This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Clegg TBP sandbox
From Proteopedia
| Line 7: | Line 7: | ||
The TATA box-binding protein (TBP) is required by all three eukaryotic RNA polymerases for correct initiation of transcription of ribosomal, messenger, small nuclear, and transfer RNAs. The cocrystal structure of the C-terminal/core region of human TBP complexed with the TATA element of the adenovirus major late promoter has been determined at 1.9 angstroms resolution. Structural and functional analyses of the protein-DNA complex are presented, with a detailed comparison to our 1.9-angstroms resolution structure of Arabidopsis thaliana TBP2 bound to the same TATA box. | The TATA box-binding protein (TBP) is required by all three eukaryotic RNA polymerases for correct initiation of transcription of ribosomal, messenger, small nuclear, and transfer RNAs. The cocrystal structure of the C-terminal/core region of human TBP complexed with the TATA element of the adenovirus major late promoter has been determined at 1.9 angstroms resolution. Structural and functional analyses of the protein-DNA complex are presented, with a detailed comparison to our 1.9-angstroms resolution structure of Arabidopsis thaliana TBP2 bound to the same TATA box. | ||
| - | The TBP has two imperfect direct repeats that form a <scene name='Clegg_TBP_sandbox/Saddle/1'>saddle</scene> shaped structure forming a concave like DNA binding surface allowing for an induced-fit mechanism. The imperfect direct repeats differ by two residues. The first is a deletion of a resiude at the end of helix H1. The second is a proline in helix H2' that enables a <scene name='Clegg_TBP_sandbox/Proline/1'>bend</scene> that does not occur on H2. When the TBP binds to the TATA box a conformation change takes place and the DNA strand is <scene name='Clegg_TBP_sandbox/Bend/1'>bent</scene> 80 degrees. This conformational change promotes the recruitment of other general transcription factors. | + | The TBP has two imperfect direct repeats that form a <scene name='Clegg_TBP_sandbox/Saddle/1'>saddle</scene> shaped structure forming a concave like DNA binding surface allowing for an induced-fit mechanism. The imperfect direct repeats differ by two residues. The first is a deletion of a resiude at the end of helix H1. The second is a proline in helix H2' that enables a <scene name='Clegg_TBP_sandbox/Proline/1'>bend</scene> that does not occur on H2. When the TBP binds to the TATA box a conformation change takes place and the DNA strand is <scene name='Clegg_TBP_sandbox/Bend/1'>bent</scene> 80 degrees. This conformational change promotes the recruitment of other general transcription factors. A total of seven <scene name='Clegg_TBP_sandbox/Lysines/1'>lysine</scene> residues interact with DNA backbone. Four of these residues Lys-204, Lys-214, Lys-295, and Lys 305 allow for the patial charge neutralization within the TATA box. |
==About this Structure== | ==About this Structure== | ||
Revision as of 05:52, 7 November 2012
| |||||||||
| 1cdw, resolution 1.90Å () | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| |||||||||
| |||||||||
| Resources: | FirstGlance, OCA, RCSB, PDBsum | ||||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||||
Contents |
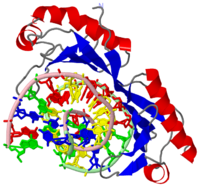
HUMAN TBP CORE DOMAIN COMPLEXED WITH DNA
The TATA box-binding protein (TBP) is required by all three eukaryotic RNA polymerases for correct initiation of transcription of ribosomal, messenger, small nuclear, and transfer RNAs. The cocrystal structure of the C-terminal/core region of human TBP complexed with the TATA element of the adenovirus major late promoter has been determined at 1.9 angstroms resolution. Structural and functional analyses of the protein-DNA complex are presented, with a detailed comparison to our 1.9-angstroms resolution structure of Arabidopsis thaliana TBP2 bound to the same TATA box.
The TBP has two imperfect direct repeats that form a shaped structure forming a concave like DNA binding surface allowing for an induced-fit mechanism. The imperfect direct repeats differ by two residues. The first is a deletion of a resiude at the end of helix H1. The second is a proline in helix H2' that enables a that does not occur on H2. When the TBP binds to the TATA box a conformation change takes place and the DNA strand is 80 degrees. This conformational change promotes the recruitment of other general transcription factors. A total of seven residues interact with DNA backbone. Four of these residues Lys-204, Lys-214, Lys-295, and Lys 305 allow for the patial charge neutralization within the TATA box.
About this Structure
1cdw is a 3 chain structure with sequence from Homo sapiens. The July 2005 RCSB PDB Molecule of the Month feature on TATA-Binding Protein by David S. Goodsell is 10.2210/rcsb_pdb/mom_2005_7. Full crystallographic information is available from OCA.
See Also
Reference
- Nikolov DB, Chen H, Halay ED, Hoffman A, Roeder RG, Burley SK. Crystal structure of a human TATA box-binding protein/TATA element complex. Proc Natl Acad Sci U S A. 1996 May 14;93(10):4862-7. PMID:8643494
- Cokol M, Nair R, Rost B. Finding nuclear localization signals. EMBO Rep. 2000 Nov;1(5):411-5. PMID:11258480 doi:10.1093/embo-reports/kvd092
- Hicks JM, Hsu VL. The extended left-handed helix: a simple nucleic acid-binding motif. Proteins. 2004 May 1;55(2):330-8. PMID:15048824 doi:10.1002/prot.10630