This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
1mae
From Proteopedia
(Difference between revisions)
(New page: 200px <!-- The line below this paragraph, containing "STRUCTURE_1mae", creates the "Structure Box" on the page. You may change the PDB parameter (which sets the PD...) |
m (Protected "1mae" [edit=sysop:move=sysop]) |
Revision as of 08:59, 2 June 2012
| |||||||||
| 1mae, resolution 2.80Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Ligands: | |||||||||
| Non-Standard Residues: | |||||||||
| Activity: | Amine dehydrogenase, with EC number 1.4.99.3 | ||||||||
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, PDBsum, RCSB | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
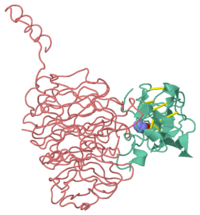
The Active Site Structure of Methylamine Dehydrogenase: Hydrazines Identify C6 as the Reactive Site of the Tryptophan Derived Quinone Cofactor
Template:ABSTRACT PUBMED 1390754
About this Structure
1mae is a 2 chain structure of Methylamine dehydrogenase with sequence from Paracoccus versutus. Full crystallographic information is available from OCA.
See Also
Reference
- Huizinga EG, van Zanten BA, Duine JA, Jongejan JA, Huitema F, Wilson KS, Hol WG. Active site structure of methylamine dehydrogenase: hydrazines identify C6 as the reactive site of the tryptophan-derived quinone cofactor. Biochemistry. 1992 Oct 13;31(40):9789-95. PMID:1390754