This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
User:Brian Hernandez/DOPA Decarboxylase
From Proteopedia
(Difference between revisions)
| Line 11: | Line 11: | ||
==Function== | ==Function== | ||
==='''The Active Site'''=== | ==='''The Active Site'''=== | ||
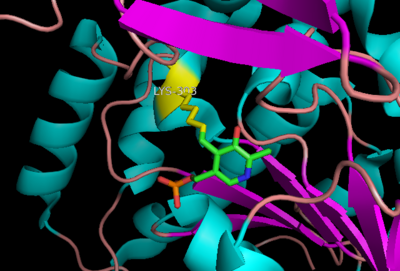
| - | DDC's active site is located in a <scene name='DOPA_decarboxylase/Dimer_interface/2'>cleft</scene> between the two monomer subunits, but is composed mainly of residues from one monomer.The <scene name='DOPA_decarboxylase/Active_site/1'>active site</scene> is composed of several key residues, including Lys-303, Asp-271, His-192, Thr-82, Ile-101, and Phe-103. In the ligand free form, PLP binds to Lys 303 via a '''Schiff base linkage'''. A [http://en.wikipedia.org/wiki/Salt_bridge_(protein) salt bridge] forms between the carboxylate group of Asp 271 and the protonated pyridine nitrogen of PLP yielding a strong '''electron sink''' capable of stabilizing the carbanionic intermediates. The only two active site residues from the adjacent monomer, Ile-101 and Phe-103, are part of the substrate binding pocket. | + | DDC's active site is located in a <scene name='DOPA_decarboxylase/Dimer_interface/2'>cleft</scene> between the two monomer subunits, but is composed mainly of residues from one monomer.The <scene name='DOPA_decarboxylase/Active_site/1'>active site</scene> is composed of several key residues, including Lys-303, Asp-271, His-192, Thr-82, Ile-101, and Phe-103. In the ligand free form, PLP binds to Lys 303 via a '''Schiff base linkage'''. A [http://en.wikipedia.org/wiki/Salt_bridge_(protein) salt bridge] forms between the carboxylate group of Asp 271 and the protonated pyridine nitrogen of PLP yielding a strong '''electron sink''' capable of stabilizing the carbanionic intermediates. The only two active site residues from the adjacent monomer, Ile-101 and Phe-103, are part of the substrate binding pocket that stabilizes the substrate via <scene name='DOPA_decarboxylase/Vanderwaals/1'>van der waals forces</scene>. |
[[image:plp bound.png|thumb|center|400px|'''Schiff base linkage of PLP to Lys303 in the active site''']] | [[image:plp bound.png|thumb|center|400px|'''Schiff base linkage of PLP to Lys303 in the active site''']] | ||
Revision as of 07:30, 29 November 2011
| |||||||||||
3D structures of DOPA decarboxylase
| |||||||||
| 1js3, resolution 2.25Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | , , | ||||||||
| Activity: | Aromatic-L-amino-acid decarboxylase, with EC number 4.1.1.28 | ||||||||
| Related: | 1js6 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
3k40 – DDC – Drosophila melanogaster
1js3 – pDDC + inhibitor – pig
1js6 - pDDC
3rbf, 3rbl – hDDC – human
3rch – hDDC + vitamin B6 phosphate + pyridoxal phosphate
References
- ↑ Christenson JG, Dairman W, Udenfriend S. On the identity of DOPA decarboxylase and 5-hydroxytryptophan decarboxylase (immunological titration-aromatic L-amino acid decarboxylase-serotonin-dopamine-norepinephrine). Proc Natl Acad Sci U S A. 1972 Feb;69(2):343-7. PMID:4536745
- ↑ Schneider G, Kack H, Lindqvist Y. The manifold of vitamin B6 dependent enzymes. Structure. 2000 Jan 15;8(1):R1-6. PMID:10673430
- ↑ 3.0 3.1 3.2 Burkhard P, Dominici P, Borri-Voltattorni C, Jansonius JN, Malashkevich VN. Structural insight into Parkinson's disease treatment from drug-inhibited DOPA decarboxylase. Nat Struct Biol. 2001 Nov;8(11):963-7. PMID:11685243 doi:http://dx.doi.org/10.1038/nsb1101-963
- ↑ Ishii S, Mizuguchi H, Nishino J, Hayashi H, Kagamiyama H. Functionally important residues of aromatic L-amino acid decarboxylase probed by sequence alignment and site-directed mutagenesis. J Biochem. 1996 Aug;120(2):369-76. PMID:8889823