This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
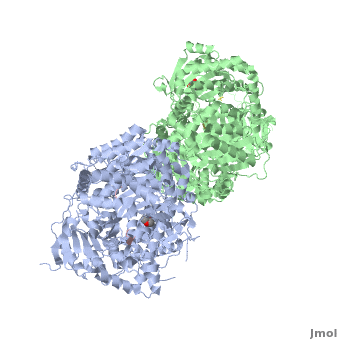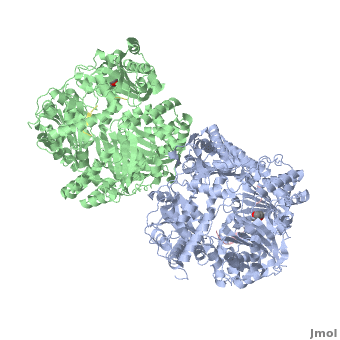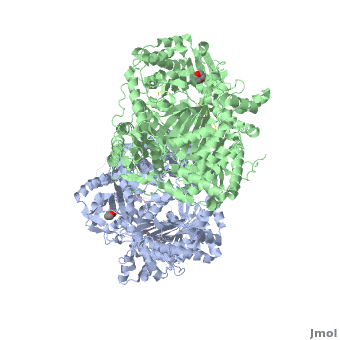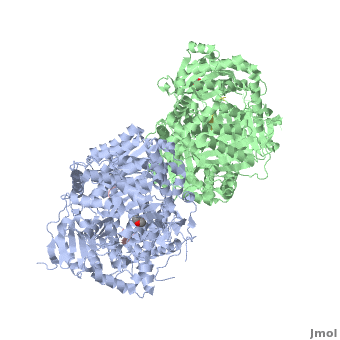
Insulin-Degrading Enzyme
From Proteopedia
(Difference between revisions)
| Line 62: | Line 62: | ||
==3D structures of insulin-degrading enzyme== | ==3D structures of insulin-degrading enzyme== | ||
| - | ''Updated | + | ''Updated June 2012'' |
| - | + | [[2jg4]] - hIDE (mutant) – human<br /> | |
| - | [[3n56]], [[3n57]] – hIDE | + | [[3qz2]] - hIDE<br /> |
| - | [[ | + | [[3n56]], [[3n57]] – hIDE (mutant) + B-type natriuretic peptide <br /> |
| - | [[3h44]] - hIDE | + | [[2yb3]] - hIDE (mutant) + inhibitor<br /> |
| - | [[3hgz]], [[2g48]] - hIDE | + | [[3ofi]] - hIDE + ubiquitin<br /> |
| - | [[3e4z]] - hIDE | + | [[3h44]] - hIDE (mutant) + C-C motif chemokine 3<br /> |
| - | [[3e50]] - hIDE | + | [[3hgz]], [[2g48]] - hIDE (mutant) + amylin<br /> |
| + | [[3e4z]] - hIDE (mutant) + insulin-like growth factor II<br /> | ||
| + | [[3e50]] - hIDE (mutant) + Transforming growth factor α<br /> | ||
[[3e4a]] – hIDE (mutant) + hydroxamate peptide II1<br /> | [[3e4a]] – hIDE (mutant) + hydroxamate peptide II1<br /> | ||
| - | [[2wby]], [[2wc0]] - hIDE | + | [[2wby]], [[2wc0]] - hIDE (mutant) + insulin<br /> |
| - | [[3cww]] - hIDE | + | [[3cww]] - hIDE (mutant) + Bradykinin N terminal peptide<br /> |
| - | [[2jbu]] - hIDE | + | [[2jbu]] - hIDE (mutant) + co-purified peptide<br /> |
| - | [[2g47]], [[2wk3]] - hIDE | + | [[2g47]], [[2wk3]] - hIDE (mutant) + amyloid β A4<br /> |
| - | [[2g49]] - hIDE | + | [[2g49]] - hIDE (mutant) + glucagons<br /> |
| - | [[2g54]] - hIDE | + | [[2g54]] - hIDE (mutant) + insulin β chain + Zn<br /> |
| - | [[2g56]] - hIDE | + | [[2g56]] - hIDE + insulin β chain residues 25-54<br /> |
| - | [[3p7l]] - rIDE | + | [[3p7l]] - rIDE – rat<br /> |
| - | [[3p7o]] - rIDE | + | [[3p7o]] - rIDE (mutant) + peptide<br /> |
| - | [[3tuv]] - rIDE | + | [[3tuv]] - rIDE + peptide + ATP |
</StructureSection> | </StructureSection> | ||
== References == | == References == | ||
Revision as of 09:52, 14 June 2012
| |||||||||||
References
- Im H, Manolopoulou M, Malito E, Shen Y, Zhao J, Neant-Fery M, Sun CY, Meredith SC, Sisodia SS, Leissring MA, Tang WJ. Structure of substrate-free human insulin-degrading enzyme (IDE) and biophysical analysis of ATP-induced conformational switch of IDE. J Biol Chem. 2007 Aug 31;282(35):25453-63. Epub 2007 Jul 5. PMID:17613531 doi:10.1074/jbc.M701590200
- Shen Y, Joachimiak A, Rosner MR, Tang WJ. Structures of human insulin-degrading enzyme reveal a new substrate recognition mechanism. Nature. 2006 Oct 19;443(7113):870-4. Epub 2006 Oct 11. PMID:17051221 doi:10.1038/nature05143
- Li P, Kuo WL, Yousef M, Rosner MR, Tang WJ. The C-terminal domain of human insulin degrading enzyme is required for dimerization and substrate recognition. Biochem Biophys Res Commun. 2006 May 19;343(4):1032-7. Epub 2006 Mar 22. PMID:16574064 doi:10.1016/j.bbrc.2006.03.083
Proteopedia Page Contributors and Editors (what is this?)
Michal Harel, Alexander Berchansky, Élodie Weider, Karsten Theis, Joel L. Sussman, Jaime Prilusky