This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
RA Mediated T-reg Differentiation
From Proteopedia
| Line 3: | Line 3: | ||
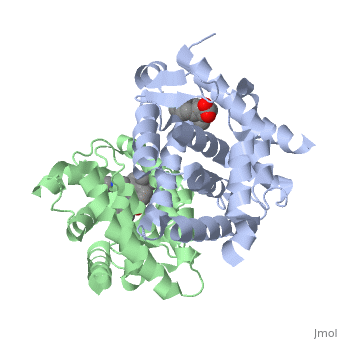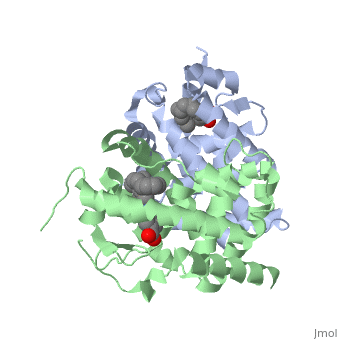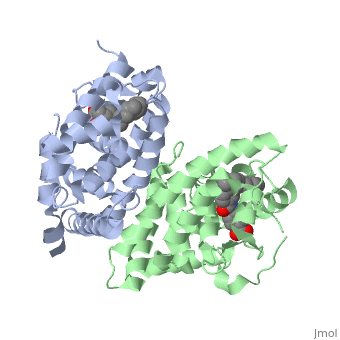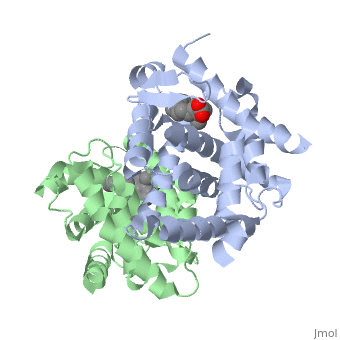
<StructureSection load='1dkf' size='350' side='left' caption='RARα-RXRα interaction (PDB entry [[1dkf]])' scene=''> | <StructureSection load='1dkf' size='350' side='left' caption='RARα-RXRα interaction (PDB entry [[1dkf]])' scene=''> | ||
==Ligand Binding Domain== | ==Ligand Binding Domain== | ||
| - | The Ligand binding domain for each piece of the dimer has a | + | The Ligand binding domain for each piece of the dimer has a nearly identical structure of an <scene name='RA_Mediated_T-reg_Differentiaition/Alpha-helical_domains/2'>Tα-helical sandwich</scene>. These alpha helices form a total of 12 domains per protein (referred to as H1-12), with an additional 2 beta sheets as well. Both monomers contain two regions of activity, the <scene name='RA_Mediated_T-reg_Differentiaition/Dimerization_interface/2'>dimerization interface</scene> and the ligand binding domain.<ref> PMID: 10882070 </ref> |
| + | |||
=== RARα-RXRα Heterodimer === | === RARα-RXRα Heterodimer === | ||
| Line 9: | Line 10: | ||
| - | The residues of | + | The residues of <scene name='RA_Mediated_T-reg_Differentiaition/Rar_dimer_interface/1'> RAR-</scene>α that are interacting in the heterodimer are as follows: |
Hydrophobic residues: L356, F374, P375, L378, M379, I381 and A389 (yellow); | Hydrophobic residues: L356, F374, P375, L378, M379, I381 and A389 (yellow); | ||
Negatively charged residues: D338, D349, E353, E357, D383, and E393 (red); | Negatively charged residues: D338, D349, E353, E357, D383, and E393 (red); | ||
Revision as of 06:23, 14 November 2012
Introduction
T-regulatory cells (T-regs) are a small subset of CD4+ T-cells that exhibit strong down regulation of immune system activity in their local environment. They are distinguished from other CD4+ T-cells by the expression of FOXP3, a gene regulator. [1] These cells have been shown to differentiate from CD4+ T-helper cells upon activation and exposure to the following cytokines: tumor growth factor β (TGF-β), Interleukin-2 (IL-2) and retinoic acid (RA). [2] Both TGF-β and IL-2 are used in other immune system differentiation, however, RA has been shown to bias T-cells to the T-reg phenotype. [3] When acting upon T-reg cells, RA acts as the ligand for the Retinoic Acid Receptor-α (RARα) / Retinoid X Receptor-α (RXRα) heterodimer. This heterodimer is of the nuclear receptor family, and each chain consists of the same three part structure: a Ligand binding domain (LBD), a DNA binding domain (DBD), and a hinge region connecting the two binding domains. [4]
| |||||||||||
| |||||||||||
Biological Significance
References
- ↑ Ochs HD, Oukka M, Torgerson TR. TH17 cells and regulatory T cells in primary immunodeficiency diseases. J Allergy Clin Immunol. 2009 May;123(5):977-83; quiz 984-5. PMID:19410687 doi:10.1016/j.jaci.2009.03.030
- ↑ Moore C, Fuentes C, Sauma D, Morales J, Bono MR, Rosemblatt M, Fierro JA. Retinoic acid generates regulatory T cells in experimental transplantation. Transplant Proc. 2011 Jul-Aug;43(6):2334-7. PMID:21839265 doi:10.1016/j.transproceed.2011.06.057
- ↑ Moore C, Fuentes C, Sauma D, Morales J, Bono MR, Rosemblatt M, Fierro JA. Retinoic acid generates regulatory T cells in experimental transplantation. Transplant Proc. 2011 Jul-Aug;43(6):2334-7. PMID:21839265 doi:10.1016/j.transproceed.2011.06.057
- ↑ Kumar R, Thompson EB. The structure of the nuclear hormone receptors. Steroids. 1999 May;64(5):310-9. PMID:10406480
- ↑ Bourguet W, Vivat V, Wurtz JM, Chambon P, Gronemeyer H, Moras D. Crystal structure of a heterodimeric complex of RAR and RXR ligand-binding domains. Mol Cell. 2000 Feb;5(2):289-98. PMID:10882070
- ↑ Bourguet W, Vivat V, Wurtz JM, Chambon P, Gronemeyer H, Moras D. Crystal structure of a heterodimeric complex of RAR and RXR ligand-binding domains. Mol Cell. 2000 Feb;5(2):289-98. PMID:10882070
Proteopedia Page Contributors and Editors (what is this?)
William Bailey, Alexander Berchansky, Michal Harel, Jaime Prilusky