This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2bo5
From Proteopedia
m (Protected "2bo5" [edit=sysop:move=sysop]) |
|||
| Line 1: | Line 1: | ||
| - | [[ | + | ==BOVINE OLIGOMYCIN SENSITIVITY CONFERRAL PROTEIN N-TERMINAL DOMAIN== |
| + | <StructureSection load='2bo5' size='340' side='right' caption='[[2bo5]], [[NMR_Ensembles_of_Models | 44 NMR models]]' scene=''> | ||
| + | == Structural highlights == | ||
| + | <table><tr><td colspan='2'>[[2bo5]] is a 1 chain structure with sequence from [http://en.wikipedia.org/wiki/Bos_taurus Bos taurus]. Full experimental information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2BO5 OCA]. For a <b>guided tour on the structure components</b> use [http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2BO5 FirstGlance]. <br> | ||
| + | </td></tr><tr><td class="sblockLbl"><b>Activity:</b></td><td class="sblockDat"><span class='plainlinks'>[http://en.wikipedia.org/wiki/H(+)-transporting_two-sector_ATPase H(+)-transporting two-sector ATPase], with EC number [http://www.brenda-enzymes.info/php/result_flat.php4?ecno=3.6.3.14 3.6.3.14] </span></td></tr> | ||
| + | <tr><td class="sblockLbl"><b>Resources:</b></td><td class="sblockDat"><span class='plainlinks'>[http://oca.weizmann.ac.il/oca-docs/fgij/fg.htm?mol=2bo5 FirstGlance], [http://oca.weizmann.ac.il/oca-bin/ocaids?id=2bo5 OCA], [http://www.rcsb.org/pdb/explore.do?structureId=2bo5 RCSB], [http://www.ebi.ac.uk/pdbsum/2bo5 PDBsum]</span></td></tr> | ||
| + | <table> | ||
| + | == Evolutionary Conservation == | ||
| + | [[Image:Consurf_key_small.gif|200px|right]] | ||
| + | Check<jmol> | ||
| + | <jmolCheckbox> | ||
| + | <scriptWhenChecked>select protein; define ~consurf_to_do selected; consurf_initial_scene = true; script "/wiki/ConSurf/bo/2bo5_consurf.spt"</scriptWhenChecked> | ||
| + | <scriptWhenUnchecked>script /wiki/extensions/Proteopedia/spt/initialview01.spt</scriptWhenUnchecked> | ||
| + | <text>to colour the structure by Evolutionary Conservation</text> | ||
| + | </jmolCheckbox> | ||
| + | </jmol>, as determined by [http://consurfdb.tau.ac.il/ ConSurfDB]. You may read the [[Conservation%2C_Evolutionary|explanation]] of the method and the full data available from [http://bental.tau.ac.il/new_ConSurfDB/chain_selection.php?pdb_ID=2ata ConSurf]. | ||
| + | <div style="clear:both"></div> | ||
| + | <div style="background-color:#fffaf0;"> | ||
| + | == Publication Abstract from PubMed == | ||
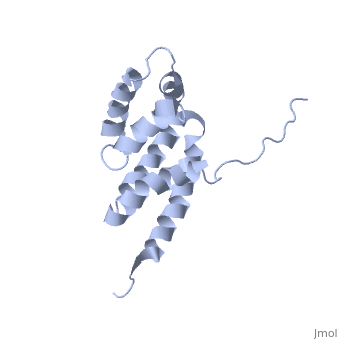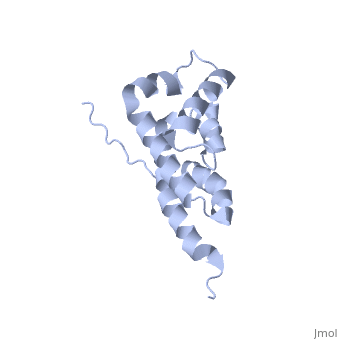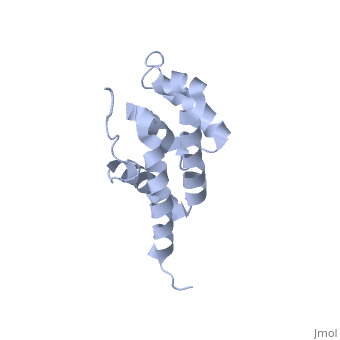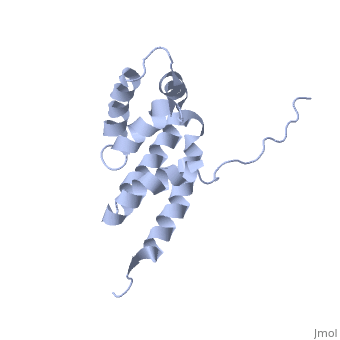
| + | The peripheral stalk of ATP synthase holds the alpha3beta3 catalytic subcomplex stationary against the torque of the rotating central stalk. In bovine mitochondria, the N-terminal domain of the oligomycin sensitivity conferral protein (OSCP-NT; residues 1-120) anchors one end of the peripheral stalk to the N-terminal tails of one or more alpha-subunits of the F1 subcomplex. Here we present the solution structure of OSCP-NT and an NMR titration study of its interaction with peptides representing N-terminal tails of F1 alpha-subunits. The structure comprises a bundle of six alpha-helices, and its interaction site contains adjoining hydrophobic surfaces of helices 1 and 5; residues in the region 1-8 of the alpha-subunit are essential for the interaction. The OSCP-NT is similar to the N-terminal domain of the delta-subunit from Escherichia coli ATP synthase (delta-NT), except that their surface charges differ (basic and acidic, respectively). As the charges of the adjacent crown regions in their alpha3beta3 complexes are similar, the OSCP-NT and delta-NT probably do not contact the crowns extensively. The N-terminal tails of alpha-subunit tails are probably alpha-helical, and so this interface, which is essential for the rotary mechanism of the enzyme, appears to consist of helix-helix interactions. | ||
| - | + | Structure of the F1-binding domain of the stator of bovine F1Fo-ATPase and how it binds an alpha-subunit.,Carbajo RJ, Kellas FA, Runswick MJ, Montgomery MG, Walker JE, Neuhaus D J Mol Biol. 2005 Aug 26;351(4):824-38. PMID:16045926<ref>PMID:16045926</ref> | |
| - | + | From MEDLINE®/PubMed®, a database of the U.S. National Library of Medicine.<br> | |
| - | + | </div> | |
| - | + | == References == | |
| - | + | <references/> | |
| - | + | __TOC__ | |
| - | + | </StructureSection> | |
| - | + | ||
| - | == | + | |
| - | < | + | |
[[Category: Bos taurus]] | [[Category: Bos taurus]] | ||
[[Category: Carbajo, R J.]] | [[Category: Carbajo, R J.]] | ||
Revision as of 01:22, 30 September 2014
BOVINE OLIGOMYCIN SENSITIVITY CONFERRAL PROTEIN N-TERMINAL DOMAIN
| |||||||||||
Categories: Bos taurus | Carbajo, R J. | Kellas, F A. | Montgomery, M G. | Neuhaus, D. | Runswick, M J. | Walker, J E. | Alpha-subunit | Atp synthase | Beta-subunit | Binding interface | Chemical shift mapping | Chemical shift perturbation | Hydrogen ion transport | Hydrolase | Ion transport | Mitochondrion | Oscp | Peripheral stalk | Protein-protein interaction | Titration | Transit peptide | Transport