From Proteopedia
(Difference between revisions)
proteopedia linkproteopedia link
|
|
| Line 67: |
Line 67: |
| | ==3D structures of hemagglutinin== | | ==3D structures of hemagglutinin== |
| | | | |
| - | ''Updated February 2013''
| + | Updated on {{REVISIONDAY2}}-{{MONTHNAME|{{REVISIONMONTH}}}}-{{REVISIONYEAR}} |
| | | | |
| | [[3aj5]], [[3ah1]], [[3ah2]], [[3ah4]] – CbHA1 + glucoside - ''Clostridium botulinum''<br /> | | [[3aj5]], [[3ah1]], [[3ah2]], [[3ah4]] – CbHA1 + glucoside - ''Clostridium botulinum''<br /> |
Revision as of 08:59, 10 March 2013
| Hemagglutinins (HA) are one of the two antigenic Glycoproteins inserted into the influenza virus' membrane. There are at least 16 different forms of HA Antigens classified from H1 to H16. H1, H2, H3 are specific to human influenza. HAs have two main functions crucial for the viral infection cycle:
1. On the target cell, HA bind the receptor which is composed by Sialic acid [1] allowing the virus/cell interaction.
2. HA induce the fusion between the host cell and the virus which can entry into the cytoplasm.
Because the HA are the major Antigens of the Virus, Antibodies recognized them and for this reason HA often change.
For discussion of influenza hemagglutinin see Influenza hemagglutinin, User:Michael Strong/H1N1/HA and User:Michael Strong/H1N1/HA/MSA for multiple sequence alignment.
Hemagglutinin Structure
HA is an integral membrane glycoprotein. Each monomer is synthesized like a single polypeptide chain almost 550 amino acids. This precursor is then glycosilated and cleaved into two smaller polypeptides by removal of Arginin 329; at the same time, a conformational change occurs in the monomer.
The HA1 and HA2 subunits are covalently attached by a from HA1 position 14 to HA2 position 467(*). These two chains form one monomer, and the noncovalentely association of three monomers forms one hemagglutinin molecule: (HA1+HA2)3. It is principally stabilized by packing of the alpha-helixes. All molecules are 135 Angström long.
The (328 amino acids) has a globular form outside the virus’ membrane: it is composed by an eight-stranded beta-sheet, associated with a little alpha-helix, separating the strands 3 and 4.
The amino acids of this alpha-helix, and some others around it, included in the beta-sheet, compose the binding site for the receptor’s sialic acid. Thus, for one molecule of hemagglutinin, there are three binding sites to the host cell’s receptor.
The (221 amino acids) has a hair spin structure composed by two antiparallel alpha-helixes. One of these belongs to the longest alpha-helixes in globular protein: it is 50 angstrom long. The hydrophobic N-terminus part of HA2, called fusion peptide, is close to a protease’s cleavage site: this protease is implied in the virus’ entry in the host cell.
HA1 and HA2 are bound each other by a disulfide bind. The three long alpha-helixes (of the three HA2) are coiled-coil to form a central region of 40 Angström, and thank to the hydrophobic amino acids and those which form salt bridges bound, we obtain a HA stabilized.[2]
2WRE
To begin, this code corresponds to the structure of influenza H2 Avian Hemagglutinin from the “Asian Influenza” of 1957.[3] Human receptor.
- At the position 226, Avian Hemagglutinin has a Glutamine residue whereas the Human one has Leucine
- At the position 228, there is a Glycin for Avian HA whereas a Serine for Human HA.
These two mutations prevent the human H2 binding on the avian receptor whereas the human receptor can be bound by avian hemagglutinin that lacks the human specific mutation of H2 pandemic viruses.
There are three sites for sialic acid binding, which are at the membrane-distal tips of the identical monomers that form the HA trimer:
- The first one is the composed by the residues from 225 to 228
- The second is called and contains the residues 131 to 137
- the last is named 190-helix
The residues of these loops 130 and 220 have carbonyl oxygens and amide nitrogens of peptide bonds exposed with potential to interact with the receptor, which is composed by Gal-1 and Sia-2.
The avian H2 can bind the human receptor because of intramolecular Hydrogen bond network created by Gln 226 and by Asn 186.
Four amino acids compose : Tyr 98, Ser 136, Trp 153(**), His 183, (which are identical in the all different HA)(*). This site forms a pocket on the distal end of the molecule. The binding structure is stabilized by other conserved which cannot interact with the receptor
(*)
The Sia-1-Gal-2 glycosidic bond adopts a cis conformation. Extensive hydrogen bond direction are settled between avian and Gal-2 of the receptor: two water molecules (Wat-1 and Wat-2) have a significant role in mediating these interactions(see fig.1).
The site chain of Lys-222 and the main chain carbonyl at 225 are linked by Wat-1 to the 3'OH of Gal-2. Gln-226 and Asn 186 form hydrogen bonds with the hydroxyl groups of 4'C of Gal-2 and 9'C of Sia-1.[4]
Oligosaccharides
Some of HA1 (8, 22, 38, 81, 165, 285)(*) have oligosaccharides chains attached to them. The seventh oligasaccharide is linked to an Aspargine of HA2 (484). All of them are complex oligosaccharides and are on the lateral surfaces of the molecule, except for site and 165. No precise functions have been assigned to them, even if the oligosaccharide at 165 seems to allow the stabilisation of oligomeric contact between globular units at the top of the protein's structure.
