This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Hemagglutinin
From Proteopedia
| Line 53: | Line 53: | ||
The substitutions of amino acids in these sites allow the influenza virus to escape from the immune system and to spread. Major changes in these antigenic regions can produce lethal influenza pandemics, like in 1957. | The substitutions of amino acids in these sites allow the influenza virus to escape from the immune system and to spread. Major changes in these antigenic regions can produce lethal influenza pandemics, like in 1957. | ||
| + | |||
| + | ==How Influenza Escapes Vaccines== | ||
| + | |||
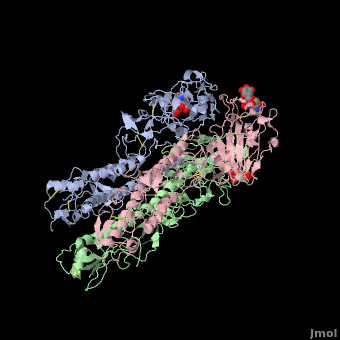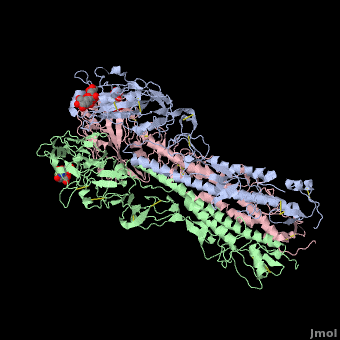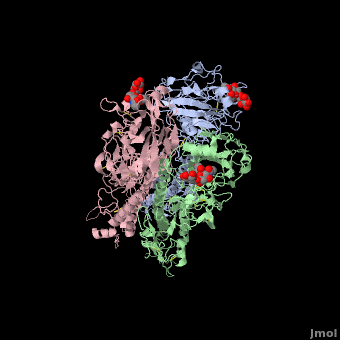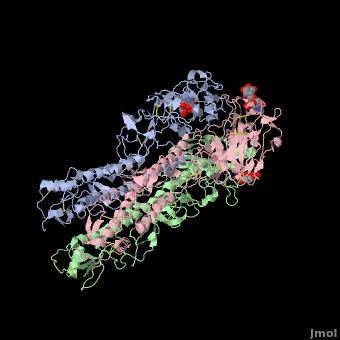
| + | Influenza hemagglutinin (''e.g.'' [[1hgf]]) is a glycoprotein on the surface of [http://en.wikipedia.org/wiki/Influenza_virus influenza virus] particles that enables them to attach to and infect host cells. [[Antibody|Antibodies]] that bind to hemagglutinin are a major defense mechanism that prevent infection. The RNA genome of influenza is characterized by a high mutation rate. Mutations on the surface of hemagglutinin tend to be protective for the virus. They tend to be retained because they tend to reduce the binding strength, and hence the host defensive capability, of antibodies that recognize the un-mutated hemagglutinin<ref name="skehel_review">PMID: 16925526</ref>. Influenza vaccines include hemagglutinins and they induce anti-hemagglutinin antibodies in vaccinated individuals. Often, however, by the time the vaccines can be designed, produced, and disseminated, mutant influenza viruses have arisen that can cause disease in vaccinated individuals<ref name="fluwikipedia">See [http://en.wikipedia.org/wiki/Influenza Influenza] in Wikipedia.</ref>. | ||
| + | If you '''show''' the '''Evolutionary Conservation''' of the structure on this page (using the blue bars below the molecule), you will see highly variable surface amino acids. These represent sites of mutations that have been retained in wild strains of influenza because they improve virus survival<ref name="skehel_review" />. (Added by [[User:Eric Martz|Eric Martz]]). | ||
==Notes== | ==Notes== | ||
Revision as of 08:42, 4 June 2012
| |||||||||||
Contents |
3D structures of hemagglutinin
Updated November 2011
3aj5, 3ah1, 3ah2, 3ah4 – CbHA1 + glucoside - Clostridium botulinum
1qxm - CbHA1 (mutant)
3aj6 - CbHA1 (mutant) + glucoside
2zoe, 2zs6 – CbHA3 + glucoside
2e4m – CbHA1 + HA17
Influenza hemagglutinin
3qqi, 3mlh - IvHA1 receptor-binding domain – Influenza virus
2jrd, 1ibn, 1ibo – IvHA fusion domain – NMR
2dci, 2xoo, 2xop - IvHA fusion domain (mutant) – NMR
2ibx, 1ti8, 2wr0, 2wr1, 2wr2, 2wr3, 2wr4, 2wr5, 2wr7, 2wrb, 2wrc, 2wrd, 2wre, 2wrf, 2wrg, 2wrh – IvHA
1ha0 – IvHA precursor (mutant)
3m5g, 3al4, 3lzg, 3m6s, 3ku3, 3ku5, 3hto, 3eyj, 3bt6, 1ru7, 1ruy, 1ruz, 1rd8, 1mql, 1htm, 2hmg, 3hmg, 4hmg, 5hmg - IvHA1 + HA2
