This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Grb10 SH2 Domain
From Proteopedia
(Difference between revisions)
| Line 11: | Line 11: | ||
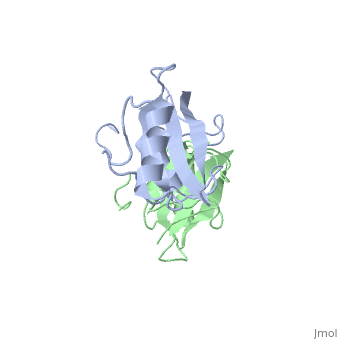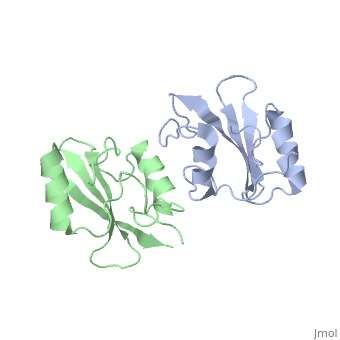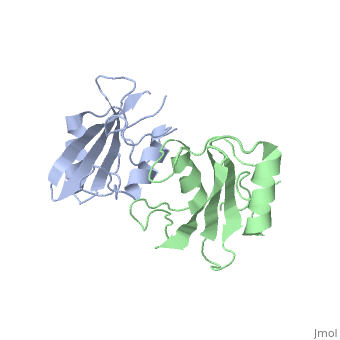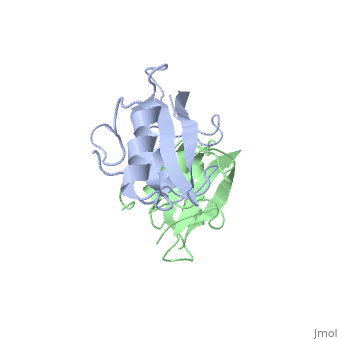
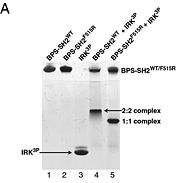
To ensure the crystallographic structure of Grb10 SH2 is indeed a dimer in solution, Evan Stein and colleagues substituted Phe515 at the dimer interface with arginine (electrically charged side chain) and found, using gel filtration chromatography, that the Grb10 SH2 dimer had indeed become independent monomers. | To ensure the crystallographic structure of Grb10 SH2 is indeed a dimer in solution, Evan Stein and colleagues substituted Phe515 at the dimer interface with arginine (electrically charged side chain) and found, using gel filtration chromatography, that the Grb10 SH2 dimer had indeed become independent monomers. | ||
| - | + | [[Image:Figure_6.jpeg | thumb | alt=text | asdflj]] | |
| + | |||
</StructureSection> | </StructureSection> | ||
Revision as of 05:52, 8 November 2012
| |||||||||||
Interaction Between Grb10 and E3 Ubiquitin Ligase NEDD4
WOOOOOOOO
|
Grb10 Gene Inhibition Affects Body Composition, and Insulin Signaling
References
- ↑ 1.0 1.1 1.2 Stein EG, Ghirlando R, Hubbard SR. Structural basis for dimerization of the Grb10 Src homology 2 domain. Implications for ligand specificity. J Biol Chem. 2003 Apr 11;278(15):13257-64. Epub 2003 Jan 27. PMID:12551896 doi:http://dx.doi.org/10.1074/jbc.M212026200