Antigenic variability
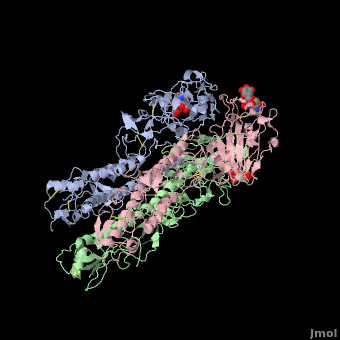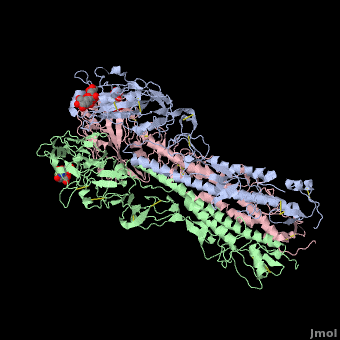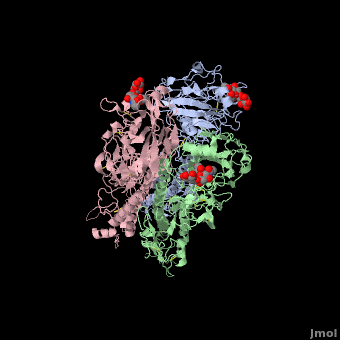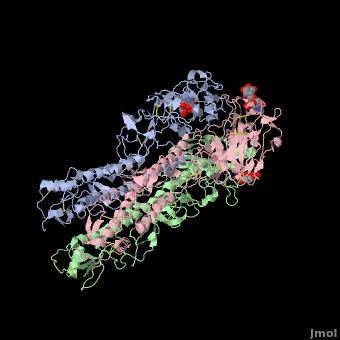
Each Hemagglutinin monomer contains four antigenic sites called . "A"(in red) site is a loop that protrudes 8 Angström distally from the surface of the target membrane. "B"(in green) site is composed by the external amino acids of one alpha helix and several amino acids of the receptor binding site which contains the sialic acid. "C"(in blue) site is a 60 Angström bulge from the distal extremity of HA. To finish, "D"(in mangenta) site is located in two beta-sheet of the jelly roll of HA1 globular extremity.
The substitutions of amino acids in these sites allow the influenza virus to escape from the immune system and to spread. Major changes in these antigenic regions can produce lethal influenza pandemics, like in 1957.
How Influenza Escapes Vaccines
Influenza hemagglutinin (e.g. 1hgf) is a glycoprotein on the surface of influenza virus particles that enables them to attach to and infect host cells. Antibodies that bind to hemagglutinin are a major defense mechanism that prevent infection. The RNA genome of influenza is characterized by a high mutation rate. Mutations on the surface of hemagglutinin tend to be protective for the virus. They tend to be retained because they tend to reduce the binding strength, and hence the host defensive capability, of antibodies that recognize the un-mutated hemagglutinin[5]. Influenza vaccines include hemagglutinins and they induce anti-hemagglutinin antibodies in vaccinated individuals. Often, however, by the time the vaccines can be designed, produced, and disseminated, mutant influenza viruses have arisen that can cause disease in vaccinated individuals[6].
If you show the Evolutionary Conservation of the structure on this page (using the blue bars below the molecule), you will see highly variable surface amino acids. These represent sites of mutations that have been retained in wild strains of influenza because they improve virus survival[5]. (Added by Eric Martz).
Notes
(*)These four amino-acids don't have the same numbering in the text and on the picture because there is a frame shift between the PDB and the Swissprot sequence numbering.
(**) The amino acid 153 lacks for the A monomer chain.
