Sandbox 35
From Proteopedia
| Line 15: | Line 15: | ||
| - | This protease belongs to an extended family of aminopeptidases, dipeptidyl peptidases, endopeptidases, and other enzymes having both exo- and endo-peptidase activity. The inactivated zymogen with N-terminal propeptide regions - providing stability in alkaline environments and enabling proper folding - is activated through removal of the propeptide regions <ref>PMID: 7845226</ref><ref>PMID: 12188906</ref>. | + | This protease belongs to an extended family of aminopeptidases, dipeptidyl peptidases, endopeptidases, and other enzymes having both exo- and endo-peptidase activity. The inactivated zymogen with N-terminal propeptide regions - providing stability in alkaline environments and enabling proper folding - is activated through removal of the propeptide regions <ref>PMID: 7845226</ref><ref>PMID: 12188906</ref>. Papain. Lights. Camera. Action! |
Revision as of 02:08, 14 November 2011
| Please do NOT make changes to this Sandbox. Sandboxes 30-60 are reserved for use by Biochemistry 410 & 412 at Messiah College taught by Dr. Hannah Tims during Fall 2012 and Spring 2013. |
Contents |
Papain
Introduction
DID YOU KNOW?
. Meat tenderizer. Old time home remedy for insect, jellyfish, and stingray stings[1]. Who would have thought that a sulfhydryl protease from the latex of the papaya fruit, Carica papaya and Vasconcellea cundinamarcensis would have such a practical application beyond proteopedia?
This protease belongs to an extended family of aminopeptidases, dipeptidyl peptidases, endopeptidases, and other enzymes having both exo- and endo-peptidase activity. The inactivated zymogen with N-terminal propeptide regions - providing stability in alkaline environments and enabling proper folding - is activated through removal of the propeptide regions [2][3]. Papain. Lights. Camera. Action!
|
Structure
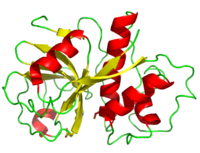
Papain's polypeptide chain consists of 212 amino acid residues which fold to form a groove containing the active site between its two domains. Its consists of 17 strands and 7 giving it a composition 21% and 25% respectively. [4] The hydrogen bonds within the alpha helices are shorter than the typical alpha helix because of C=O being directed further away from the helical axis. Moreover, the beta sheet hydrogen bonding constraints and structural angles show great variation; hydrogen bonds in the sheets central tend to be shorter than on the fringes. Three disulfide bonds (yellow) serve to hold papain's tertiary structure together.
The primarily consist of three main residues Cys25-His159-Asn175 that resemble the catalytic triad of chymotrypsin [5][6]. However growing studies are showing that the mechanism behind catalysis may actually involve a double catalytic site - consisting of Cys25-His159-Asn175 and Cys25-His159-
! It is postulated that "a two-state mechanism" takes place instead of a "single steric mechanism." [7] In addition, replacement of Asn 175 with other residues such as Ala mutants, reveals a decrease in kcat (less efficient), but the rate of hydrolysis is still significantly larger than non-catalytic rates suggesting a less essential role the residue plays than originally thought. It should be noted however, that alteration to 175 side chain resulted in less thermal stability lending thought to the residues structural conservative role rather than catalytic. [8]
Crystallization of the protease under conditions of 62% (w/w) methanol in water reveals water playing a crucial role in providing structural stability. The 21 internal water molecules surrounding adjacent papain molecules appear to form an encasement that limit protein to protein interaction. [9]
Distribution of Residues
Papain has a scattered distribution of , but can be seen to have more basic residues than acidic, shedding understanding into the application of its use as a digestive supplement. [10] Its build on this picture with resting more on the outside and sequestering near the center. Observations have revealed that the proteins atomic positions are more ordered going from the center toward the outside [11]
Ligands interactions and Pseudo Substrates
Papain is said to have 29 methanol molecules that encircle around it as . The polarity of the ligands results in hydrogen bonding interaction. [12]
is part of a series of CLIK . [13] Primarily hydrogen bonds with non-water and hydrophobic interactions
Catalytic Mechanism
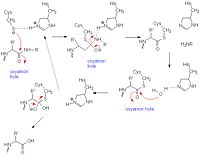
References
- ↑ [1] Ameridan International
- ↑ Rawlings ND, Barrett AJ. Families of cysteine peptidases. Methods Enzymol. 1994;244:461-86. PMID:7845226
- ↑ Yamamoto Y, Kurata M, Watabe S, Murakami R, Takahashi SY. Novel cysteine proteinase inhibitors homologous to the proregions of cysteine proteinases. Curr Protein Pept Sci. 2002 Apr;3(2):231-8. PMID:12188906
- ↑ [2]9PAP PDB
- ↑ Wang J, Xiang YF, Lim C. The double catalytic triad, Cys25-His159-Asp158 and Cys25-His159-Asn175, in papain catalysis: role of Asp158 and Asn175. Protein Eng. 1994 Jan;7(1):75-82. PMID:8140097
- ↑ Ménard R, Khouri HE, Plouffe C, Dupras R, Ripoll D, Vernet T, Tessier DC, Lalberté F, Thomas DY, Storer AC. A protein engineering study of the role of aspartate 158 in the catalytic mechanism of papain. Biochemistry. 1990 Jul 17;29(28):6706-13. PMID:2397208 doi:10.1021/bi00480a021
- ↑ Wang J, Xiang YF, Lim C. The double catalytic triad, Cys25-His159-Asp158 and Cys25-His159-Asn175, in papain catalysis: role of Asp158 and Asn175. Protein Eng. 1994 Jan;7(1):75-82. PMID:8140097
- ↑ [3] The Journal of Biological Chemistry
- ↑ Kamphuis IG, Kalk KH, Swarte MB, Drenth J. Structure of papain refined at 1.65 A resolution. J Mol Biol. 1984 Oct 25;179(2):233-56. PMID:6502713
- ↑ [4] WebMD
- ↑ Kamphuis IG, Kalk KH, Swarte MB, Drenth J. Structure of papain refined at 1.65 A resolution. J Mol Biol. 1984 Oct 25;179(2):233-56. PMID:6502713
- ↑ Kamphuis IG, Kalk KH, Swarte MB, Drenth J. Structure of papain refined at 1.65 A resolution. J Mol Biol. 1984 Oct 25;179(2):233-56. PMID:6502713
- ↑ Tsuge H, Nishimura T, Tada Y, Asao T, Turk D, Turk V, Katunuma N. Inhibition mechanism of cathepsin L-specific inhibitors based on the crystal structure of papain-CLIK148 complex. Biochem Biophys Res Commun. 1999 Dec 20;266(2):411-6. PMID:10600517 doi:10.1006/bbrc.1999.1830
- ↑ [5] University of Maine
http://www.pdb.org/pdb/explore/explore.do?structureId=2PAD
• Show the secondary structures.
• Compare the distribution of polar residues to that of nonpolar residues.
• Highlight the active site.
• If you can find a PDB file of the enzyme that contains a pseudo-substrate (may be inhibitor), highlight it.
• Show the contacts or attractions that are present between the pseudo-substrate and the protein, and if the enzyme has multiple subunits, show the contacts between the subunits.
• Identify any other ligands that are present in the structure and the types of contacts that are present between them and the protein
http://proteopedia.org/wiki/index.php/Sandbox_55#cite_note-18 Table of contents Pictures References (cross links)