This old version of Proteopedia is provided for student assignments while the new version is undergoing repairs. Content and edits done in this old version of Proteopedia after March 1, 2026 will eventually be lost when it is retired in about June of 2026.
Apply for new accounts at the new Proteopedia. Your logins will work in both the old and new versions.
2axi
From Proteopedia
| Line 1: | Line 1: | ||
| - | [[Image:2axi.gif|left|200px]] | + | [[Image:2axi.gif|left|200px]] |
| - | + | ||
| - | '''HDM2 in complex with a beta-hairpin''' | + | {{Structure |
| + | |PDB= 2axi |SIZE=350|CAPTION= <scene name='initialview01'>2axi</scene>, resolution 1.40Å | ||
| + | |SITE= | ||
| + | |LIGAND= <scene name='pdbligand=SO4:SULFATE+ION'>SO4</scene> and <scene name='pdbligand=MPO:3[N-MORPHOLINO]PROPANE SULFONIC ACID'>MPO</scene> | ||
| + | |ACTIVITY= | ||
| + | |GENE= MDM2 ([http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&srchmode=5&id=9606 Homo sapiens]) | ||
| + | }} | ||
| + | |||
| + | '''HDM2 in complex with a beta-hairpin''' | ||
| + | |||
==Overview== | ==Overview== | ||
| Line 7: | Line 16: | ||
==About this Structure== | ==About this Structure== | ||
| - | 2AXI is a [ | + | 2AXI is a [[Single protein]] structure of sequence from [http://en.wikipedia.org/wiki/Homo_sapiens Homo sapiens]. Full crystallographic information is available from [http://oca.weizmann.ac.il/oca-bin/ocashort?id=2AXI OCA]. |
==Reference== | ==Reference== | ||
| - | Structure-activity studies in a family of beta-hairpin protein epitope mimetic inhibitors of the p53-HDM2 protein-protein interaction., Fasan R, Dias RL, Moehle K, Zerbe O, Obrecht D, Mittl PR, Grutter MG, Robinson JA, Chembiochem. 2006 Mar;7(3):515-26. PMID:[http:// | + | Structure-activity studies in a family of beta-hairpin protein epitope mimetic inhibitors of the p53-HDM2 protein-protein interaction., Fasan R, Dias RL, Moehle K, Zerbe O, Obrecht D, Mittl PR, Grutter MG, Robinson JA, Chembiochem. 2006 Mar;7(3):515-26. PMID:[http://www.ncbi.nlm.nih.gov/pubmed/16511824 16511824] |
[[Category: Homo sapiens]] | [[Category: Homo sapiens]] | ||
[[Category: Single protein]] | [[Category: Single protein]] | ||
| Line 21: | Line 30: | ||
[[Category: drug design]] | [[Category: drug design]] | ||
[[Category: p53]] | [[Category: p53]] | ||
| - | [[Category: protein-protein | + | [[Category: protein-protein interaction]] |
| - | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Thu | + | ''Page seeded by [http://oca.weizmann.ac.il/oca OCA ] on Thu Mar 20 15:55:01 2008'' |
Revision as of 13:55, 20 March 2008
| |||||||
| , resolution 1.40Å | |||||||
|---|---|---|---|---|---|---|---|
| Ligands: | and | ||||||
| Gene: | MDM2 (Homo sapiens) | ||||||
| Coordinates: | save as pdb, mmCIF, xml | ||||||
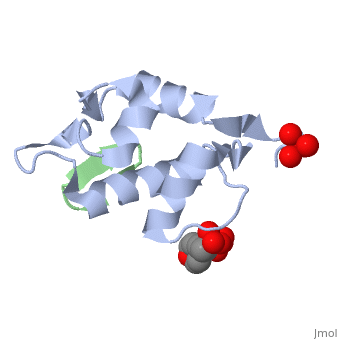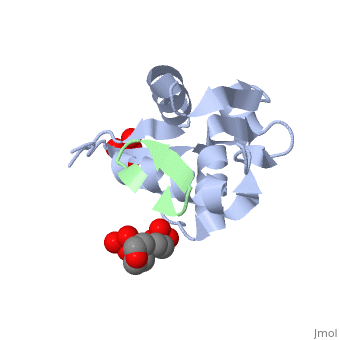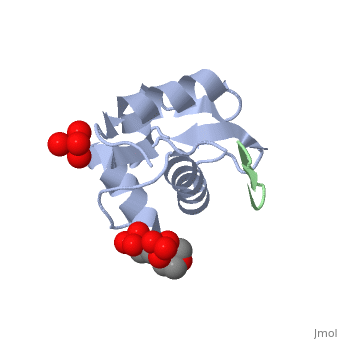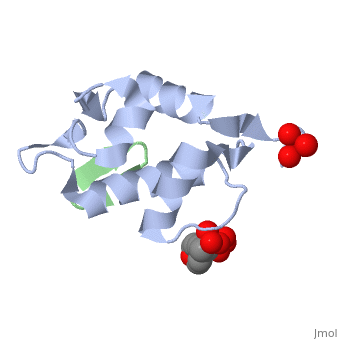
HDM2 in complex with a beta-hairpin
Overview
Inhibitors of the interaction between the p53 tumor-suppressor protein and its natural human inhibitor HDM2 are attractive as potential anticancer agents. In earlier work we explored designing beta-hairpin peptidomimetics of the alpha-helical epitope on p53 that would bind tightly to the p53-binding site on HDM2. The beta-hairpin is used as a scaffold to display energetically hot residues in an optimal array for interaction with HDM2. The initial lead beta-hairpin mimetic, with a weak inhibitory activity (IC(50)=125 microM), was optimized to afford cyclo-(L-Pro-Phe-Glu-6ClTrp-Leu-Asp-Trp-Glu-Phe-D-Pro) (where 6ClTrp=L-6-chlorotryptophan), which has an affinity almost 1,000 times higher (IC(50)=140 nM). In this work, insights into the origins of this affinity maturation based on structure-activity studies and an X-ray crystal structure of the inhibitor/HDM2(residues 17-125) complex at 1.4 A resolution are described. The crystal structure confirms the beta-hairpin conformation of the bound ligand, and also reveals that a significant component of the affinity increase arises through new aromatic/aromatic stacking interactions between side chains around the hairpin and groups on the surface of HDM2.
About this Structure
2AXI is a Single protein structure of sequence from Homo sapiens. Full crystallographic information is available from OCA.
Reference
Structure-activity studies in a family of beta-hairpin protein epitope mimetic inhibitors of the p53-HDM2 protein-protein interaction., Fasan R, Dias RL, Moehle K, Zerbe O, Obrecht D, Mittl PR, Grutter MG, Robinson JA, Chembiochem. 2006 Mar;7(3):515-26. PMID:16511824
Page seeded by OCA on Thu Mar 20 15:55:01 2008