This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
Halophilic malate dehydrogenase
From Proteopedia
| Line 1: | Line 1: | ||
== ''Structural features that stabilize halophilic malate dehydrogenase from an archaebacterium'' == | == ''Structural features that stabilize halophilic malate dehydrogenase from an archaebacterium'' == | ||
| - | <StructureSection load='1HLP' size='500' side='center' caption='3D Structure of halophilic malate dehydrogenase ' | + | <StructureSection load='1HLP' size='500' side='center' caption='3D Structure of halophilic malate dehydrogenase ' scene='Insert optional scene name here' /> |
== '''Scheme''' == | == '''Scheme''' == | ||
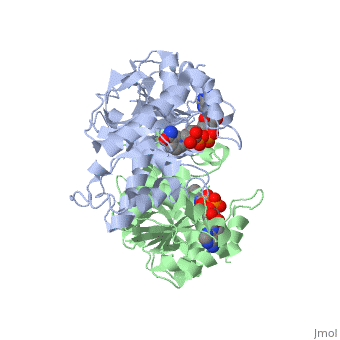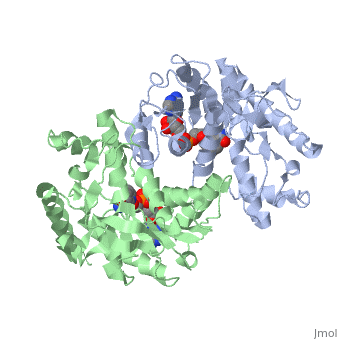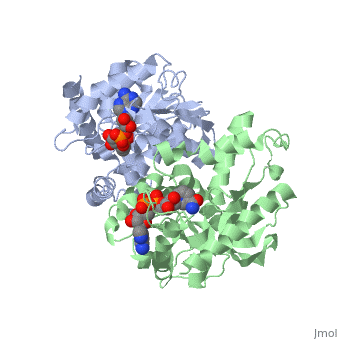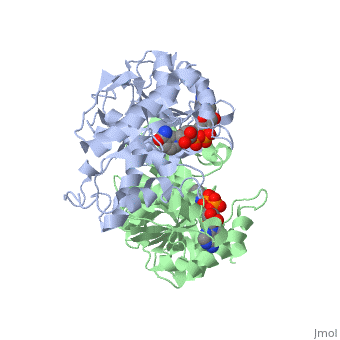
| - | The high-resolution structure of halophilic malate dehydrogenase (hMDH) from the archaebacterium Haloarcula marismortui [[http://www.highbeam.com/doc/1G1-16845034.html]] was determined by x-ray crystallography. Comparison of the three-dimensional structures of hMDH and its nonhalophilic congeners reveals structural features that may promote the stability of hMDH at high salt concentrations. These features include an excess of acidic over basic residues distributed on the enzyme surface and more salt bridges present in hMDH compared with its nonhalophilic counterparts. Other features that contribute to the stabilization of thermophilic lactate dehydrogenase and thermophilic MDH-the incorporation of alanine into alpha helices and the introduction of negatively charged amino acids near their amino termini, both of which stabilize the alpha helix as a result of interaction with the positive part of the alpha-helix dipole-also were observed in hMDH | + | The <scene name='WISPSF201208/Cationic_and_anionic_residues/1'>TextToBeDisplayed</scene>high-resolution structure of halophilic malate dehydrogenase (hMDH) from the archaebacterium Haloarcula marismortui [[http://www.highbeam.com/doc/1G1-16845034.html]] was determined by x-ray crystallography. Comparison of the three-dimensional structures of hMDH and its nonhalophilic congeners reveals structural features that may promote the stability of hMDH at high salt concentrations. These features include an excess of acidic over basic residues distributed on the enzyme surface and more salt bridges present in hMDH compared with its nonhalophilic counterparts. Other features that contribute to the stabilization of thermophilic lactate dehydrogenase and thermophilic MDH-the incorporation of alanine into alpha helices and the introduction of negatively charged amino acids near their amino termini, both of which stabilize the alpha helix as a result of interaction with the positive part of the alpha-helix dipole-also were observed in hMDH |
Revision as of 19:34, 24 December 2012
Structural features that stabilize halophilic malate dehydrogenase from an archaebacterium
|
Scheme
The high-resolution structure of halophilic malate dehydrogenase (hMDH) from the archaebacterium Haloarcula marismortui [[1]] was determined by x-ray crystallography. Comparison of the three-dimensional structures of hMDH and its nonhalophilic congeners reveals structural features that may promote the stability of hMDH at high salt concentrations. These features include an excess of acidic over basic residues distributed on the enzyme surface and more salt bridges present in hMDH compared with its nonhalophilic counterparts. Other features that contribute to the stabilization of thermophilic lactate dehydrogenase and thermophilic MDH-the incorporation of alanine into alpha helices and the introduction of negatively charged amino acids near their amino termini, both of which stabilize the alpha helix as a result of interaction with the positive part of the alpha-helix dipole-also were observed in hMDH