|
3D structures of hemagglutinin
Updated on 10-March-2013
3aj5, 3ah1, 3ah2, 3ah4 – CbHA1 + glucoside - Clostridium botulinum
1qxm - CbHA1 (mutant)
3aj6 - CbHA1 (mutant) + glucoside
2zoe, 2zs6, 4en6, 4en7, 4en8, 4en9 – CbHA3 + glucoside
2e4m – CbHA1 + HA17
3a5p – HA I – slime mold
4fbr – MxHA – Myxococcus xanthus
4fbv - MxHA + glucoside
Influenza hemagglutinin
3qqi, 3mlh - IvHA1 receptor-binding domain – Influenza virus
2jrd, 1ibn, 1ibo – IvHA fusion domain – NMR
2dci, 2xoo, 2xop - IvHA fusion domain (mutant) – NMR
2ibx, 1ti8, 2wr0, 2wr1, 2wr2, 2wr3, 2wr4, 2wr5, 2wr7, 2wrb, 2wrc, 2wrd, 2wre, 2wrf, 2wrg, 2wrh, 3uyw – IvHA
2yp2, 2yp3, 2yp4, 2yp5, 2yp7, 2yp8, 2yp9, 3uyx, 4fiu - IvHA1 (mutant)
4f23 - IvHA precursor
1ha0 – IvHA precursor (mutant)
1qu1 – IvHA2
3m5g, 3al4, 3lzg, 3m6s, 3ku3, 3ku5, 3hto, 3eyj, 3bt6, 1ru7, 1ruy, 1ruz, 1rd8, 1mql, 1htm, 2hmg, 3hmg, 4hmg, 5hmg, 1jsd, 1jsm, 3s11, 3s12, 3s13, 2ypg, 4dj6, 4dj7, 4dj8- IvHA1 + HA2
1jsh, 1jsi, 1jsn, 1jso - IvHA1 + HA2 + sialic acid
3ubj, 3ube, 3ubn, 3ubq - IvHA1 (mutant) + HA2 + sialic acid
3htp, 3htq, 3htt, 2rft, 2rfu, 1rv0, 1rvt, 1rvx, 1rvz, 1mqm, 1mqn, 1hgd, 1hge, 1hgf, 1hgg, 1hgh, 1hgi, 1hgj, 3m5h, 3m5i, 3m5j - IvHA1 + HA2 + polysaccharide
3eyk, 3eym - IvHA1 + HA2 + membrane-fusion inhibitor
3ku6, 4f3z - IvHA1 (mutant) + HA2
2viu, 3vun - IvHA1 (mutant) + HA2 (mutant)
3r2x – IvHA1 receptor-binding domain + HA2 fusion subunit + designed HA-binding protein
3qqb, 3qqe, 3qqo - IvHA1 + HA2 ectodomain (mutant)
2fk0 - IvHA14 receptor-binding domain + HA2 membrane-fusion domain
3lzf, 3gbm, 3gbn, 3fku, 1ken, 1eo8, 1qfu, 2vir, 3sdy, 3sm5, 3ztj, 3ztn, 4gms - IvHA1 + HA2 + antibody
2vis, 2vit - IvHA1 (mutant) + antibody
4fqj, 4hkx - IvHA1 + antibody
Viral hemagglutinin-neuraminidase
1usx – NvPIV head domain + thiosialoside – Newcastle diasease virus
1usr, 1e8t, 1e8u, 1e8v - NvPIV head domain
3t1e - NvPIV ectodomain
1v2i, 2v3b, 1v3b – hPIV – Human parainfluenza virus
2v3c, 2v3d, 1v3c, 1v3d, 1v3e – hPIV + glucoside
2v3e – hPIV + zanamavir
1z4v, 1z4w, 1z4z, 1z50 – SvPIV extracellular domain + DANA – Simian virus
1z4x, 1z4y - SvPIV extracellular domain + glucoside
2vsk, 2vsm – HN ephrin-binding domain + ephrin – Hendra virus
2vwd, 3d11 - NivHN receptor-binding domain – Nipah virus
3d12 - NivHN receptor-binding domain + ephrin
Measles hemagglutinin
2rkc – MvHA extracellular domain – Measles virus
2zb5, 2zb6 - MvHA head domain
3alw, 3alx, 3alz – MvHA/SLAM receptor
3inb – MvHA head domain + hMembrane cofactor protein
1rwr – HA filamentous – Bordetella pertussis
Dr hemagglutinin
2w5p – EcDrae adhesin subunit – Escherichia coli
1ut1 - EcDrae adhesin subunit (mutant)
2jkj, 2jkl, 2jkn, 1usq - EcDrae adhesin subunit + chloramphenicol derivative
Phytohemagglutinin
1g8w, 1fat – PHA-L – kidney bean
Various viral hemagglutinin
1kke – RvSigma1 – human orthoreovirus
3eoy – RvSigma1 + junctional adhesion molecule
2aen – VP8 – human rotavirus
Additional Resources
For additional information, see: Influenza
References
- ↑ Gottschalk A. Chemistry of virus receptors. The Viruses. 1959;3:51–61.
- ↑ Wiley DC, Skehel JJ. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu Rev Biochem. 1987;56:365-94. PMID:3304138 doi:http://dx.doi.org/10.1146/annurev.bi.56.070187.002053
- ↑ Liu J, Stevens DJ, Haire LF, Walker PA, Coombs PJ, Russell RJ, Gamblin SJ, Skehel JJ. Structures of receptor complexes formed by hemagglutinins from the Asian Influenza pandemic of 1957. Proc Natl Acad Sci U S A. 2009 Oct 6;106(40):17175-80. Epub 2009 Sep 28. PMID:19805083
- ↑ Cell Binding protein in Avian Influenza; Jack Cerchiara,'06 and Brendan Holsberry, 07;http://biology.kenyon.edu/BMB/Chime2/2005/Cerchiara-Holsberry/FRAMES/start.htm
- ↑ 5.0 5.1 Knossow M, Skehel JJ. Variation and infectivity neutralization in influenza. Immunology. 2006 Sep;119(1):1-7. PMID:16925526 doi:http://dx.doi.org/10.1111/j.1365-2567.2006.02421.x
- ↑ See Influenza in Wikipedia.