This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Sandbox Reserved 329
From Proteopedia
| Line 8: | Line 8: | ||
== INTRODUCTION == | == INTRODUCTION == | ||
| - | Terminal uridylyl transferases (TUTases) belong to a superfamily of polymerase ß nucleotidyl transferases.<ref name="primary citation">PMID:17785418</ref> TUTases have been isolated from ''Trypanosoma brucei'' and also ''Leishmania'' ssp, parasites causing diseases in humans such as African Sleeping Sickness.<ref>PMID:11893335</ref> TUTases function in RNA editing; more specifically | + | Terminal uridylyl transferases (TUTases) belong to a superfamily of polymerase ß nucleotidyl transferases.<ref name="primary citation">PMID:17785418</ref> TUTases have been isolated from ''Trypanosoma brucei'' and also ''Leishmania'' ssp, parasites causing diseases in humans such as African Sleeping Sickness.<ref>PMID:11893335</ref> TUTases can function in RNA editing; more specifically TUTase4 catalyzes a reaction that adds a uridine monophosphate (UMP) from UTP to a RNA substrate. Trypanosomal TUTases have RNA substrates that are shown to select for cognate nucleosides and provide a metal ion binding site for Mg<sup>2+</sup> ions.<ref name="primary citation">PMID:17785418</ref> |
{{STRUCTURE_2q0d | PDB=2q0d | SCENE=Reserved_Sandbox_329/Scene1/1}} | {{STRUCTURE_2q0d | PDB=2q0d | SCENE=Reserved_Sandbox_329/Scene1/1}} | ||
Revision as of 23:43, 30 March 2011
| This Sandbox is Reserved from January 10, 2010, through April 10, 2011 for use in BCMB 307-Proteins course taught by Andrea Gorrell at the University of Northern British Columbia, Prince George, BC, Canada. |
To get started:
More help: Help:Editing |
Contents |
Uridylyl transferases
INTRODUCTION
Terminal uridylyl transferases (TUTases) belong to a superfamily of polymerase ß nucleotidyl transferases.[1] TUTases have been isolated from Trypanosoma brucei and also Leishmania ssp, parasites causing diseases in humans such as African Sleeping Sickness.[2] TUTases can function in RNA editing; more specifically TUTase4 catalyzes a reaction that adds a uridine monophosphate (UMP) from UTP to a RNA substrate. Trypanosomal TUTases have RNA substrates that are shown to select for cognate nucleosides and provide a metal ion binding site for Mg2+ ions.[1]
| |||||||||
| 2q0d, resolution 2.00Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , | ||||||||
| Gene: | TUT4 (Trypanosoma brucei) | ||||||||
| Activity: | RNA uridylyltransferase, with EC number 2.7.7.52 | ||||||||
| Related: | 2ikf, 2nom | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
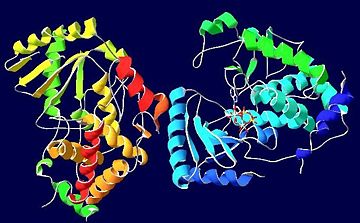
STRUCTURE
The uridylyl transferase bound ligand is an with two Mg2+ ions, however many TUTases involved in RNA editing are shown to exhibit preference for binding to UTP instead.[1] Three are conserved in TUTases, and are required for coordinating the Mg2+ ions in some TUTases. [1] Thus, these are vital in catalyzing this reaction.
REFERENCES
- ↑ 1.0 1.1 1.2 1.3 Stagno J, Aphasizheva I, Aphasizhev R, Luecke H. Dual role of the RNA substrate in selectivity and catalysis by terminal uridylyl transferases. Proc Natl Acad Sci U S A. 2007 Sep 11;104(37):14634-9. Epub 2007 Sep 4. PMID:17785418
- ↑ Aphasizhev R, Sbicego S, Peris M, Jang SH, Aphasizheva I, Simpson AM, Rivlin A, Simpson L. Trypanosome mitochondrial 3' terminal uridylyl transferase (TUTase): the key enzyme in U-insertion/deletion RNA editing. Cell. 2002 Mar 8;108(5):637-48. PMID:11893335