3htp, 3htq, 3htt, 2rft, 2rfu, 1rv0, 1rvt, 1rvx, 1rvz, 1mqm, 1mqn, 1hgd, 1hge, 1hgf, 1hgg, 1hgh, 1hgi, 1hgj, 3m5h, 3m5i, 3m5j - IvHA1 + HA2 + polysaccharide
3eyk, 3eym - IvHA1 + HA2 + membrane-fusion inhibitor
3ku6 - IvHA1 (mutant) + HA2
2viu - IvHA1 (mutant) + HA2 (mutant)
3r2x – IvHA1 receptor-binding domain + HA2 fusion subunit + designed HA-binding protein
3qqb, 3qqe, 3qqo - IvHA1 + HA2 ectodomain (mutant)
2fk0 - IvHA14 receptor-binding domain + HA2 membrane-fusion domain
3lzf, 3gbm, 3gbn, 3fku, 1ken, 1eo8, 1qfu, 2vir, 3sdy, 3sm5, 3ztj - IvHA1 + HA2 + antibody
2vis, 2vit - IvHA1 (mutant) + antibody
Viral hemagglutinin-neuraminidase
1usx – NvPIV head domain + thiosialoside – Newcastle diasease virus
1usr, 1e8t, 1e8u, 1e8v - NvPIV head domain
3t1e - NvPIV ectodomain
1v2i, 2v3b, 1v3b – hPIV – Human parainfluenza virus
2v3c, 2v3d, 1v3c, 1v3d, 1v3e – hPIV + glucoside
2v3e – hPIV + zanamavir
1z4v, 1z4w, 1z4z, 1z50 – SvPIV extracellular domain + DANA – Simian virus
1z4x, 1z4y - SvPIV extracellular domain + glucoside
2vsk, 2vsm – HN ephrin-binding domain + ephrin – Hendra virus
2vwd, 3d11 - NivHN receptor-binding domain – Nipah virus
3d12 - NivHN receptor-binding domain + ephrin
Measles hemagglutinin
2rkc – MvHA extracellular domain – Measles virus
2zb5, 2zb6 - MvHA head domain
3alw, 3alx, 3alz – MvHA/SLAM receptor
3inb – MvHA head domain + hMembrane cofactor protein
1rwr – HA filamentous – Bordetella pertussis
Dr hemagglutinin
2w5p – EcDrae adhesin subunit – Escherichia coli
1ut1 - EcDrae adhesin subunit (mutant)
2jkj, 2jkl, 2jkn, 1usq - EcDrae adhesin subunit + chloramphenicol derivative
Phytohemagglutinin
1g8w, 1fat – PHA-L – kidney bean
Various viral hemagglutinin
1kke – RvSigma1 – human orthoreovirus
3eoy – RvSigma1 + junctional adhesion molecule
2aen – VP8 – human rotavirus
3a5p – HA I – slime mold
Additional Resources
For additional information, see: Influenza
References
- ↑ Gottschalk A. Chemistry of virus receptors. The Viruses. 1959;3:51–61.
- ↑ Wiley DC, Skehel JJ. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu Rev Biochem. 1987;56:365-94. PMID:3304138 doi:http://dx.doi.org/10.1146/annurev.bi.56.070187.002053
- ↑ Liu J, Stevens DJ, Haire LF, Walker PA, Coombs PJ, Russell RJ, Gamblin SJ, Skehel JJ. Structures of receptor complexes formed by hemagglutinins from the Asian Influenza pandemic of 1957. Proc Natl Acad Sci U S A. 2009 Oct 6;106(40):17175-80. Epub 2009 Sep 28. PMID:19805083
- ↑ Cell Binding protein in Avian Influenza; Jack Cerchiara,'06 and Brendan Holsberry, 07;http://biology.kenyon.edu/BMB/Chime2/2005/Cerchiara-Holsberry/FRAMES/start.htm
- ↑ 5.0 5.1 Knossow M, Skehel JJ. Variation and infectivity neutralization in influenza. Immunology. 2006 Sep;119(1):1-7. PMID:16925526 doi:http://dx.doi.org/10.1111/j.1365-2567.2006.02421.x
- ↑ See Influenza in Wikipedia.
Proteopedia Page Contributors and Editors (what is this?)
Julien Madouasse, Michal Harel, David Canner, Mark Hoelzer, Ann Taylor, Alexander Berchansky, Joel L. Sussman