochrane JC, Lipchock SV, Strobel SA. Plasticity of the RNA kink turn structural motif. RNA. 2010 Apr;16(4):762-8. Epub 2010 Feb 9. PMID:20145044 doi:10.1261/rna.1883810
- ↑ Chao JA, Williamson JR. Joint X-ray and NMR refinement of the yeast L30e-mRNA complex. Structure. 2004 Jul;12(7):1165-76. PMID:15242593 doi:10.1016/j.str.2004.04.023
- ↑ Harms J, Schluenzen F, Zarivach R, Bashan A, Gat S, Agmon I, Bartels H, Franceschi F, Yonath A. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell. 2001 Nov 30;107(5):679-88. PMID:11733066
- ↑ Chen YW, Bycroft M, Wong KB. Crystal structure of ribosomal protein L30e from the extreme thermophile Thermococcus celer: thermal stability and RNA binding. Biochemistry. 2003 Mar 18;42(10):2857-65. PMID:12627951 doi:10.1021/bi027131s
- ↑ Wong KB, Lee CF, Chan SH, Leung TY, Chen YW, Bycroft M. Solution structure and thermal stability of ribosomal protein L30e from hyperthermophilic archaeon Thermococcus celer. Protein Sci. 2003 Jul;12(7):1483-95. PMID:12824494 doi:10.1110/ps.0302303
- ↑ Gagnon MG, Steinberg SV. The adenosine wedge: a new structural motif in ribosomal RNA. RNA. 2010 Feb;16(2):375-81. Epub 2009 Dec 28. PMID:20038632 doi:10.1261/rna.1550310
- ↑ Steinberg SV, Boutorine YI. G-ribo: a new structural motif in ribosomal RNA. RNA. 2007 Apr;13(4):549-54. Epub 2007 Feb 5. PMID:17283211 doi:10.1261/rna.387107
- ↑ Lee JC, Cannone JJ, Gutell RR. The lonepair triloop: a new motif in RNA structure. J Mol Biol. 2003 Jan 3;325(1):65-83. PMID:12473452
- ↑ Tamura M, Holbrook SR. Sequence and structural conservation in RNA ribose zippers. J Mol Biol. 2002 Jul 12;320(3):455-74. PMID:12096903
Additional Literature and Resources
- Moore PB. The ribosome returned. J Biol. 2009;8(1):8. Epub 2009 Jan 26. PMID:19222865 doi:10.1186/jbiol103
- Steitz TA. A structural understanding of the dynamic ribosome machine. Nat Rev Mol Cell Biol. 2008 Mar;9(3):242-53. PMID:18292779 doi:10.1038/nrm2352
- Rodnina MV, Wintermeyer W. The ribosome goes Nobel. Trends Biochem Sci. 2010 Jan;35(1):1-5. Epub 2009 Dec 2. PMID:19962317 doi:10.1016/j.tibs.2009.11.003
- Schmeing TM, Ramakrishnan V. What recent ribosome structures have revealed about the mechanism of translation. Nature. 2009 Oct 29;461(7268):1234-42. Epub 2009 Oct 18. PMID:19838167 doi:10.1038/nature08403
- Ramakrishnan V, Moore PB. Atomic structures at last: the ribosome in 2000. Curr Opin Struct Biol. 2001 Apr;11(2):144-54. PMID:11297922
- Schroeder KT, McPhee SA, Ouellet J, Lilley DM. A structural database for k-turn motifs in RNA. RNA. 2010 Aug;16(8):1463-8. Epub 2010 Jun 18. PMID:20562215 doi:10.1261/rna.2207910
- RCSB Protein Data Bank coverage of the 2009 Nobel Prizes in Chemistry
- 70S Ribosome: January 2010 Molecule of the Month as part of the series of tutorials that are at the RCSB Protein Data Bank and written by David Goodsell
- Ribosome: October 2000 Molecule of the Month as part of the series of tutorials that are at the RCSB Protein Data Bank and written by David Goodsell