Ribonuclease A Catalysis
From Proteopedia
| Line 8: | Line 8: | ||
==Introduction== | ==Introduction== | ||
| - | Acid Base Catalysis is the acceleration of a chemical reaction by the addition of an acid or a base and is mainly used in organic chemical reactions. The acid or base is not consumed in the reaction itself. An acid transfers protons to a reactant and a base accepts protons from the reactant. An acid is often thought of as a proton and the base as a hydroxyl. When the acids or bases donate or accept protons, they stabilize the developing charges in the transition state. This usually creates a better leaving group, making the reaction more | + | Acid Base Catalysis is the acceleration of a chemical reaction by the addition of an acid or a base and is mainly used in organic chemical reactions. The acid or base is not consumed in the reaction itself. An acid transfers protons to a reactant and a base accepts protons from the reactant. An acid is often thought of as a proton and the base as a hydroxyl. When the acids or bases donate or accept protons, they stabilize the developing charges in the transition state. This usually creates a better leaving group, making the reaction more energetically favorable. Additionally, this has an effect on the activity of the nucleophile and electrophile groups. Histidine is a very common residue involved in acid-base cataylsis due to the fact that is has a pKa close to neutral; therefore, it can both accept and donate protons. |
Revision as of 15:38, 31 March 2011
| This Sandbox is Reserved from Feb 02, 2011, through Jul 31, 2011 for use by the Biochemistry II class at the Butler University at Indianapolis, IN USA taught by R. Jeremy Johnson. This reservation includes Sandbox Reserved 191 through Sandbox Reserved 200. |
To get started:
More help: Help:Editing |
Contents |
Ribonuclease A Catalysis
| |||||||||
| 3cin, resolution 1.70Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , , | ||||||||
| Gene: | TM1419, TM_1419 (Thermotoga maritima MSB8) | ||||||||
| Activity: | Inositol-3-phosphate synthase, with EC number 5.5.1.4 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum, TOPSAN | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Introduction
Acid Base Catalysis is the acceleration of a chemical reaction by the addition of an acid or a base and is mainly used in organic chemical reactions. The acid or base is not consumed in the reaction itself. An acid transfers protons to a reactant and a base accepts protons from the reactant. An acid is often thought of as a proton and the base as a hydroxyl. When the acids or bases donate or accept protons, they stabilize the developing charges in the transition state. This usually creates a better leaving group, making the reaction more energetically favorable. Additionally, this has an effect on the activity of the nucleophile and electrophile groups. Histidine is a very common residue involved in acid-base cataylsis due to the fact that is has a pKa close to neutral; therefore, it can both accept and donate protons.
The acid base mechanism can extensively alter the pKa depending on the environment of the residue. PKa will increase for an acidic residue if the environment is hydrophobic or if the adjacent residues are of similar charges. In the same environmental conditions, a basic residue will decrease the pKa. PKa will decrease for an acidic residue and increase for a basic residue if there is a salt bridge.
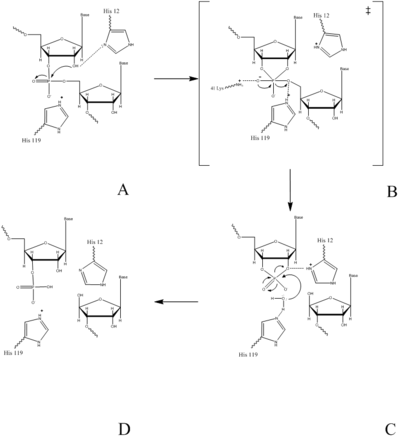
Mechanism
|
References
Works Cited
Proteopedia Page Contributors and Editors (what is this?)
Nathan Clarke, David Canner, Alexander Berchansky, R. Jeremy Johnson, OCA